Novel Anti-Cancer Agents: Potential of Protease Enzymes from Vibrio alginolyticus as a Breast Cancer Therapy for MCF-7 Cells
DOI:
https://doi.org/10.48048/tis.2025.10450Keywords:
Vibrio alginolyticus, Protease, Cytotoxicity, MCF-7 cell, Molecular dockingAbstract
This study explores the potential of Vibrio alginolyticus protease enzyme as a cytotoxic agent against MCF-7 breast cancer cells, using molecular docking and cytotoxicity assays. The docking analysis revealed that the native ligand interacts strongly with key residues at the active site of the target protein, including hydrogen bonds with residues ARG394 (2.034 Å) and GLU353 (1.920 Å), as well as hydrophobic interactions with residues PHE404, LEU391, MET421, and ILE424, with distances ranging from 4.664 Å to 5.355 Å. These interactions demonstrate a high affinity between the ligand and the target protein, indicating potential for enzyme-based cancer therapy development. Cytotoxicity testing using the resazurin assay showed that V. alginolyticus protease enzyme exhibited an IC50 value of 1.63 μg/mL, indicating significant cytotoxic effects on MCF-7 cells. At concentrations above 2.5 μg/mL, the enzyme demonstrated strong cytotoxic effects, with morphological changes in the cells, such as shrinkage and detachment. In contrast, at lower concentrations, the cytotoxic activity was limited. The observed decrease in cell viability at high concentrations suggests that the enzyme works by disrupting key proteins or pathways involved in cancer cell proliferation and survival. Both in silico and in vitro analyses support the possibility that V. alginolyticus protease targets essential proteins involved in apoptosis regulation and structural protein degradation in cancer cells. These findings provide new insights into the development of enzyme-based cancer therapies, warranting further research to understand the specific mechanisms behind cancer cell death induction.
HIGHLIGHTS
- Vibrio alginolyticus protease demonstrated strong binding affinity to MCF-7 breast cancer target proteins through molecular docking, showing key interactions at ARG394 and GLU353.
- Hydrophobic and hydrogen bond interactions between the protease and residues PHE404, LEU391, MET421, and ILE424 support its potential as a protein-targeting therapeutic.
- Cytotoxicity assays revealed a low IC50 value of 1.63 μg/mL, indicating potent cytotoxic effects against MCF-7 cells at relatively low concentrations.
- Both in silico and in vitro results suggest apoptosis regulation and structural protein degradation as possible mechanisms of cancer cell death induction by the enzyme.
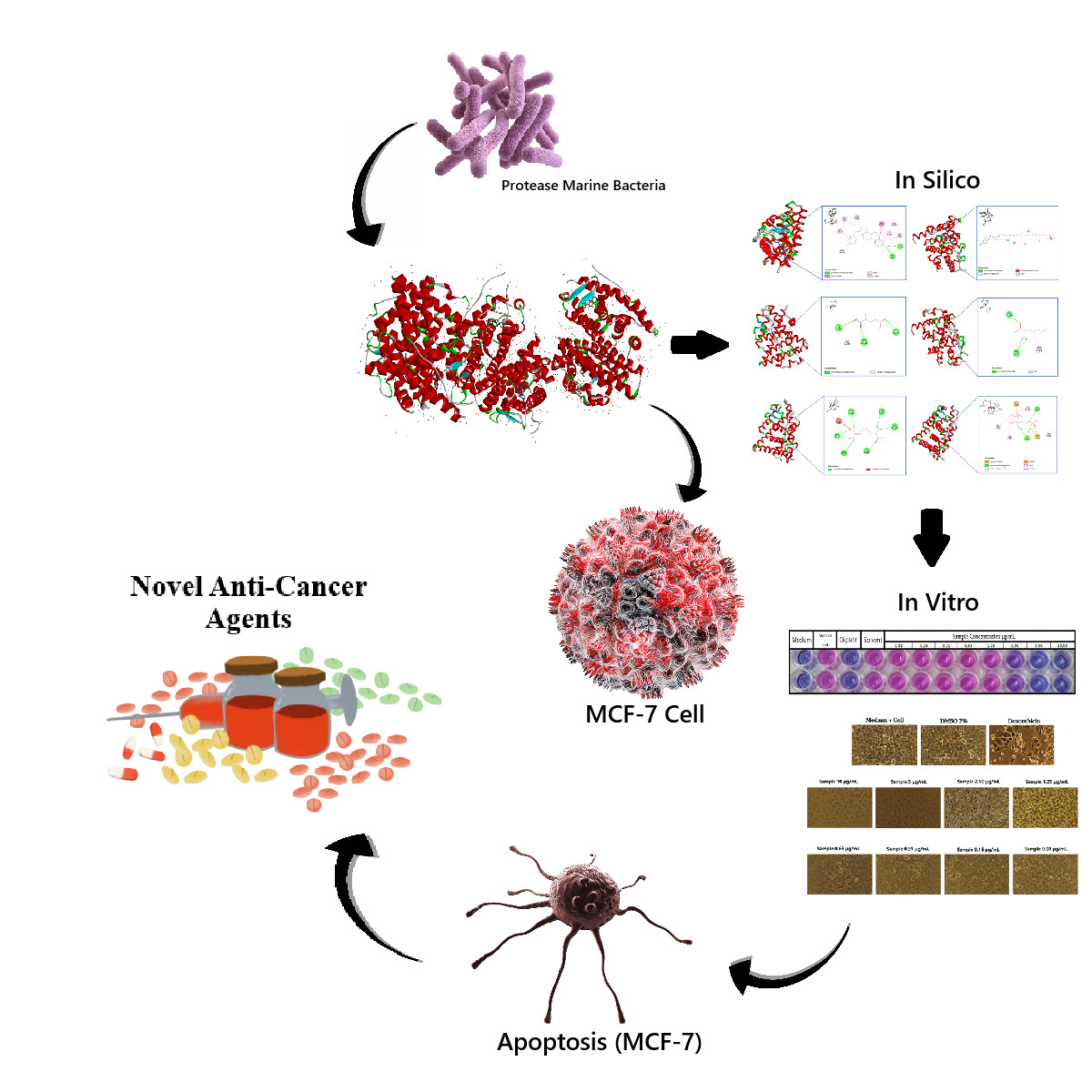
GRAPHICAL ABSTRACT
Downloads
References
A Saini, M Kumar, S Bhatt, V Saini and A Malik. Cancer : Cancer is a disorder. International Journal of Pharmaceutical Sciences and Research 2020; 11, 3121-3134.
FH Al-Ostoot, S Salah and SA Khanum. An overview of cancer biology, pathophysiological development and It’s treatment modalities: Current challenges of cancer anti-angiogenic therapy. Cancer Investigation 2024; 42(7), 559-604.
M He, X Zhou, and X Wang. Glycosylation: Mechanisms, biological functions and clinical implications. Signal Transduction and Targeted Therapy 2024; 9(1), 2024.
GR Bhat, I Sethi, HQ Sadida, B Rah, R Mir, N Algehainy, IA Albalawi, T Masoodi, GK Subbaraj, F Jamal, M Singh, R Kumar, MA Macha, S Uddin, ASA Akil, M Haris and AA Bhat Cancer cell plasticity: From cellular, molecular, and genetic mechanisms to tumor heterogeneity and drug resistance. Cancer and Metastasis Reviews 2024; 43(1), 197-228.
SK Kim, E Brotslaw, V Thome, J Mitchell, R Ventrella, C Collins and B Mitchell. A role for Cep70 in centriole amplification in multiciliated cells. Developmental Biology 2021; 471, 10-17.
X An, W Yu, J Liu, D Tang, L Yang and X Chen. Oxidative cell death in cancer: Mechanisms and therapeutic opportunities. Cell Death & Disease 2024; 15(8), 556.
U Anand, A Dey, AKS Chandel, R Sanyal, A Mishra, DK Pandey, VD Falco, A Upadhyay, R Kandimalla, A Chaudhary, JK Dhanjal, S Dewanjee, J Vallamkondu and JMPLD Lastra. Cancer chemotherapy and beyond: Current status, drug candidates, associated risks and progress in targeted therapeutics. Genes 2023; 10(4), 1367-1401.
DT Debela, SGY Muzazu, KD Heraro, MT Ndalama, BW Mesele, DC Haile, SK Kitui and T Manyazewal. New approaches and procedures for cancer treatment: Current perspectives. SAGE Open Medicine 2021; 9, 20503121211034366.
A Saini, M Kumar, S Bhatt, V Saini and A Malik. Cancer causes and treatments. International Journal of Pharmaceutical Sciences and Research 2020; 11(7), 3121-3134.
N Behranvand, F Nasri, RZ Emameh, P Khani, A Hosseini, J Garssen and R Falak. Chemotherapy: A double-edged sword in cancer treatment. Cancer Immunology, Immunotherapy 2022; 71(3), 507-526.
A Shah, A Mushtaq, F Mandokhail, S Munir and SA Ali. A review on breast cancer , risk factors , symptoms and some common treatments. SBK Journal of Basic Sciences and Innovative Research 2021; 1(1), 34-41.
AK Das, SK Biswas, A Bhattacharya and E Alam. Introduction to breast cancer and awareness. In: Proceedings of the 7th International Conference on Advanced Computing and Communication Systems, Coimbatore, India, 2021.
S Sharma, M Chakraborty, D Yadav, A Dhullap, R Singh, RK Verma, S Bhattacharya and S Singh. Strategic developments in polymer-functionalized liposomes for targeted colon cancer therapy: An updated review of clinical trial data and future horizons. Biomacromolecules 2024; 25(9), 5650-5669.
KA Kasgri, M Abazari, SM Badeleh, KM Badeleh and N Peyman. Comprehensive review of breast cancer consequences for the patients and their coping strategies: A systematic review. Cancer Control 2024; 31, 10732748241249355.
S Comsa, AM Cimpean and M Raica. The story of MCF-7 breast cancer cell line: 40 Years of experience in research. Anticancer Research 2015; 35(6), 3147-3154.
AS Levenson and VC Jordan. MCF-7: The first hormone-responsive breast cancer cell line. Cancer Research 1997; 57(15), 3071-3078.
H Natsir, A Ahmad, N Massi, P Taba, Anita, and W Rauf. Isolation, production of protease, and antimicrobial activities from marine sediment gamma-proteobacteria of MBS-L3 isolate. Research Journal of Pharmacy and Technology 2024; 17(6), 2855-2862.
E Pratiwi, I Raya, H Natsir, R Irfandi, P Taba, R Arfah, H Rasyid, Y Hala, S Kasim, AB Khaerunnisa, B Ilham, M Mazaya, Y Tanzil and D Luthfiana. Investigations of Ni(II)cysteine-tyrosine dithiocarbamate complex: Synthesis, characterization, molecular docking, molecular dynamic, and anticancer activity on MCF-7 breast cancer cell line. Asian Pacific Journal of Cancer Prevention 2024; 25(4), 1301-1313.
TT Ogunjobi, IC Okorie, CD Gigam-Ozuzu, JV Olorunleke, FI Ogunleye, EO Irimoren, DO Atanda, AM Okafor, CE Agbo, FO Okunbi, OD Umoren, AD Adidu and EO Ojo. Bioinformatics tools in protein analysis: Structure prediction, interaction modelling, and function relationship. European Journal of Sustainable Development Research 2024; 8(1), em0298.
SU Khan, H Wuryastuty, MH Wibowo, S Sarmin and SH Irianingsih. Genetic analyses of the structural protein E2 bovine viral diarrhea virus isolated from dairy cattle in Yogyakarta, Indonesia. Veterinary World 2024; 17(7), 1562-1574.
Yuliza, A Salamah and H Puspitaningrum. Exploration of Oryza sativa drought-responsive element binding protein 2A (OsDREB2A) gene in several local Indonesian rice varieties. Biodiversitas 2024; 25(2), 574-582.
SV Pawar, WSK Banini, MM Shamsuddeen, TA Jumah, NNO Dolling, A Tiamiyu and OI Awe. Prostruc: An open-source tool for 3D structure prediction using homology modeling. Frontiers in Chemistry 2024; 12, 1-13.
S Zarougui, M Er-Rajy, A Faris, H Imtara, ME Fadili, AA Qurtam, FA Nasr, M Al-Zharani and M Elhallaoui. 3D computer modeling of inhibitors targeting the MCF-7 breast cancer cell line. Frontiers in Chemistry 2024; 12, 1-18.
SE Rhabori, M Alaqarbeh, AE Aissouq, M Bouachrine, S Chtita and F Khalil. Design, 3D-QSAR, molecular docking, ADMET, molecular dynamics and MM-PBSA simulations for new anti-breast cancer agents. Chemical Physics Impact 2024; 8, 100455.
P Kamankesh and Z Ghayedi. Archive of SID. International Journal of Engineering Science 2023; 12(47), 84-92.
K Kavitha, JV Jothikanth, KP Kaniga, KS Manoj, RP Revathi, ER Esakiprasanth, MS Mohan, SNL Gowri, JR Jubilee, SM Srinithi and KH Lokesh. Synthesis, molecular docking, in silico druglikeness: In vitro cytotoxicity study on MCF-7 cell line of quinazolin-4-one scaffold. Journal of Young Pharmacists 2024; 16(4), 725-734.
M Azmal, JK Paul, FS Prima, OF Talukder and A Ghosh. An in silico molecular docking and simulation study to identify potential anticancer phytochemicals targeting the RAS signaling pathway. PLoS One 2024; 19(9), e0310637.
NM Abd-El-Aziz, MS Hifnawy, RA Lotfy and IY Younis. LC/MS/MS and GC/MS/MS metabolic profiling of Leontodon hispidulus, in vitro and in silico anticancer activity evaluation targeting hexokinase 2 enzyme. Scientific Reports 2024; 14(1), 6872.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






