Novel Hydrolytic Enzymes and Protein Modelling from Marine Sedimentary Bacteria Losari Coastal, Makassar South Sulawesi, Indonesia
DOI:
https://doi.org/10.48048/tis.2025.10574Keywords:
Marine sediment, Bacteria, Amylase, Lipase, Protease, Losari coastalAbstract
Marine ecosystems are invaluable sources of biodiversity and unique chemical compounds, especially for the discovery of new natural products, including enzymes. Marine sediment bacteria from the Losari Coastal, Makassar City, Indonesia, show great potential for biotechnological applications due to their ability to produce enzymes with extraordinary properties such as salt tolerance, heat stability, and adaptability to extreme conditions. This study investigated the enzymatic capabilities of marine sediment bacteria isolated from the Losari Coastal, Makassar City, Indonesia, focusing on 3 enzymes: Amylase, lipase, and protease. Bacterial samples were collected from 4 coastal locations, and a comprehensive analysis of their morphological, physiological, and molecular characteristics was performed, combining 16S rRNA sequencing and protein modeling using SWISS-MODEL. This study yielded 15 bacterial isolates characterized by a gram-negative, rod-shaped morphology and distinctive, round, yellow colonies with serrated edges. Among these, 10 produced amylase, 9 produced lipases, and 10 showed protease activity; however, 2 isolates stood out because they could produce all three enzymes: amylase, lipase, and protease. The MB-SL 5 isolate showed 97.82% similarity to Shewanella algae strain DW01, and the MB-SL 6 isolate had 90.53% similarity to Vibrio alginolyticus NBRC 15630 strain, with predicted protein structure code A0A3M5GRJ4_1 A. These findings highlight the significant biotechnological potential of marine sediment bacteria from the Losari Coastal producing hydrolytic enzymes for industrial applications, which opens new avenues for the development of new biocatalysts.
HIGHLIGHTS
- Pioneering Exploration of the Ecosystem as an Optimal Source for Enzyme-Producing Bacteria: This study represents the 1st comprehensive investigation of marine sedimentary bacteria from the unique Losari Coastal ecosystem in Makassar City, Indonesia, revealing previously uncharacterized enzymatic capabilities.
- Multi-Enzyme Production in Indigenous Bacterial Strains: The discovery of local bacterial strains capable of producing all three industrially valuable enzymes (amylase, lipase, and protease) simultaneously represents a significant advancement in biocatalyst research.
- Integration of Advanced Structural Prediction: The novel combination of 16S rRNA sequencing with SWISS-MODEL protein modeling provides unprecedented structural insights into the enzymes produced by these marine bacteria.
- Identification of Specific Protein Structure Codes: The research uniquely assigns specific protein structure codes (A0A3M5GRJ41.A protein) to the enzymes produced by algae and V. alginolyticus, enabling precise characterization for future applications.
- Enzyme Properties from a Tropical Coastal Source: The study reveals the unexpected presence of enzyme properties (salt tolerance, temperature stability) in bacteria from a tropical coastal environment rather than traditional extreme habitats, expanding our understanding of where such valuable biocatalysts can be sourced.
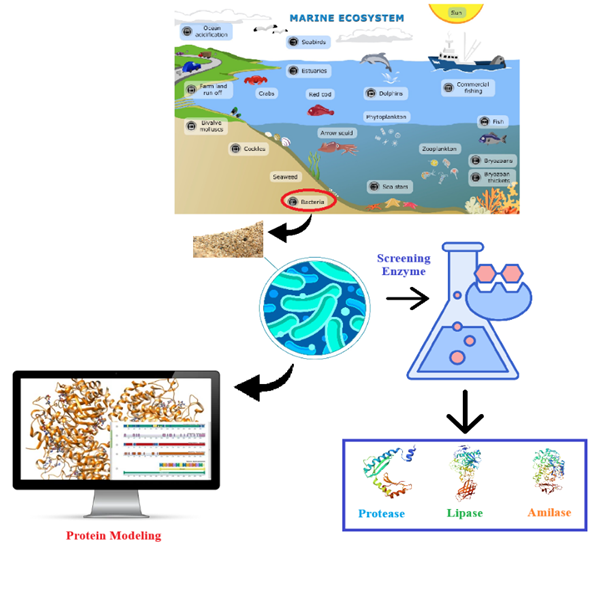
GRAPHICAL ABSTRACT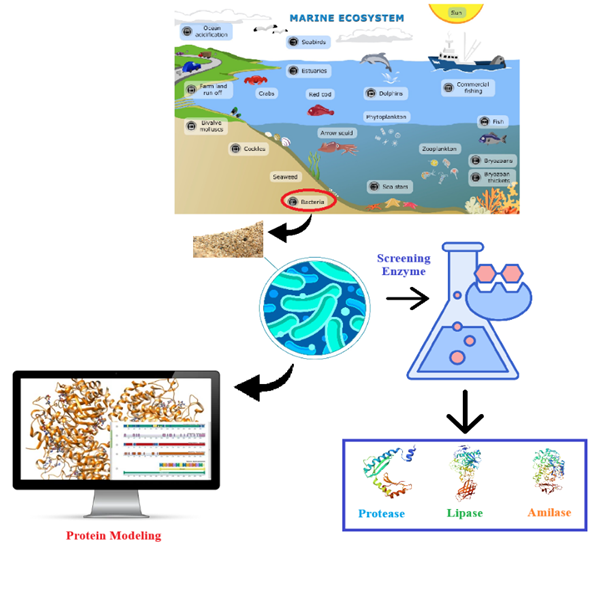
Downloads
References
AD Rogers, W Appeltans, J Assis, LT Ballance, P Cury, C Duarte, F Favoretto, LA Hynes, JA Kumagai, CE Lovelock, P Miloslavich, A Niamir, D Obura, BC O’Leary, E Ramirez-Llodra, G Reygondeau, C Roberts, Y Sadov, O Steeds, T Sutton, …, O Aburto-Oropeza. Discovering marine biodiversity in the 21st century. Advances in Marine Biology 2022; 93, 23-115.
A Karthikeyan, A Joseph and BG Nair. Promising bioactive compounds from the marine environment and their potential effects on various diseases. Journal of Genetic Engineering and Biotechnology 2022; 20(1), 14.
DL Dayanidhi, BC Thomas, JS Osterberg, M Vuong, G Vargas, SK Kwartler, E Schmaltz, MM Dunphy-Daly, TF Schultz, D Rittschof, WC Eward, C Roy and JA Somarelli. Exploring the diversity of the marine environment for new anti-cancer compounds. Frontiers in Marine Science 2021; 7, 614766.
F Ameen, S AlNadhari and AA Al-Homaidan. Marine microorganisms as an untapped source of bioactive compounds. Saudi Journal of Biological Sciences 2021; 28(1), 224-231.
S Ghattavi and A Homaei. Marine enzymes: Classification and application in various industries. International Journal of Biological Macromolecules 2023; 230, 123136.
AH Banday, NU Azha, R Farooq, SA Sheikh, MA Ganie, MN Parray, H Mushtaq, I Hameed and MA Lone. Exploring the potential of marine natural products in drug development: A comprehensive review. Phytochemistry Letters 2024; 59, 124-135.
J Zhang, L Jiang, X Chen, K Lv, M Basiony, G Zhu, L Karthik, L Ouyang, L Zhang and X Liu. Recent advances in biotechnology for marine enzymes and molecules. Current Opinion in Biotechnology 2021; 69, 308-315.
S Bhandari, DK Poudel, R Marahatha, S Dawadi, K Khadayat, S Phuyal, S Shrestha, S Gaire, K Basnet, U Khadka and N Parajuli. Microbial enzymes used in bioremediation. Journal of Chemistry 2021; 2021(1), 8849512.
S Raveendran, B Parameswaran, SB Ummalyma, A Abraham, AK Mathew, A Madhavan, S Rebello and A Pandey. Applications of microbial enzymes in food industry. Food Technology and Biotechnology 2018; 56(1), 16-30.
N Kochhar, IK Kavya, S Shrivastava, A Ghosh, VS Rawat, KK Sodhi and M Kumar. Perspectives on the microorganisms of extreme environments and their applications. Current Research in Microbial Sciences 2022; 3, 100134.
A Trincone. Enzymatic processes in marine biotechnology. Marine Drugs 2017; 15(4), 93.
K Maldonado-Ruiz, R Pedroza-Islas and L Pedraza-Segura. Blue biotechnology: Marine bacteria bioproducts. Microorganisms 2024; 12(4), 697.
E Nikolaivits, M Dimarogona, N Fokialakis and E Topakas. Marine-derived biocatalysts: Importance, accessing, and application in aromatic pollutant bioremediation. Frontiers in Microbiology 2017; 8, 265.
P Solanki, C Putatunda, A Kumar, R Bhatia and A Walia. Microbial proteases: Ubiquitous enzymes with innumerable uses. 3 Biotech 2022; 11(10), 428.
A Kumar, S Dhiman, B Krishan, M Samtiya, A Kumari, N Pathak, A Kumari, RE Aluko and T Dhewa. Microbial enzymes and major applications in the food industry: A concise review. Food Production, Processing and Nutrition 2024; 6, 85.
DS Kocabaş, J Lyne and Z Ustunol. Hydrolytic enzymes in the dairy industry: Applications, market and future perspectives. Trends in Food Science and Technology 2022; 119, 467-475.
A Fasim, VS More and SS More. Large-scale production of enzymes for biotechnology uses. Current Opinion in Biotechnology 2021; 69, 68-76.
W Yang, F Lu and Y Liu. Recent advances of enzymes in the food industry. Foods 2023; 12(24), 4506.
J John. Biochemical and biotechnological aspects of microbial amylases. In: R Goldbeck and P Poletto (Eds.). Polysaccharide-degrading biocatalysts. Academic Press, Cambridge, 2023, p. 191-204.
R Seth and A Meena. Enzyme-based nanomaterial synthesis: An eco-friendly and green synthesis approach. Clean Technologies and Environmental Policy 2024. https://doi.org/10.1007/s10098-024-02854-7.
AA Al-Ghanayem and B Joseph. Current prospects in using cold-active enzymes as eco-friendly detergent additives. Applied Microbiology and Biotechnology 2020; 104(7), 2871-2882.
D Kumar, R Bhardwaj, S Jassal, T Goyal, A Khullar and N Gupta. Application of enzymes for an eco-friendly approach to textile processing. Environmental Science and Pollution Research International 2023; 30(28), 71838-71848.
Y Khambhaty. Applications of enzymes in leather processing. Environmental Chemistry Letters 2020; 18(3), 747-769.
GK Gupta, M Dixit, RK Kapoor and P Shukla. Xylanolytic enzymes in pulp and paper industry: New technologies and perspectives. Molecular Biotechnology 2022; 64(2), 130-143.
KKR Shah, S Devanshi, GB Patel and VD Patel. Application of microbial enzymes: Biodegradation of paper and pulp waste, in innovations in environmental biotechnology. In: S Arora, A Kumar, S Ogita and YY Yau (Eds.). Springer Nature Singapore, Singapore, 2022, p. 283-304.
N Soni and MS Madhusudhan. Computational modeling of protein assemblies. Currnt Opinion in Structural Biology 2017; 44, 179-189.
Y Kumar, H Ramesh, K Dhabade, M Shahare and B Kalra. Structural study and molecular docking insights into laccase-mediated dye degradation. Sustainable Chemistry for the Environment 2024; 8, 100175.
K Nam, Y Shao, DT Major and M Wolf-Watz. Perspectives on computational enzyme modeling: From mechanisms to design and drug development. ACS Omega 2024; 9(7), 7393-7412.
K Grigorakis, C Ferousi and E Topakas. Protein engineering for industrial biocatalysis: Principles, approaches, and lessons from engineered PETases. Catalysts 2025; 15(2), 147.
S Mao, J Jiang, K Xiong, Y Chen, Y Yao, L Liu, H Liu and X Li. Enzyme engineering: Performance optimization, novel sources and applications in the food industry. Foods 2024; 13(23), 3846.
L Lin, D Xiao, W Song and W Lu. A breakthrough computational strategy for efficient enzymatic digestion of walnut protein to prepare antioxidant peptides. Food Chemistry 2025; 476, 143311.
R Ruginescu, P Lavin, L Iancu, S Menabit and C Purcarea. Bioprospecting for novel bacterial sources of hydrolytic enzymes and antimicrobials in the romanian littoral zone of the black sea. Microorganisms 2022; 10(12), 12-26.
SV Jagannathan, EM Manemann, SE Rowe, MC Callender and W Soto. Marine actinomycetes, new sources of biotechnological products. Marine Drugs 2021; 19(7), 365.
H Natsir, A Ahmad, N Massi, P Taba, Anita and W Rauf. Isolation, production of protease, and antimicrobial activities from marine sediment gamma - proteobacteria of MBS-L3 isolate. Research Journal of Pharmacy and Technology 2024; 17(6), 2855-2862.
SA Fasiku, OF Ogunsola, A Fakunle and AA Olanbiwoninu. Isolation of bacteria with potential of producing extracellular enzymes (amylase, cellulase and protease) from soil samples. Journal of Advances in Microbiology 2020; 20(3), 21-26.
P Gómez-Villegas, J Vigara, L Romero, C Gotor, S Raposo, B Gonçalves and R Léon. Biochemical characterization of the amylase activity from the new Haloarchaea strain Haloarcula sp. HS isolated in the odiel marshlands. Biology 2021; 10(4), 337.
J Carrazco-Palafox, BE Rivera-Chavira, N Ramírez-Baca, LI Manzanares-Papayanopoulos and GV Nevárez-Moorillón. Improved method for qualitative screening of lipolytic bacterial strains. MethodsX 2018; 5, 68-74.
OI Ilesanmi, AE Adekunle, JA Omolaiye, EM Olorode and AL Ogunkanmi. Isolation, optimization and molecular characterization of lipase-producing bacteria from contaminated soil. Scientific African 2020; 8, e00279.
SK Marathe, MA Vashistht, A Prashanth, N Parveen, S Chakraborty and SS Nair. Isolation, partial purification, biochemical characterization and detergent compatibility of alkaline protease produced by Bacillus subtilis, Alcaligenes faecalis and Pseudomonas aeruginosa obtained from sea water samples. Journal of Genetic Engineering and Biotechnology 2018; 16(1), 39-46.
C Masi, G Gemechu and M Tafesse. Isolation, screening, characterization, and identification of alkaline protease-producing bacteria from leather industry effluent. Annals Microbiology 2021; 71, 24.
E Rosa, CN Ekowati, TT Handayani, A Ikhsanudin, F Apriliani and A Arifiyanto, Characterization of entomopathogenic fungi as a natural biological control of American cockroaches (Periplaneta americana). Biodiversitas 2020; 21(11), 5276-5282.
DL Church, L Cerutti, A Gürtler, T Griener, A Zelazny and S Emler. Performance and application of 16S rRNA gene cycle sequencing for routine identification of bacteria in the clinical microbiology laboratory. Clinical Microbiology Reviews 2020; 33(4), e00053-19.
R Srinivasan, U Karaoz, M Volegova, J MacKichan, M Kato-Maeda, S Miller, R Nadarajan, EL Brodie and SV Lynch. Use of 16S rRNA gene for identification of a broad range of clinically relevant bacterial pathogens. PLoS One 2015; 10(2), e0117617.
AK Farha, TR Thasneem, A Purushothaman, JA Salam and AM Hatha. Phylogenetic diversity and biotechnological potentials of marine bacteria from the continental slope of eastern Arabian Sea. Journal, Genetic Engineering and Biotechnology 2018; 16(2), 253-258.
EH Sanjaya, S Suharti, M Alvionita, I Telussa, S Febriana and H Clevanota. Isolation and characterization of amylase enzyme produced by indigenous bacteria from sugar factory waste. The Open Biotechnology Journal 2024; 18, e18740707296261.
PT Nnaji, E Adukwu, HR Morse and RU Chidugu-Ogborigbo. Amylase production from marine sponge Hymeniacidon perlevis; potential sustainability benefits. PLoS One 2023; 18(12), e0294931.
S Pesek and R Silaghi-Dumitrescu. The iodine/iodide/starch supramolecular complex. Molecules 2024; 29(3), 641.
A Budiharjo, D Wulandari, J Shabrina, RA Mawarni, AR Maulana, Nurhayati, W Wijanarka, L Hartajanie and Lindayani. Bioprospecting and molecular identification of amylase and cellulase producing thermophilic bacteria from sediment of Nglimut Hot Springs, Kendal Regency. Journal of Tropical Biodiversity and Biotechnology 2024; 9(3), jtbb86756.
P Chandra, Enespa, R Singh and PK Arora. Microbial lipases and their industrial applications: A comprehensive review. Microbial Cell Factories 2020; 19, 169.
D Bharathi and G Rajalakshmi. Microbial lipases: An overview of screening, production and purification. Biocatalysis and Agricultural Biotechnology 2019; 22, 101368.
S Lanka and JNL Latha. A short review on various screening methods to isolate potential lipase producers: Lipases-the present and future enzymes of biotech industry. International Journal of Biological Chemistry 2015; 9(5), 207-219.
YA Lemenh, TG Biru, AZ Chernet and FB Lema. Isolation and identification of protease-producing bacteria from sludge and sediment soil around Adama, Ethiopia. Indonesian Journal Biotechnology 2025; 26(4), 159.
P Song, X Zhang, S Wang, W Xu, F Wang, R Fu and F Wei. Microbial proteases and their applications. Frontiers in Microbiology 2023; 14(9), 1236368.
DE Cruz-Casas, CN Aguilar, JA Ascacio-Valdés, R Rodríguez-Herrera, ML Chávez-González and AC Flores-Gallegos. Enzymatic hydrolysis and microbial fermentation: The most favorable biotechnological methods for the release of bioactive peptides. Food Chemistry: Molecular Sciences 2021; 3, 100047.
J Su, Y Song, Z Zhu, X Huang, J Fan, J Qiao and F Mao. Cell-cell communication: New insights and clinical implications. Signal Transduction and Targeted Therapy 2024; 9, 196.
Y Hong, A Boiti, D Vallone and NS Foulkes. Reactive oxygen species signaling and oxidative stress: Transcriptional regulation and evolution. Antioxidants 2024; 13(3), 164-176.
N Prihatiningsih, A Asnani and HA Djatmiko. Extracellular protease from Bacillus subtilis B315 with antagonistic activity against bacterial wilt pathogen (Ralstonia solanacearum) of chili. Biodiversitas 2022; 22(3), 1291-1295.
M Dehimi, F Yusof, RA Raus, NF Hadry and R Nedjai. Genotypic and phenotypic characterisation of isolated marine bacteria and their potential to produce alkaline protease. Journal of Advanced Research in Applied Sciences and Engineering Technology 2021; 22(1), 16-25.
A Abdel-Latif and G Osman. Comparison of three genomic DNA extraction methods to obtain high DNA quality from maize. Plant Methods 2017; 13, 1.
W Qamar, MR Khan and A Arafah. Optimization of conditions to extract high-quality DNA for PCR analysis from whole blood using SDS-proteinase K method. Saudi Journal of Biological Sciences 2017; 24(7), 1465-1469.
HR Dash, P Shrivastava and S Das. Quantification of DNA by using agarose gel electrophoresis technique. In: HR Dash, P Shrivastava and S Das (Eds.). Principles and practices of DNA analysis: A laboratory manual for forensic DNA typing. Springer Protocols Handbooks, Humana, New York, 2020, p. 119-125.
R Veneziano, TR Shepherd, S Ratanalert, L Bellou, C Tao and M Bathe. In vitro synthesis of gene-length single-stranded DNA. Scientific Reports 2018; 8(1), 6548.
J Philips, L Procopio and IPG Marshall. Insights into the various mechanisms by which Shewanella spp. induce and inhibit steel corrosion. NPJ Materials Degradation 2023; 7, 95.
Z Wang, J Zhao, L Wang, C Li, J Liu, L Zhang and Y Zhang. A novel benthic phage infecting shewanella with strong replication ability. Viruses 2019; 11(11), 1081.
HI Sheikh, NII Alhamadin, HJ Liew, A Fadhlina, MEA Wahid, N Musa and KCA Jalal. Virulence factors of the zoonotic pathogen Vibrio alginolyticus: A review and bibliometric analysis. Applied Biochemistry and Microbiology 2024; 60, 514-531.
KMJ Slifka, AE Newton and BE Mahon. Vibrio alginolyticus infections in the USA, 1988 - 2012. Epidemiology and Infection 2017; 45(7), 1491-1499.
J Mamangkey, D Suryanto, E Munir, AZ Mustopa, MT Sibero, LW Mendes, A Hartanto, S Taniwan, MJ Ek-Ramos, A Harahap, A Verma, E Trihatmoko, WS Putranto, L Pardosi and LOAP Rudia. Isolation and enzyme bioprospection of bacteria associated with Bruguiera cylindrica, a mangrove plant of North Sumatra, Indonesia. Biotechnology Reports 2021; 30, e00617.
M Salamone, A Nicosia, G Ghersi and M Tagliavia. Vibrio proteases for biomedical applications: Modulating the proteolytic secretome of V. alginolyticus and V. parahaemolyticus for improved enzymes production. Microorganisms 2019; 7(10), 387.
R Shunmugam, SR Balusamy, V Kumar, S Menon, T Lakshmi and H Perumalsamy. Biosynthesis of gold nanoparticles using marine microbe (Vibrio alginolyticus) and its anticancer and antioxidant analysis. Journal of King Saud University - Science 2021; 33(1), 101260.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






