miR-143-3p: A Critical Regulator of KRAS and HRAS with Potential for Targeted Therapies in Breast Cancer
DOI:
https://doi.org/10.48048/tis.2025.9630Keywords:
Breast cancer, Gene regulation, HRAS, KRAS, MicroRNA, miR-143-3p, RAS mutationAbstract
Breast cancer is a heterogeneous disease with distinct subtypes, including the highly aggressive triple-negative breast cancer (TNBC), which lacks estrogen, progesterone, and HER2 receptors. The rapid recurrence, high metastatic potential, and poor clinical outcomes of breast cancer are driven by dysregulated signaling pathways, such as PI3K/AKT/mTOR, MAPK/ERK, and JAK/STAT, which contribute to breast cancer progression, therapy resistance, and metastasis. Among these, the Ras family of proteins, particularly KRAS and HRAS, is pivotal in promoting tumor survival, epithelial-to-mesenchymal transition (EMT), and chemotherapy resistance. Recent research highlights the regulatory role of microRNAs (miRNAs) in these pathways, with miR-143-3p emerging as a critical modulator of the RAS signaling cascade. miR-143-3p acts as a tumor suppressor by targeting key components of the RAS pathway, including KRAS, thereby influencing downstream effectors. This regulation impacts cellular processes such as proliferation, apoptosis, and EMT, crucial in breast cancer progression and therapeutic resistance. This review focuses on the role of miR-143-3p in breast cancer, particularly its regulation of the RAS pathway. It discusses its potential as a therapeutic target for improving outcomes in aggressive breast cancer subtypes.
HIGHLIGHTS
- This article reviewed how miR-143-3p targets KRAS and HRAS and influences breast cancer cell proliferation, apoptosis, and migration and development of miR-143-3p carrier agent.
- miR-143-3p targets KRAS and HRAS both directly and indirectly.
- Several miR-143-3p carriers, organic and inorganic, have been developed to increase their stability, bioavailability, and efficiency in targeting cancer cells.
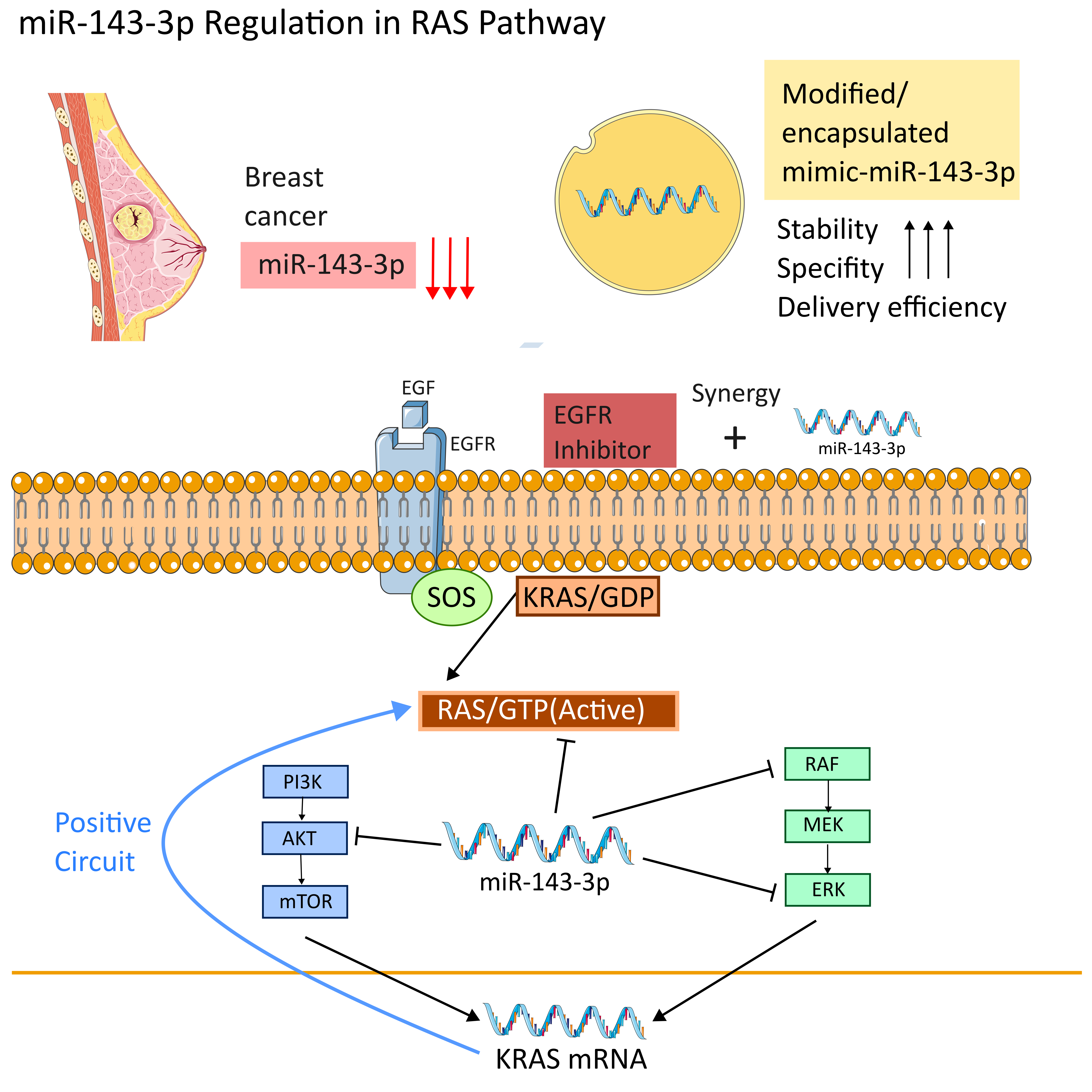
GRAPHICAL ABSTRACT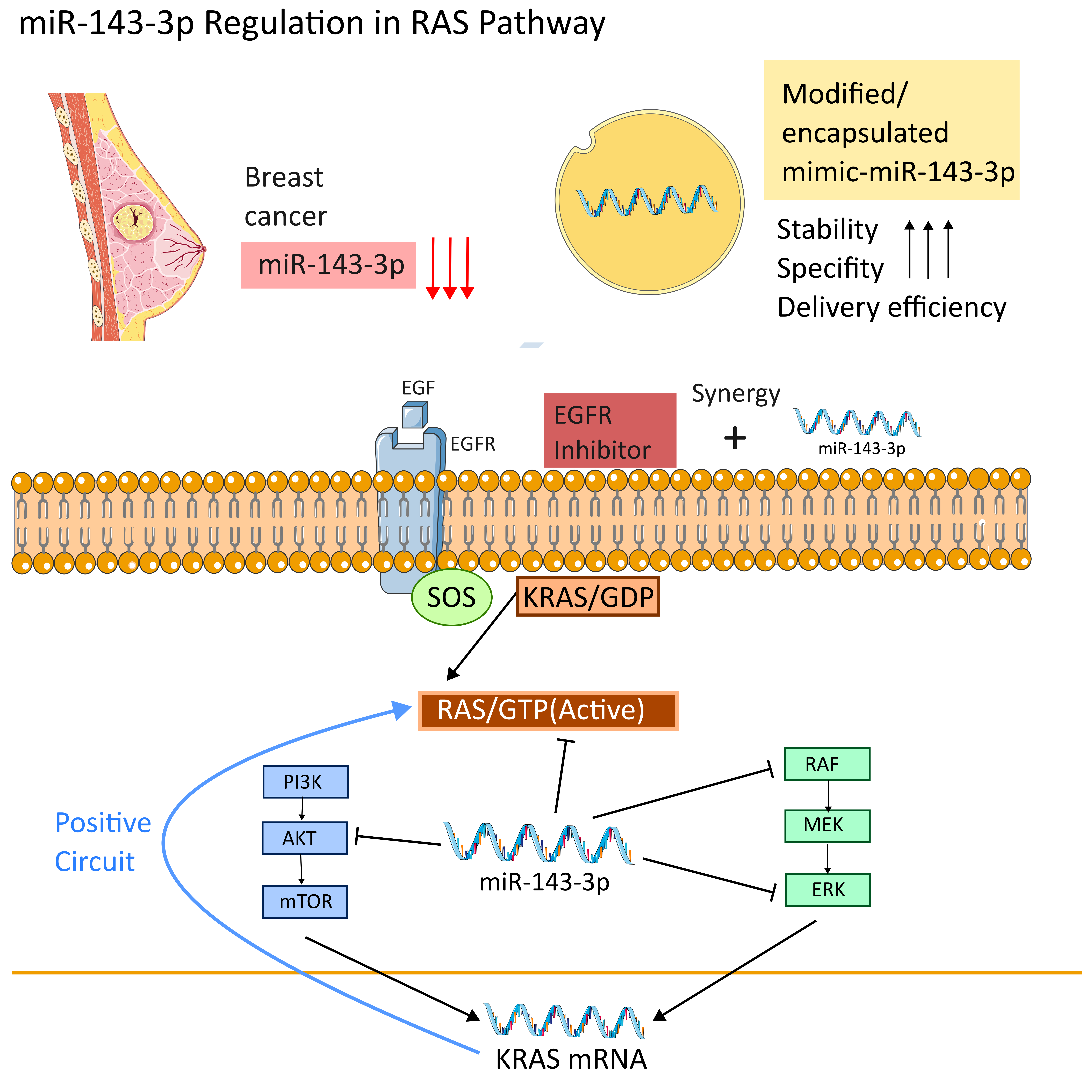
Downloads
References
MA Medina, G Oza, A Sharma, LG Arriaga, JMH Hernandez, VM Rotello and JT Ramirez. Triple-negative breast cancer: A review of conventional and advanced therapeutic strategies. International Journal of Environmental Research and Public Health 2020; 17(6), 2078.
I Muneer, MTU Qamar, K Tusleem, SA Rauf, HMJ Hussain and AR Siddiqi. Discovery of selective inhibitors for cyclic AMP response element-binding protein: A combined ligand and structure-based resources pipeline. Anti-Cancer Drugs 2019; 30(4), 363-373.
Y Lin, H Gao, X Yang, T Zhu, X Zheng, F Ji, L Zhang, C Yang, M Yang, J Li, M Cheng and K Wang. Neoadjuvant therapy in triple-negative breast cancer: A systematic review and network meta-analysis. The Breast 2022; 66, 126-135.
S Singh, A Numan, B Maddiboyina, S Arora, Y Riadi, S Md, NA Alhakamy and P Kesharwani. The emerging role of immune checkpoint inhibitors in the treatment of triple-negative breast cancer. Drug Discovery Today 2021; 26(7), 1721-1727.
O Obidiro, G Battogtokh and EO Akala. Triple negative breast cancer treatment options and limitations: Future outlook. Pharmaceutics 2023; 15(7), 1796.
N Xiong, H Wu and Z Yu. Advancements and challenges in triple-negative breast cancer: A comprehensive review of therapeutic and diagnostic strategies. Frontiers in Oncology 2024; 14, 1405491.
GV Echeverria, Z Ge, S Seth, X Zhang, S Jeter-Jones, X Zhou, S Cai, Y Tu, A McCoy, M Peoples, Y Sun, H Qiu, Q Chang, C Bristow, A Carugo, J Shao, X Ma, A Harris, P Mundi, … H Piwnica-Worms. Resistance to neoadjuvant chemotherapy in triple-negative breast cancer mediated by a reversible drug-tolerant state. Science Translational Medicine 2019; 11(488), eaav0936.
P Hou and YA Wang. Conquering oncogenic KRAS and its bypass mechanisms. Theranostics 2022; 12(13), 5691-5709.
A Ferreira, F Pereira, C Reis, MJ Oliveira, MJ Sousa and A Preto. Crucial role of oncogenic kras mutations in apoptosis and autophagy regulation: Therapeutic implications. Cells 2022; 11(14), 2183.
M Maqbool, F Bekele and G Fekadu. Treatment strategies against triple-negative breast cancer: An updated review. Breast Cancer: Targets and Therapy 2022; 14, 15-24.
BP Kundaktepe, V Sozer, C Papila, S Durmus, PC Kocael, G Simsek, R Gelisgen, K Zengin, K Ulualp and H Uzun. Associations between miRNAs and two different cancers: Breast and colon. Cancer Management and Research 2020; 12, 871-879.
L Wang, Z Shi, C Jiang, X Liu, Q Chen, X Qian, D Li, X Ge, X Wang, L Liu, Y You, N Liu and B Jiang. MiR-143 acts as a tumor suppressor by targeting N-RAS and enhances temozolomide-induced apoptosis in glioma. Oncotarget 2014; 5, 5416-5427.
Y Tokumaru, M Oshi, E Katsuta, L Yan, V Satyananda, N Matsuhashi, M Futamura, Y Akao, K Yoshida and K Takabe. KRAS signaling enriched triple negative breast cancer is associated with favorable tumor immune microenvironment and better survival. American Journal of Cancer Research 2020; 10(3), 897-907.
J Wu, Y Zhu, D Liu, Q Cong and C Bai. Biological functions and potential mechanisms of miR 143 3p in cancers (Review). Oncology Reports 2024; 52(3), 113.
J Xu, X Li, P Zhang, J Luo, E Mou and S Liu. miR-143-5p suppresses breast cancer progression by targeting the HIF-1α-related GLUT1 pathway. Oncology Letters 2022, 23(5), 147.
B Mansoori, S Kiani, AA Mezajin, P Zandi, H Banaie, D Rostamzadeh, WC Cho, PHG Duijf, C Mansoori and B Baradaran. MicroRNA-143-5p Suppresses ER-Positive Breast Cancer Development by Targeting Oncogenic HMGA2. Clinical Breast Cancer 2023; 23(7), e480-e490.
H Sanada, N Seki, K Mizuno, S Misono, A Uchida, Y Yamada, S Moriya, N Kikkawa, K Machida, T Kumamoto, T Suetsugu and H Inoue. Involvement of dual strands of miR-143 (miR-143-5p and miR-143-3p) and their target oncogenes in the molecular pathogenesis of lung adenocarcinoma. International Journal of Molecular Sciences 2019; 20(18), 4482.
V Asghariazar, M Kadkhodayi, M Sarailoo, AG Jolfayi and B Bararadan. MicroRNA-143 as a potential tumor suppressor in cancer: An insight into molecular targets and signaling pathways. Pathology - Research and Practice 2023; 250, 154792.
F Nilasari, SM Haryana, DAA Nugrahaningsih and PB Satriyo. Development of nanocomplex mimic‐hsa‐miR‐143‐3p loaded exosome (exo‐miR) to inhibit viability, migration and proliferation of triple‐negative breast cancer. Indonesian Journal of Biotechnology 2024; 29(4), 190-197.
H Wang, Q Deng, Z Lv, Y Ling, X Hou, Z Chen, S Ma, D Li, Y Wu, Y Peng, H Huang and L Chen. N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1. Molecular Cancer 2019; 18, 181.
H Adhikari, WE Kattan, S Kumar, P Zhou, JF Hancock and CM Counter. Oncogenic KRAS is dependent upon an EFR3A-PI4KA signaling axis for potent tumorigenic activity. Nature Communications 2021; 12, 5248.
B Wang, Y Xu, Y Wei, L Lv, N Liu, R Lin, X Wang and B Shi. Human mesenchymal stem cell-derived exosomal microRNA-143 promotes apoptosis and suppresses cell growth in pancreatic cancer via target gene regulation. Frontiers in Genetics 2021; 12, 581694.
W Kolch, D Berta and E Rosta. Dynamic regulation of RAS and RAS signaling. Biochemical Journal 2023; 480(1), 1-23.
R Nussinov, M Zhang, R Maloney and H Jang. Ras isoform-specific expression, chromatin accessibility, and signaling. Biophysical Reviews 2021, 13(4), 489-505.
GE Menzies, IA Prior, A Brancale, SH Reed and PD Lewis. Carcinogen-induced DNA structural distortion differences in the RAS gene isoforms; The importance of local sequence. BMC Chemistry 2021; 15, 51.
R Szalat, M Lawlor, M Fulciniti, CB Epstein, Y Xu, A Samur, C Lin, P Rao, N Farrell, S Wu, L Schwartz, K Wen, Y Tai, J Wang, N Gray, R Young, KC Anderson, M. Samur, C Ott and N Munshi. Genome wide chromatin accessibility profiling identifies chromatin signatures and novel transcription factor dependencies in multiple myeloma. Clinical Lymphoma, Myeloma and Leukemia 2019; 19(10), E8-E9.
S Zhang, P Li, J Li, J Gao, Q Qi, G Dong, X Liu, Q Jiao, Y Wang, L Du, H Zhan, S Xu and C Wang. Chromatin accessibility uncovers KRAS-driven FOSL2 promoting pancreatic ductal adenocarcinoma progression through up-regulation of CCL28. British Journal of Cancer 2023; 129, 426-443.
C Xia, Y Yang, F Kong, Q Kong and C Shan. MiR-143-3p inhibits the proliferation, cell migration and invasion of human breast cancer cells by modulating the expression of MAPK7. Biochimie 2018; 147, 98-104
R Kim, Y Suh, K Yoo, Y Cui, H Kim, MJ Kim, IG Kim and S Lee. Activation of KRAS promotes the mesenchymal features of basal-type breast cancer. Experimental & Molecular Medicine 2015; 47(1), e137.
M Teufelsbauer, S Stickler, M Eggerstorfer, DC Hammond and G Hamilton. BET-directed PROTACs in triple negative breast cancer cell lines MDA-MB-231 and MDA-MB-436. Breast Cancer Research and Treatment 2024; 208(1), 89-101.
M Koh, H Yong, E Kim, H Son, YR Jeon, J Hwang, M Kim, Y Cha, WS Choi, D Noh, K Lee, K Kim, J Lee, HJ Kim, H Kim, H Kim, EJ Kim, SY Park, HS Kim, … A Moon. A novel role for flotillin-1 in H-Ras-regulated breast cancer aggressiveness. International Journal of Cancer 2016; 138(5), 1232-1245.
N Barnabas and D Cohen. Phenotypic and molecular characterization of MCF10DCIS and SUM breast cancer cell lines. International Journal of Breast Cancer 2013; 2013, 872743.
KH Lee, M Koh and A Moon. Farnesyl transferase inhibitor FTI-277 inhibits breast cell invasion and migration by blocking H-Ras activation. Oncology Letters 2016; 12(3), 2222-2226.
M Dai, G Yan, N Wang, G Daliah, AM Edick, S Poulet, J Boudreault, S Ali, SA Burgos and JJ Lebrun. In vivo genome-wide CRISPR screen reveals breast cancer vulnerabilities and synergistic mTOR/Hippo targeted combination therapy. Nature Communications 2021; 12, 3055.
H Liang, G Zhou, L Lv, J Lu and J Peng. KRAS expression is a prognostic indicator and associated with immune infiltration in breast cancer. Breast Cancer 2021; 28(2), 379-386.
MB Myers, M Banda, KL McKim, Y Wang, MJ Powell and BL Parsons. Breast cancer heterogeneity examined by high-sensitivity quantification of PIK3CA, KRAS, HRAS, and BRAF mutations in normal breast and ductal carcinomas. Neoplasia 2016; 18(4), 253-263.
M Banys-Paluchowski, K Milde-Langosch, T Fehm, I Witzel, L Oliveira-Ferrer, B Schmalfeldt and V Muller. Clinical relevance of H-RAS, K-RAS, and N-RAS mRNA expression in primary breast cancer patients. Breast Cancer Research and Treatment 2020; 179(2), 403-414.
X Sun, G Dai, L Yu, Q Hu, J Chen and W Guo. miR-143-3p inhibits proliferation, migration and invasion in osteosarcoma by targeting FOSL2. Scientific Reports 2018; 8, 606.
J O’Brien, H Hayder, Y Zayed and C Peng. Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Frontiers in Endocrinology 2018; 9, 402.
B Smolarz, A Durczyński, H Romanowicz, K Szyllo and P Hogendorf. miRNAs in cancer (review of literature). International Journal of Molecular Sciences 2022; 23(5), 2805.
A Menon, N Abd-Aziz, K Khalid, CL Poh and R Naidu. miRNA: A promising therapeutic target in cancer. International Journal of Molecular Sciences 2022; 23(19), 11502.
SK Prajapati, N kumari, D Bhowmik and R Gupta. Recent advancements in biomarkers, therapeutics, and associated challenges in acute myeloid leukemia. Annals of Hematology 2024; 103(11), 4375-4400.
E Jordan-Alejandre, AD Campos-Parra, DL Castro-Lopez and MB Silva-Cazares. Potential miRNA Use as a biomarker: From breast cancer diagnosis to metastasis. Cells 2023; 12(4), 525.
L Karimi, B Mansoori, D Shanebandi, A Mohammadi, M Aghapour and B Baradaran. Function of microRNA-143 in different signal pathways in cancer: New insights into cancer therapy. Biomedicine & Pharmacotherapy 2017; 91, 121-131.
T Lu, T Qiu, B Han, Y Wang, X Sun, Y Qin, A Liu, N Ge and W Jiao. Circular RNA circCSNK1G3 induces HOXA10 signaling and promotes the growth and metastasis of lung adenocarcinoma cells through hsa‑miR‑143‑3p sponging. Cellular Oncology 2021; 44(2), 297-310.
R Chen, C Zhang, Y Cheng, S Wang, H Lin and H Zhang. LncRNA UCC promotes epithelial-mesenchymal transition via the miR-143-3p/SOX5 axis in non-small-cell lung cancer. Laboratory Investigation 2021; 101, 1153-1165.
Y Yang, S Li, J Cao, Y Li, H Hu and Z Wu. RRM2 regulated by LINC00667/miR-143-3p signal is responsible for non-small cell lung cancer cell progression. OncoTargets and Therapy 2019; 12, 9927-9939.
Q Li, Y Bian and Q Li. Down-regulation of TMPO-AS1 induces apoptosis in lung carcinoma cells by regulating miR-143-3p/CDK1 Axis. Technology in Cancer Research & Treatment 2021; 20, 1-10.
C Huang, C Tsai, C Yu, Y Wu, MF Ye, J Ho and D Yu. Long noncoding RNA LINC02470 sponges microRNA-143-3p and enhances SMAD3-Mediated epithelial-to-mesenchymal transition to promote the aggressive properties of bladder cancer. Cancers 2022, 14(4), 968.
L Dong, X Zhang, M Mao, Y Li, X Zhang, D Xue and Y Liu. LINC00511/miRNA-143-3p modulates apoptosis and malignant phenotype of bladder carcinoma cells via PCMT1. Frontiers in Cell and Developmental Biology 2021; 9, 650999.
T Xiang, H Jiang, B Zhang and G Liu. CircFOXO3 functions as a molecular sponge for miR-143-3p to promote the progression of gastric carcinoma via upregulating USP44. Gene 2020; 753, 144798.
LL Zhou, JL Dong, G Huang, ZL Sun and J Wu. MicroRNA-143 inhibits cell growth by targeting ERK5 and MAP3K7 in breast cancer. Brazilian Journal of Medical and Biological Research 2017; 50(8), e5891.
J Li, H Zhang and H Luo. Long non-coding RNA OIP5-as1 contributes to gallbladder cancer cell invasion and migration by miR-143-3p suppression. Cancer Management and Research 2020; 12, 12983-12992.
G Zhang, Z Liu, J Zhong and L Lin. Circ-ACAP2 facilitates the progression of colorectal cancer through mediating miR-143-3p/FZD4 axis. European Journal of Clinical Investigation 2021; 51(12), e13607.
J Chen and X Chen. MYBL2 is targeted by miR-143-3p and regulates breast cancer cell proliferation and apoptosis. Oncology Research 2018; 26(6), 913-922.
D Li, J Hu, H Song, H Xu, C Wu, B Zhao, D Xie, T Wu, J Zhao and L Fang. miR-143-3p targeting LIM domain kinase 1 suppresses the progression of triple-negative breast cancer cells. American Journal of Translational Research 2017; 9(5), 2276-2285.
Y Cui, Y Fan, G Zhao, Q Zhang, Y Bao, Y Cui, Z Ye, G Chen, X Piao, F Guo, J Wang, Y Bai and D Yu. Novel lncRNA PSMG3‑AS1 functions as a miR‑143‑3p sponge to increase the proliferation and migration of breast cancer cells. Oncology Reports 2020; 43(1), 229-239.
L Song, L Wang, X Pan and C Yang. lncRNA OIP5-AS1 targets ROCK1 to promote cell proliferation and inhibit cell apoptosis through a mechanism involving miR-143-3p in cervical cancer. Brazilian Journal of Medical and Biological Research 2018; 53(1), e8883.
H Zhao, M Bi, M Lou, X Yang and L Sun. Downregulation of SOX2-OT prevents hepatocellular carcinoma progression through miR-143-3p/MSI2. Frontiers in Oncology 2021; 11, 685912.
L Chen, H Yao, K Wang and X Liu. Long non-coding RNA MALAT1 regulates ZEB1 expression by sponging miR-143-3p and promotes hepatocellular carcinoma progression. Journal of Cellular Biochemistry 2017; 118(12), 4836-4843.
J Peng, HJ Wu, HF Zhang, SQ Fang and R Zeng. miR-143-3p inhibits proliferation and invasion of hepatocellular carcinoma cells by regulating its target gene FGF1. Clinical and Translational Oncology 2021; 23(3), 468-480.
H Fan, Y Ge, X Ma, Z Li, L Shi, L Lin, J Xiao, W Chen, P Ni, L Yang and Z Xu. Long non-coding RNA CCDC144NL-AS1 sponges miR-143-3p and regulates MAP3K7 by acting as a competing endogenous RNA in gastric cancer. Cell Death & Disease 2020; 11(7), 521.
S Wang, W Li, L yang, J Yuan, L Wang, N Li and H Zhao. CircPVT1 facilitates the progression of oral squamous cell carcinoma by regulating miR-143-3p/SLC7A11 axis through MAPK signaling pathway. Functional & Integrative Genomics 2022; 22(5), 891-903.
T Shan, Z Tian, Q Li, Y Jiang, F Liu, X Sun, Y Han, L Sun and L Chen. Long intergenic noncoding RNA 00908 promotes proliferation and inhibits apoptosis of colorectal cancer cells by regulating KLF5 expression. Journal of Cellular Physiology 2021; 236(2), 889-899.
X Ding, J Du, K Mao, X Wang, Y Ding and F Wang. MicroRNA-143-3p suppresses tumorigenesis by targeting catenin-δ1 in colorectal cancer. OncoTargets and Therapy 2019; 12, 3255-3265.
L Guo, J Fu, S Sun, M Zhu, L Zhang, H Niu, Z Chen, Y Zhang, L Guo and S Wang. MicroRNA-143-3p inhibits colorectal cancer metastases by targeting ITGA6 and ASAP3. Cancer Science 2019; 110(2), 805-816.
Liu M, J Jia, X Wang, Y Liu, C Wang and R Fan. Long non-coding RNA HOTAIR promotes cervical cancer progression through regulating BCL2 via targeting miR-143-3p. Cancer Biology & Therapy 2018; 19(5), 391-399.
J Tang, H Pan, W Wang, C Qi, C Gu, A Shang and J Zhu. MiR-495-3p and miR-143-3p co-target CDK1 to inhibit the development of cervical cancer. Clinical and Translational Oncology 2021; 23(11), 2323-2334.
T Liu, X Wang, J Zhai, Q Wang and B Zhang. Long Noncoding RNA UCA1 facilitates endometrial cancer development by regulating KLF5 and RXFP1 gene expressions. Cancer Biotherapy & Radiopharmaceuticals 2021; 36(6), 521-533.
Y Hou, H Feng, J Jiao, L Qian, B Sun, P Chen, Q Li and Z Liang. Mechanism of miR-143-3p inhibiting proliferation, migration and invasion of osteosarcoma cells by targeting MAPK7. Artificial Cells, Nanomedicine, and Biotechnology 2019; 47(1), 2065-2071.
Z Doosti, SO Ebrahimi, MS Ghahfarokhi and S Reiisi. Synergistic effects of miR-143 with miR-99a inhibited cell proliferation and induced apoptosis in breast cancer. Biotechnology and Applied Biochemistry 2024; 71(50), 993-1004
X Chen, J Zhou, L Luo, B Han, F Li, J Chen, Y Zhu, W Chen and X Yu. Black rice anthocyanins suppress metastasis of breast cancer cells by targeting RAS/RAF/MAPK Pathway. BioMed Research International 2015; 2015(1), 414250.
L Armstrong, CE Willoughby and DJ McKenna. Targeting of AKT1 by miR-143-3p suppresses epithelial-to-mesenchymal transition in prostate cancer. Cells 2023; 12(18), 2207.
Y Du, J Zhang, Y Meng, M Huang, W Yan and Z Wu. MicroRNA-143 targets MAPK3 to regulate the proliferation and bone metastasis of human breast cancer cells. AMB Express 2020; 10, 134.
F Tavanafar, R Safaralizadeh, MA Hosseinpour-Feizi, B Mansoori, D Shanehbandi, A Mohammadi and B Baradaran. Restoration of miR-143 expression could inhibit migration and growth of MDA-MB-468 cells through down-regulating the expression of invasion-related factors. Biomedicine & Pharmacotherapy 2017; 91, 920-924.
F Zhang and H Cao. MicroRNA‑143‑3p suppresses cell growth and invasion in laryngeal squamous cell carcinoma via targeting the k‑Ras/Raf/MEK/ERK signaling pathway. International Journal of Oncology 2019; 54(2), 689-701.
JP Munoz, P Perez-Moreno, Y Perez and GM Calaf. The role of microRNAs in breast cancer and the challenges of their clinical application. Diagnostics 2023; 13(19), 3072.
Y Miao, C Fu, Z Yu, L Yu, Y Tang and M Wei. Current status and trends in small nucleic acid drug development: Leading the future. Acta Pharmaceutica Sinica B 2024; 14(9), 3802-3817.
AS Alfutaimani, NK Alharbi, AS Alahmari, AA Alqabbani and AM Aldayel. Exploring the landscape of lipid nanoparticles (LNPs): A comprehensive review of LNPs types and biological sources of lipids. International Journal of Pharmaceutics: X 2024; 8, 100305.
Y Suzuki and H Ishihara. Difference in the lipid nanoparticle technology employed in three approved siRNA (Patisiran) and mRNA (COVID-19 vaccine) drugs. Drug Metabolism and Pharmacokinetics 2021; 41, 100424.
K Chen, Y Zhang, L Qian and P Wang. Emerging strategies to target RAS signaling in human cancer therapy. Journal of Hematology & Oncology 2021; 14(1), 116.
C Wang, Y Zhang, W Kong, X Rong, Z Zhong, L Jiang, S Chen, C Li, F Zhang, and J Jiang. Delivery of miRNAs using nanoparticles for the treatment of osteosarcoma. International Journal of Nanomedicine 2024; 19, 8641-8660.
Y Lu, Z Mai, L Cui and X Zhao. Engineering exosomes and biomaterial-assisted exosomes as therapeutic carriers for bone regeneration. Stem Cell Research & Therapy 2023; 14, 55.
A Zalbalza, A Pappolla, M Comabella, X Montalban and S Malhotra. MiRNA-based therapeutic potential in multiple sclerosis. Frontiers in Immunology 2024; 15, 1441733.
JS Suk, Q Xu, N Kim, J Hanes and LM Ensign. PEGylation as a strategy for improving nanoparticle-based drug and gene delivery. Advanced Drug Delivery Reviews 2016; 99(Part A), 28-51.
P Mishra, B Nayak and RK Dey. PEGylation in anti-cancer therapy: An overview. Asian Journal of Pharmaceutical Sciences 2016; 11(3), 337-348.
N Miyamoto, N Sugito, Y Kitade and Y Akao. Growth inhibition of RAS-mutated hematopoietic tumor cells using glucose-attached reversibly ionic oligonucleotide-based nanoparticles caging chemically modified microRNA143-3p. Journal of Drug Delivery Science and Technology 2023; 88, 104902.
C Chakraborty, AR Sharma, G Sharma and S Lee. Therapeutic advances of miRNAs: A preclinical and clinical update. Journal of Advanced Research 2020; 28, 127-138.
AR Teixeira, VV Ferreira, T Pereira-da-Silva and RC Ferreira. The role of miRNAs in the diagnosis of stable atherosclerosis of different arterial territories: A critical review. Frontiers in Cardiovascular Medicine 2022; 9, 1040971.
I Fernandez-Pineiro, I Badiola and A Sanchez. Nanocarriers for microRNA delivery in cancer medicine. Biotechnology Advances 2017; 35(3), 350-360.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






