Effect of Mitochondrial DNA T15663C Mutation on Type 2 Diabetes Mellitus and Cataract Patients, Molecular Dynamics Simulation Study
DOI:
https://doi.org/10.48048/tis.2024.8195Keywords:
T15663C mutation, Molecular dynamics, Cytochrome b, Type 2 diabetes mellitusAbstract
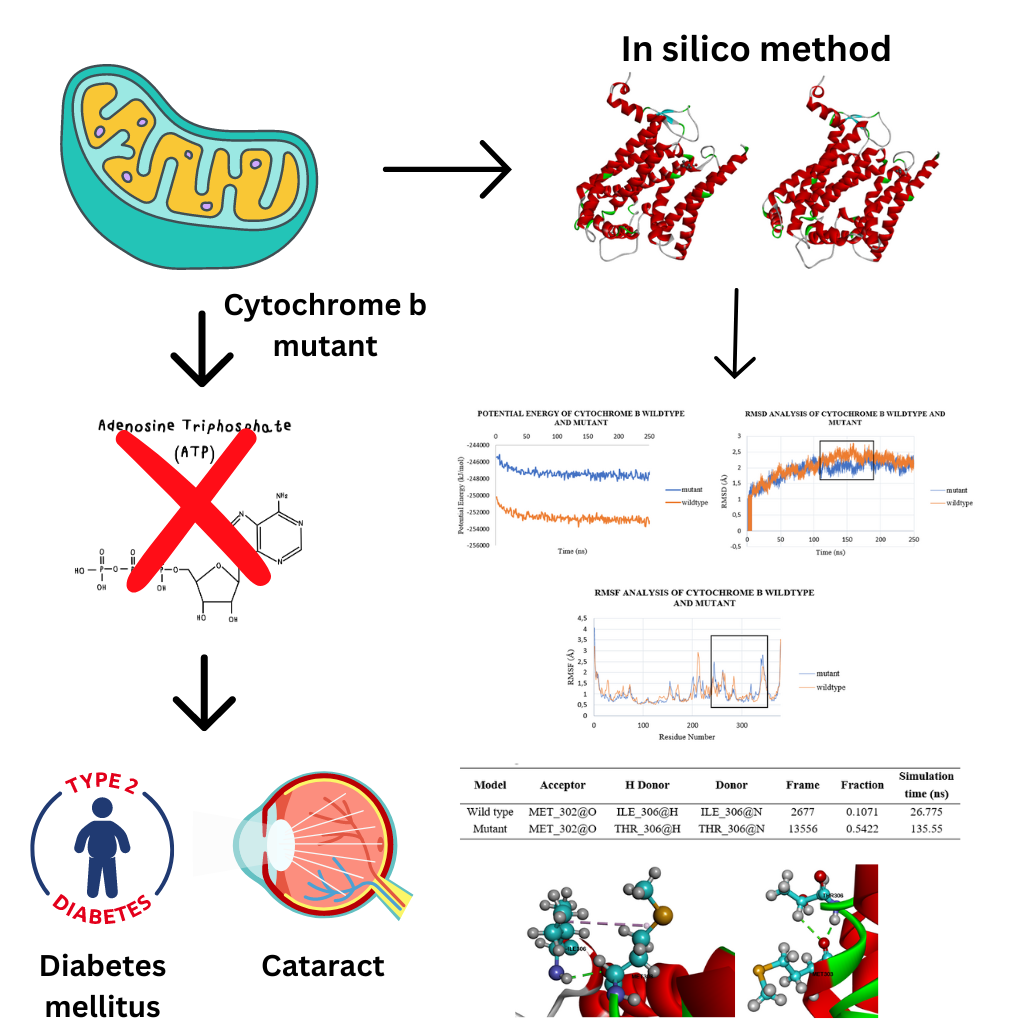
Diabetes mellitus is a metabolic disease characterized by hyperglycemia. Type 2 diabetes mellitus patients are known to have mutations in their mitochondrial DNA (mtDNA). mtDNA easily undergoes mutations because it does not have histone proteins for protection and lacks a proofreading mechanism during its replication. The mtDNA mutations T15663C found in type 2 diabetes mellitus patients sufferers with a cataract disease. This mutation was found to occur in the cytochrome b gene, one of the gene in the complex III respiratory chain. In this study we investigate the effect of this mutation on the structure of cytochrome b using molecular dynamics simulations using PDB ID: 5XTE as a template and carrying out simulations for 250 ns. The models of wild type and mutant protein structure were constructed using CHARMM-Gui so that, the protein was looked like in the mitochondrial membrane. The results of the molecular dynamics simulation show that the potential energy of the wild type model is lower than the mutant model, the RMSD value of the wild type model is more stable at 114 - 180 ns, and the RMSF value in the wild type model does not fluctuate as much as the mutant model. In addition, the hydrogen bond (H bond) analysis of the mutant model has a higher H bond number than the wild type model, which means that the mutant model is more rigid. The simulation results were visualized using VMD, and new hydrogen bonds were found in the mutant model.
HIGHLIGHTS
- Mitochondrial mutations can lead to many diseases such as type 2 diabetes mellitus and cataracts.
- Molecular dynamics simulations can be used to see the differences between wild type and mutant structures.
- Potential energy, Root Mean Square Deviation (RMSD), Root Mean Square Fluctuation (RMSF) and hydrogen bond (H bond) analysis commonly used to analyze simulation results, such as the stability or rigidity of the structure.
GRAPHICAL ABSTRACT

Downloads
References
HAL Tuppen, EL Blakely, DM Turnbull and RW Taylor. Mitochondrial DNA mutations and human disease. Biochim. Biophys. Acta Bioenerg. 2010; 1797, 113-28.
FR Rahmadanthi and IP Maksum. Transfer RNA mutation associated with type 2 diabetes mellitus. Biology 2023; 12, 871.
MITOMAP, Available at: https://www.mitomap.org/MITOMAP, accessed November 2023.
R Mulyani, W Destiarani, UM Rahmawati, M Yusuf, T Subroto and IP Maksum. An investigation of the effect of T15458C and T15663C mutations in the CYB gene on respiration via an in silico method. J. Proteonomics Bioinf. 2018; 13, 1-8.
S Casagrande, M Dzialo, L Trost, K Malkoc, ET Sadowska, M Hau, B Pierce, S McWilliams and U Bauchinger. Mitochondrial metabolism in blood more reliably predicts whole-animal energy needs compared to other tissues. iScience 2023; 26, 108321.
C Yan, X Duanmu, L Zeng, B Liu and Z Song. Mitochondrial DNA: Distribution, mutations, and elimination. Cells 2019; 8, 379.
IP Maksum, S Sriwidodo, O Suprijana, G Natadisastra, S Nuswantara and AS Noer. Identifikasi Mutasi Heteroplasmi A3243G DNA Mitokondria dan Studi Pewarisan Maternal pada Pasien Diabetes Melitus Tipe 2 (in Indonesian). Bionatura Jurnal Ilmu-ilmu Hayati dan Fisik 2010; 12, 78-85.
AM Ilmi, M Rahmaniar and MI Permana. Design and optimization of PCR-RFLP assay for detection of G9053A and T15663C mutation in mitochondrial DNA. Res. J. Chem. Environ. 2023; 27, 1-5.
A Biggin, R Henke, B Bennetts, DR Thorburn and J Christodoulou. Mutation screening of the mitochondrial genome using denaturing high-performance liquid chromatography. Mol. Genet. Metabol. 2005; 84, 61-74.
ML Smith, XY Hua, DL Marsden, D Liu, NG Kennaway, KY Ngo and RH Haas. Diabetes and mitochondrial encephalomyopathy with lactic acidosis and stroke-like episodes (MELAS): Radiolabeled polymerase chain reaction is necessary for accurate detection of low percentages of mutation. J. Clin. Endocrinol. Metabol. 1997; 82, 2826-31.
HE White, VJ Durston, A Seller, C Fratter, JF Harvey and NCP Cross. Accurate detection and quantitation of heteroplasmic mitochondrial point mutations by pyrosequencing. Genet. Test. 2005; 9, 190-9.
M Urata, M Wakiyama, M Iwase, M Yoneda, S Kinoshita, N Hamasaki and D Kang. New sensitive method for the detection of the A3243G mutation of human mitochondrial deoxyribonucleic acid in diabetes mellitus patients by ligation-mediated polymerase chain reaction. Clin. Chem. 1998; 44, 2088-93.
YW Hartati, SN Topkaya, IP Maksum and M Ozsoz. Sensitive detection of mitochondrial DNA A3243G tRNALeu mutation via an electrochemical biosensor using meldola’s blue as a hybridization indicator. Adv. Anal. Chem. 2013; 3, 20-7.
RAI Chandra, S Sriwidodo, A Diantini and IP Maksum. Restriction enzymes apai analysis to find A3243G mutation in indonesia diabetes mellitus type II patients. J. Med. Bioeng. 2015; 4, 492-6.
S Sriwidodo, O Suprijana, T Subroto and IP Maksum. Studi mutasi titik A3243G DNA mitokondria penyebab maternally (in Indonesian). Majalah Ilmu Kefarmasian 2008; 3, 121-9.
IP Maksum, A Farhani, SD Rachman and Y Ngili. Making of the A3243G mutant template through site directed mutagenesis as positive control in PASA-Mismatch three bases. Int. J. Pharmtech Res. 2013; 5, 441-50.
E Rong, H Wang, S Hao, Y Fu, Y Ma and T Wang. Heteroplasmy detection of mitochondrial DNA A3243G mutation using quantitative real-time PCR assay based on TaqMan-MGB probes. Biomed. Res. Int. 2018; 2018, 1286480.
IP Maksum, AF Maulana, M Yusuf, R Mulyani, W Destiarani and R Rustaman. Molecular dynamics simulation of a tRNA-leucine dimer with an A3243G heteroplasmy mutation in human mitochondria using a secondary structure prediction approach. Ind. J. Chem. 2022; 22, 1043-51.
W Destiarani, R Mulyani, M Yusuf and IP Maksum. Molecular dynamics simulation of T10609C and C10676G mutations of mitochondrial ND4L gene associated with proton translocation in type 2 diabetes mellitus and cataract patients. Bioinf. Biol. Insights 2020; 14, 1-8.
IP Maksum, R Mulyani, K Hasan, MI Azizah, W Destiarani, AF Maulana, M Yusuf and T Subroto. Study on the mitochondrial genome of variants carrying mt.3243A>G from type-2 diabetes mellitus and cataract patients in Indonesia. Hayati 2023; 30, 1017-24.
IP Maksum, S Safitri, M Yusuf, SN Nuswantara and O Suprijana. Structure and function of CYB in type-2 diabetes mellitus and cataract patients which associated with mitochondrial DNA mutation. In: Proceeding of the 1st International Conference on Computation for Science and Technology, Chiang Mai, Thailand. 2010, p. 103-10.
HI Ingólfsson, C Arnarez, X Periole and SJ Marrink. Computational ‘microscopy’ of cellular membranes. J. Cell Sci. 2016; 129, 257-68.
EL Wu, WX Cheng, S Jo, H Rui, KC Song, EM Davilia-Contreras, Y Qi, J Lee, V Monje-Galvan, RM Venable, JB Klauda and W Im. CHARMM-GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 2014; 35, 1997-2004.
S Jo, T Kim, VG Iyer and W Im. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008; 29, 1859-65.
EM Mejia and GM Hatch. Mitochondrial phospholipids: Role in mitochondrial function. J. Bioenerg. Biomembr. 2016; 48, 99-112.
M Kondrashova, M Zakharchenko and N Khunderyakova. Preservation of the in vivo state of mitochondrial network for ex vivo physiological study of mitochondria. Int. J. Biochem. Cell Biol. 2009; 41, 2036-50.
X Meng, H Liu, N Zhao, Y Yang, K Zhao and Y Dai. Molecular dynamics study of the effect of charge and glycosyl on superoxide anion distribution near lipid membrane. Int. J. Mol. Sci. 2023; 24, 10926.
AF Maulana, IP Maksum, S Sriwidodo and Y Rukayadi. Proposed molecular mechanism of non-competitive inhibition using molecular dynamics simulations between α-glucosidase enzyme and mangostin compound as antidiabetic. J. Mol. Model. 2024; 30, 136.
G Pintilie, DH Chen, CA Haase-Pettingell, JA King and W Chiu. Resolution and probabilistic models of components in CryoEM maps of mature P22 bacteriophage. Biophys. J. 2016; 110, 827-39.
RN Beni, P Elyasi-Ebli, S Gharaghani and A Seyedarabi. In silico studies of anti-oxidative and hot temperament-based phytochemicals as natural inhibitors of SARS-CoV-2 Mpro. PLoS One 2023; 18, e0295014.
ASMJ Islam, MdS Akbar, MdS Islam, C Stampfl, AG Bhuiyan and J Park. Atomistic details of grain, crack, and notch effect on the mechanical behavior and fracture mechanisms of monolayer silicon carbide. Mater. Chem. Phys. 2024; 316, 129068.
X Zhang, Y Zhang, T Fan, Z Feng and L Yang. Structure-guided engineered urethanase from Candida parapsilosis with pH and ethanol tolerance to efficiently degrade ethyl carbamate in Chinese rice wine. Ecotoxicol. Environ. Saf. 2024; 276, 116335.
J Sharma, VK Bhardwaj, R Singh, V Rajendran, R Purohit and S Kumar. An in-silico evaluation of different bioactive molecules of tea for their inhibition potency against non-structural protein-15 of SARS-CoV-2. Food Chem. 2021; 346, 128933.
K Kavitha and A Mohanapriya. Insights into the structure–function relationship of missense mutations in the human TOP2A protein in ovarian cancer. Front. Phys. 2024; 12, 01-17.
M Shojapour, F Fatemi, S Farahmand and MD Shasaltaneh. Investigation of Cyc1 protein structure stability after H53I mutation using computational approaches to improve redox potential. J. Mol. Graph. Model. 2021; 105, 107864.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2024 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






