Computational Prediction of Cinnamomum zeylanicum Bioactive Compounds as Potential Antifungal by Inhibit Biofilm Formation of Candida albicans
DOI:
https://doi.org/10.48048/tis.2024.7986Keywords:
Agglutinin-like sequence, Antifungal, Biofilm formation, Molecular docking, Molecular dynamicsAbstract
The issue of Candida albicans resistance to antifungal agents in the management of candidiasis has emerged as a significant concern, necessitating the development of novel antifungal alternatives to ensure efficacy in the treatment of candidiasis. A notable manifestation of C. albicans pathogenicity is the development of biofilms, which are facilitated by the presence of certain genes known as agglutinin-like sequence (ALS) genes, specifically ALS1 and ALS3. The utilisation of Cinnamomum zeylanicum extract as a potential alternative antifungal agent has promise in addressing fungal resistance, particularly in the context of C. albicans, for the treatment of candidiasis. Bioactive compounds such as cinnamonaldehyde, cyclopentane, eugenic acid, hexadecenoid acid and Pyrantel Hydrochloride are present in C. zeylanicum. This research investigates the mechanistic plausibility of the bioactive chemicals found in C. zeylanicum as agents for combating candidiasis. The study specifically examines the inhibitory effects of these compounds on the Agglutinin-like sequence-1 (ALS1) and ALS3 proteins. The study employs a combination of molecular docking and molecular dynamics simulations to elucidate the complex interactions between the bioactive constituents of C. zeylanicum and ALS1/ALS3 proteins. The ligand exhibited notable binding affinity towards the molecules Cinnamaldehyde, Pyrantel Hydrochloride and Hexadecenoic Acid derived from C. zeylanicum, as seen by their binding affinity values ranging from –4.873 to –6.03 kcal/mol. This indicates their potential effectiveness in altering the activities of the target proteins. The investigation reveals distinct interaction patterns between each chemical and the proteases. Significantly, Pyrantel Hydrochloride establishes binding interactions with essential catalytic residues in both ALS1 and ALS3. The stability of the Pyrantel Hydrochloride-ALS1/ALS3 complexes is further supported by molecular dynamics simulations, which suggest that these complexes possess a strong inhibitory capacity. The preservation of protein structures appears to be minimally impacted, indicating a prolonged inhibitory effect. The effectiveness of Pyrantel Hydrochloride is emphasized by the numerous hydrogen bonds that are formed and confirmed using free-binding energy calculations. Collectively, the results highlight the plausibility of C. zeylanicum anti-candidiasis action, principally mediated through the inhibition of ALS1 and ALS3. This activity is predominantly facilitated by Pyrantel Hydrochloride. This comprehensive understanding provides a valuable foundation for the development of innovative antifungal strategies in the battle against C. albicans infections.
HIGHLIGHTS
- The preservation of the protein structures appears to be mostly unaffected, indicating a prolonged inhibitory effect
- The effectiveness of Pyrantel Hydrochloride is emphasized by the numerous hydrogen bonds that are formed and confirmed using free-binding energy calculations
- The collective findings emphasize the plausibility of Zeylanicum’s anti-candidiasis activity, principally achieved by the inhibition of ALS1 and ALS3
- The comprehensive understanding provides a valuable foundation for the development of innovative antifungal strategies in the battle against C. albicans infections
GRAPHICAL ABSTRACT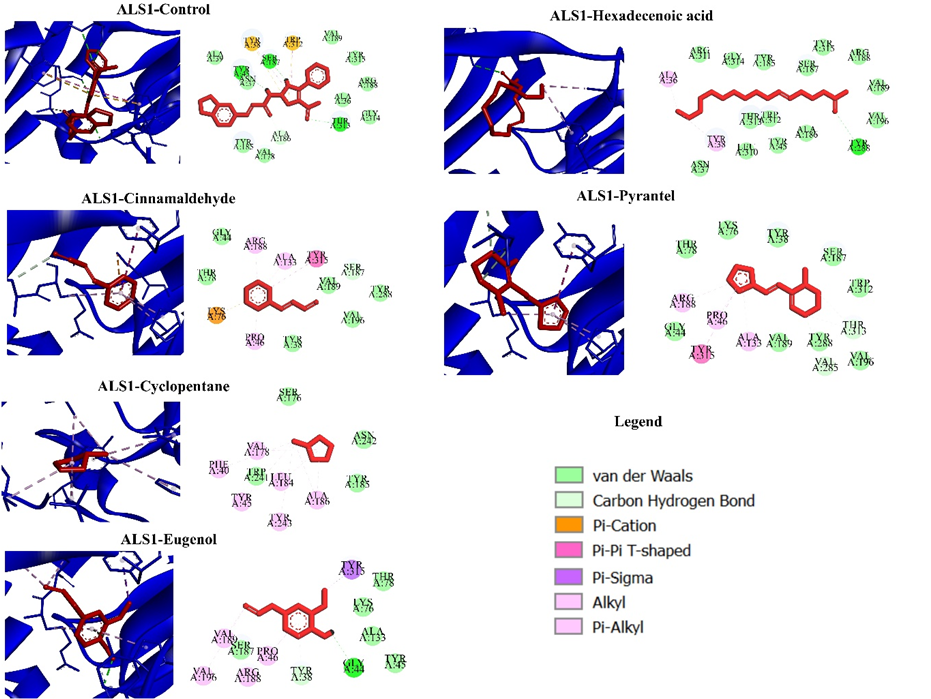
Downloads
References
S Saravani, T Nosratzeh, H Kadeh and S Mir. Oral manifestations and related factors of HIV positive patients in south-east of Iran. J. Dent. Mater. Tech. 2017; 6, 11-8.
V Meylani, L Sembiring, A Fudholi and T Wibawa. Differentiated sap (4 - 6) gene expression of Candida albicans isolates from HIV-positive patients with oral candidiasis and commensals in healthy individuals. Microb. Pathog. 2021; 158, 105075.
C Firacative. Invasive fungal disease in humans: Are we aware of the real impact? Memorias Do Instituto Oswaldo Cruz 2020; 115, e200430.
HE Hallen-Adams and MJ Suhr. Fungi in the healthy human gastrointestinal tract. Virulence 2017; 8, 352-8.
MY Wani, A Ahmad, FM Aqlan and AS Al-Bogami. Citral derivative activates cell cycle arrest and apoptosis signaling pathways in Candida albicans by generating oxidative stress. Bioorg. Chem. 2021; 115, 105260.
DA Hogan and R Kolter. Pseudomonas-Candida interactions: An ecological role for virulence factors. Science 2002; 296, 2229-32.
CSY Lim, R Rosli, HF Seow and PP Chong. Candida and invasive candidiasis: Back to basics. Eur. J. Clin. Microbiol. Infect. Dis. 2012; 31, 21-31.
PE Sudbery. Growth of Candida albicans hyphae. Nat. Rev. Microbiol. 2011; 9, 737-48.
J Liu, HME Willems, EA Sansevere, S Allert, KS Barker, DJ Lowes, AC Dixson, Z Xu, J Miao, C DeJarnette, H Tournu, GE Palmer, JP Richardson, FN Barrera, B Hube, JR Naglik and BM Peters. A variant ECE1 allele contributes to reduced pathogenicity of Candida albicans during vulvovaginal candidiasis. PLoS Pathog. 2021; 17, e1009884.
AR Pereira, AGDO Paranhos, SFD Aquino and SDQ Silva. Distribution of genetic elements associated with antibiotic resistance in treated and untreated animal husbandry waste and wastewater. Environ. Sci. Pollut. Res. 2021; 28, 26380-403.
FL Mayer, D Wilson and B Hube. Candida albicans pathogenicity mechanisms. Virulence 2013; 4, 119-28.
M Henriques and S Silva. Candida albicans virulence factors and its pathogenicity. Microorganisms 2021; 9, 704.
MC Garcia, JT Lee, CB Ramsook, D Alsteens, YF Dufrêne and PN Lipke. A role for amyloid in cell aggregation and biofilm formation. PLoS One 2011; 6, e17632.
KJ Verstrepen and FM Klis. Flocculation, adhesion and biofilm formation in yeasts. Mol. Microbiol. 2006; 60, 5-15.
SS Hosseini, E Ghaemi, A Noroozi and F Niknejad. Zinc oxide nanoparticles inhibition of initial adhesion and ALS1 and ALS3 gene expression in Candida albicans strains from urinary tract infections. Mycopathologia 2019; 184, 261-71.
L Esteban-Tejeda, C Prado, B Cabal, J Sanz, R Torrecillas and JS Moya. Antibacterial and antifungal activity of ZnO containing glasses. PLoS One 2015; 10, e0132709.
H Nails, S Kucharíková, M Ricicová, PV Dijck, D Deforce, H Nelis and T Coenye. Real-time PCR expression profiling of genes encoding potential virulence factors in Candida albicans biofilms: Identification of model-dependent and - independent gene expression. BMC Microbiol. 2010; 10, 114.
A Rabes, S Zimmermann, K Reppe, R Lang, PH Seeberger, N Suttorp, M Witzenrath, B Lepenies and B Opitz. The C-type lectin receptor mincle binds to streptococcus pneumoniae but plays a limited role in the anti-pneumococcal innate immune response. PLoS One 2015; 10, e0117022.
T Sultan and SA Ali. Psoralea corylifolia extracts stimulate cholinergic-like psoralen receptors of tadpole-tail melanophores, leading to skin darkening. J. Recept. Signal Transduction Res. 2011; 31, 39-44.
SL Kelly, DC Lamb, DE Kelly, NJ Manning, J Loeffler, H Hebart, U Schumacher and H Einsele. Resistance to fluconazole and cross-resistance to amphotericin B in Candida albicans from AIDS patients caused by defective sterol delta 5,6-desaturation. FEBS Lett. 1997; 400, 80-2.
D Oro, A Heissler, EM Rossi, D Scapin, PDS Malheiros and E Boff. Antifungal activity of natural compounds against Candida species isolated from HIV-positive patients. Asian Pac. J. Trop. Biomed. 2015; 5, 781-4.
B Khanzada, N Akhtar, MK Okla, SA Alamri, A Al-Hashimi, MW Baig, S Rubnawaz, H AbdElgawad, AH Hirad, IU Haq and B Mirza. Profiling of antifungal activities and in silico studies of natural polyphenols from some plants. Molecules 2021; 26, 7164.
V Meylani, RR Putra, M Miftahussurur, S Sukardiman, FE Hermanto and A Abdullah. Molecular docking analysis of Cinnamomum zeylanicum phytochemicals against Secreted Aspartyl Proteinase 4 - 6 of Candida albicans as anti-candidiasis oral. Results Chem. 2022; 5, 100721.
BA Behbahani, F Falah, FL Arab, M Vasiee and FT Yazdi. Chemical composition and antioxidant, antimicrobial, and antiproliferative activities of Cinnamomum zeylanicum bark essential oil. Evidence Based Complement Altern. Med. 2020; 29, 5190603.
C Giordani, G Simonetti, D Natsagdorj, G Choijamts, F Ghirga, A Calcaterra, D Quaglio, GD Angelis, C Toniolo and G Pasqua. Antifungal activity of Mongolian medicinal plant extracts. Nat. Prod. Res. 2020; 34, 449-55.
IntechOpen. The world’ s leading publisher of Open Access books built by scientists, for scientists TOP 1 %, 2012, p. 13, Available at: https://www.intechopen.com/chapters/17746, accessed December 2023.
N Keikha, M Shafaghat, SM Mousavi, M Moudi and F Keshavarzi. Antifungal effects of ethanolic and aqueous extracts of vitex agnus-castus against vaginal isolates of Candida albicans. Curr. Med. Mycol. 2018; 4, 1-5.
MSA Aboody and S Mickymaray. Anti-fungal efficacy and mechanisms of flavonoids. Antibiotics 2020; 9, 45.
S Rampone, C Pagliarulo, C Marena, A Orsillo, M Iannaccone, C Trionfo, D Sateriale and M Paolucci. In silico analysis of the antimicrobial activity of phytochemicals: Towards a technological breakthrough. Comput. Methods Programs Biomed. 2021; 200, 105820.
OB Rocha, LDC Silva, MABDC Júnior, AAD Oliveira, CMDA Soares and M Pereira. In vitro and in silico analysis reveals antifungal activity and potential targets of curcumin on Paracoccidioides spp. Braz. J. Microbiol. 2021; 52, 1897-911.
Z Shahina, A Molaeitabari, T Sultana and TES Dahms. Cinnamon leaf and clove essential oils are potent inhibitors of Candida albicans virulence traits. Microorganisms 2022; 10, 1989.
S Choonharuangdej, T Srithavaj and S Thummawanit. Fungicidal and inhibitory efficacy of cinnamon and lemongrass essential oils on Candida albicans biofilm established on acrylic resin: An in vitro study. J. Prosthet. Dent. 2021; 125, 707.e1-707.e6.
S Kim, J Chen, T Cheng, A Gindulyte, J He, S He, Q Li, BA Shoemaker, PA Thiessen, B Yu, L Zaslavsky, J Zhang and EE Bolton. PubChem 2023 update. Nucleic Acids Res. 2023; 6, 1373-80.
A Waterhouse, M Bertoni, S Bienert, G Studer, G Tauriello, R Gumienny, FT Heer, TAPD Beer, C Rempfer, L Bordoli, R Lepore and T Schwede. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018; 46, W296-W303.
FE Hermanto, W Warsito, M Rifa’i and N Widodo. Understanding hypocholesterolemic activity of soy isoflavones: Completing the puzzle through computational simulations. J. Biomol. Struct. Dyn. 2022; 41, 9931-7.
NM O’Boyle, M Banck, CA James, C Morley, T Vandermeersch and GR Hutchison. Open Babel: An open chemical toolbox. J. Cheminf. 2011; 3, 33.
S Dallakyan and AJ Olson. Small-molecule library screening by docking with PyRx. In: Chemical biology: Methods and protocols. Humana Press, New York, 2015, p. 243-50.
AK Rappe, CJ Casewir, KS Colwell, WAG III and WM Skiff. UFF, a full periodic table force field for molecular mevhanics and molecular dynamics simulatons. J. Am. Chem. Soc. 1992; 114, 10024-35.
G. Marc, C Araniciu, SD Oniga, L Vlase, A Pîrnău, M Duma, L Măruțescu, MC Chifiriuc and O Oniga. New N-(oxazolylmethyl)-thiazolidinedione active against Candida albicans biofilm: Potential Als proteins inhibitors. Molecules 2018; 23, 2522.
J Eberhardt, D Santos-Martins, AF Tillack and S Forli. AutoDock Vina 1.2.0: New docking methods, expanded force field, and python bindings. J. Chem. Inf. Modell. 2021; 61, 3891-8.
O Trott and AJ Olson. AutoDock Vina: Improving the speed and accurancy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2012; 31, 455-61.
E Krieger and G Vriend. New ways to boost molecular dynamics simulations. J. Comput. Chem. 2015; 36, 996-1007.
JA Maier, C Martinez, K Kasavajhala, L Wickstrom, KE Hauser and C Simmerling. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 2015; 11, 3696-713.
FE Hermanto, W Warsito, M Rifa’i and N Widodo. On the hypolipidemic activity of elicited soybeans: Evidences based on computational analysis. Indones. J. Chem. 2022; 22, 1626-36.
MI Choudhary, M Shaikh, A Tul-Wahab and A Ur-Rahman. In silico identification of potential inhibitors of key SARS-CoV-2 3CL hydrolase (Mpro) via molecular docking, MMGBSA predictive binding energy calculations, and molecular dynamics simulation. PLoS One 2020; 15, e0235030.
R Patil, S Das, A Stanley, L Yadav, A Sudhakar and AK Varma. Optimized hydrophobic interactions and hydrogen bonding at the target-ligand interface leads the pathways of drug-designing. PLoS One 2010; 5, e12029.
D Chen, N Oezguen, P Urvil, C Ferguson, SM Dann and TC Savidge. Regulation of protein-ligand binding affinity by hydrogen bond pairing. Sci. Adv. 2016; 2, e1501240.
E Barratt, RJ Bingham, DJ Warner, CA Laughton, SEV Phillips and SW Homans. Van der Waals interactions dominate ligand-protein association in a protein binding site occluded from solvent water. J. Am. Chem. Soc. 2005; 127, 11827-34.
LL Hoyer and E Cota. Candida albicans agglutinin-like sequence (Als) family vignettes: A review of als protein structure and function. Front. Microbiol. 2016; 7, 280.
HN Otoo, KG Lee, W Qiu and PN Lipke. Candida albicans Als adhesins have conserved amyloid-forming sequences. Eukaryotic Cell 2008; 7, 776-82.
PN Lipke. What we do not know about fungal cell adhesion molecules. J. Fungi 2018; 4, 59.
SH Oh and LL Hoyer. Assessing Als3 peptide-binding cavity and amyloid-forming region contributions to Candida albicans invasion of human oropharyngeal epithelial cells. Front. Cell. Infect. Microbiol. 2022; 12, 890839.
QT Phan, CL Myers, Y Fu, DC Sheppard, MR Yeaman, WH Welch, AS Ibrahim, JEJ Edwards and SG Filler. Als3 is a Candida albicans invasin that binds to cadherins and induces endocytosis by host cells. PloS Biol. 2007; 5, e64.
A Susilo, M Cahyati, Nurjannah, D Pranowo, FE Hermanto and EP Primandasari. Chrysin inhibits indonesian serotype foot-and-mouth-disease virus replication: Insights from DFT, molecular docking, and molecular dynamics analyses. J. Trop. Biodiversity Biotechnol. 2024; 9, jtbb.83140.
S Roudbarmohammadi, M Roudbary, B Bakhshi, F Katiraee, R Mohammadi and M Falahati. ALS1 and ALS3 gene expression and biofilm formation in Candida albicans isolated from vulvovaginal candidiasis. Adv. Biomed. Res. 2016; 5, 105.
K Deng, W Jiang, Y Jiang, Q Deng, J Cao, W Yang and X Zhao. ALS3 expression as an indicator for Candida albicans biofilm formation and drug resistance. Front. Microbiol. 2021; 12, 655242.
EW Bachtiar, S Dewiyani, SMS Akbar and BM Bachtiar. Inhibition of Candida albicans biofilm development by unencapsulated Enterococcus faecalis cps2. J. Dent. Sci. 2016; 11, 323-30.
D Darmadi, D Lindarto, J Siregar, T Widyawati, M Rusda, MM Amin, F Yusuf, PC Eyanoer, M Lubis and I Rey. Study of the molecular dynamics stability in the inhibitory interaction of tenofovir disoproxil fumarate against CTLA-4 in chronic hepatitis B patients. Med. Arch. 2023; 77, 227-30.
JS Finkel and AP Mitchell. Genetic control of Candida albicans biofilm development. Nat. Rev. Microbiol. 2011; 9, 109-18.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2024 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






