OncomiR Structure and Network Prediction on Adenomatosis Polyposis Coli (APC) Gene Silencing Regulation in Colorectal Cancer
DOI:
https://doi.org/10.48048/tis.2023.6168Keywords:
ß-catenin, Cancer cell proliferation, miR-135a/b, miR-494, WntAbstract
The emergence of colorectal cancer cells is associated with the inactivation of the adenomatosis polyposis coli (APC) gene which increases the activity of ß-catenin, one of which is due to oncomiRNA (cancer-inducing microRNA). miR-135a/b-5p and miR-494-3p were thought to be involved in silencing the APC gene and increasing cell proliferation and could potentially be used as anti-miR targets. However, there is a need for an in-depth evaluation of the involvement of the oncomiR as a therapeutic target in preventing the formation of CRC. Therefore, this study aimed to predict the mechanism of inhibition of oncomiR hsa-miR-135a/b, and has-miR-494-3p against APC in the Wnt/ß-catenin signaling pathway. This research was conducted through in silico analysis using a web-based application to describe the stability of the secondary structure, binding position on mRNA, and conserved nucleotides that support biological activity. The data obtained were then used to develop miRNA interaction networks with APCs on the CRC-associated Wnt/ß-catenin signaling pathway. This study suggests that miR-135a-5p, and miR-135b-5p probably evolved earlier in the evolutionary evolution of the conserved oncomiR CRC in various vertebrate species, whereas miR-494-3p is more conserved and commonly found in mammals. The biological activity of miR-494-3p is likely to be more stable and patent to bind to APC gene mRNA and trigger CRC cell proliferation. Furthermore, miR-135a/b-5p and miR-494-3p have the potential to be developed as targets for anti-miR-based transcriptomic therapy as well as for early diagnosis of CRC development. Anti-miR therapy will likely need to involve more than 1 miRNA, as each gene has more than 1 miRNA binding site.
HIGHLIGHTS
Important findings in this study include:
- Adenomatosis polyposis coli (APC) gene silencing in CRC cases correlates to oncomiR activity of miR-135a/b-3p and miR-494-3p
- miR-494 tends to inhibit the translation of APC mRNA more strongly than miR-135a/b-5p represented by lower context++ score in mRNA binding simulation
- In some cases, CRC formation was also caused by silencing activity by oncomiRs, such as miR-135a-5p, miR-135b-5p and miR-494-3p which inhibited APC gene mRNA translation
- This study suggests that miR-135a-5p, and miR-135b-5p likely evolved earlier in the evolution of conserved CRC development in various vertebrate species, whereas miR-494-3p is more conserved and common in mammals
- Based on the biological activity miR-494-3p is likely to be more stable and potent to bind to the mRNA of the APC
- miR-135a/b-5p and miR-494-3p is potentially developed as targets for transcriptomic anti-miR-based therapy as well as for early diagnosis of CRC development
GRAPHICAL ABSTRACT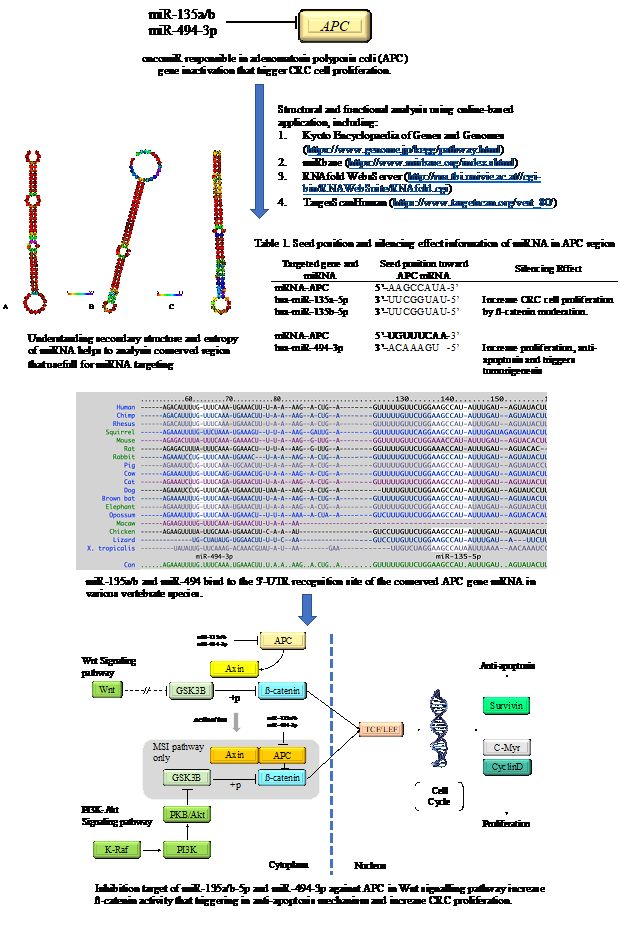
Downloads
References
F Bray, J Ferlay, I Soerjomataram, RL Siegel, LA Torre and A Jemal. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018; 68, 394-424.
H Sung, J Ferlay, RL Siegel, M Laversanne, I Soerjomataram, A Jemal and F Bray. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021; 71, 209-49.
C Langner. Serrated and non-serrated precursor lesions of colorectal cancer. Dig. Dis. 2014; 33, 28-37.
L Novellasdemunt, P Antas and VSW Li. Targeting Wnt signaling in colorectal cancer. A review in the theme: Cell signaling: Proteins, pathways and mechanisms. Am. J. Physiol. Cell Physiol. 2015; 309, 511-21.
N Shirafkan, B Mansoori, A Mohammadi, N Shomali, M Ghasbi and B Baradaran. MicroRNAs as novel biomarkers for colorectal cancer: New outlooks. Biomed. Pharmacother. 2018; 97, 1319-30.
A Saberinia, A Alinezhad, F Jafari, S Soltany and RA Sigari. Oncogenic miRNAs and target therapies in colorectal cancer. Clin. Chim. Acta 2020; 508, 77-91.
AS Aghabozorgi, A Bahreynic, A Soleimani, A Bahrami, M Khazaei, GA Ferns, A Avan and SM Hassanian. Role of adenomatous polyposis coli (APC) gene mutations in the pathogenesis of colorectal cancer; current status and perspectives. Biochimie 2019; 157, 64-71.
R Kariv, M Caspi, N Fliss-Isakov, Y Shorer, Y Shor, G Rosner, E Brazowski, G Beer, S Cohen and R Rosin-Arbesfeld. Resorting the function of the colorectal cancer gatekeeper adenomatous polyposis coli. Int. J. Cancer 2020; 146, 1064-74.
M Zhu, Z Huang, D Zhu, X Zhou, X Shan, LW Qi, L Wu, W Cheng, J Zhu, L Zhang, H Zhang, Y Chen, W Zhu, T Wang and P Liu. A panel of microRNA signature in serum for colorectal cancer diagnosis. Oncotarget 2017; 8, 17081-91.
M Hollis, K Nair, A Vyas, LS Chaturvedi, S Gambhir and D Vyas. MicroRNAs potential utility in colon cancer: Early detection, prognosis, and chemosensitivity. World J. Gastroenterol. 2015; 21, 8284-92.
B Chen, Z Xia, YN Deng, Y Yang, P Zhang, H Zhu, N Xu and S Liang. Emerging microRNA biomarkers for colorectal cancer diagnosis and prognosis. Open Biol. 2019; 9, 180212.
VL Tsikitis, A Potter, M Mori, JA Buckmeier, CR Preece, CA Harrington, AN Bartley, AK Bhattacharyya, SR Hamilton, MP Lance and PA Thompson. MicroRNA signatures of colonic polyps on screening and histology. Cancer Prev. Res. 2016; 9, 942-9.
YF Lu, Y Liu, WM Fu, J Xu, B Wang, YX Sun, TY Wu, LL Xu, KM Chan, JF Zhang and G Li. Long noncoding RNA H19 accelerates tenogenic differentiation & promotes tendon healing through targeting miR-29b-3p & activating TGF-β1 signaling. FASEB J. 2017; 31, 954-64.
H Rezaei, M Motovali-Bashi and S Khalilian. MicroRNA prediction in the FVIII gene locus: A step towards hemophilia a control. Gene Cell Tissue 2020; 7, e103096.
A Kozomara and S Griffiths-Jones. MiRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014; 42, D68-D73.
AR Gruber, R Lorenz, SH Bernhart, R Neuböck and IL Hofacker. The vienna RNA websuite. Nucleic Acids Res. 2008; 36, W70-W74.
M Kertesz, N Iovino, U Unnerstall, U Gaul and E Segal. The role of site accessibility in microRNA target recognition. Nat. Genet. 2007; 39, 1278-84.
DH Mathews, MD Disney, JL Childs, SJ Schroeder, M Zuker and DH Turner. Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proc. Natl. Acad. Sci. USA 2004; 101, 7287-92.
V Agarwal, GW Bell, JW Nam and DP Bartel. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015; 4, e05005.
RC Friedman, KKH Farh, CB Burge and DP Bartel. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009; 19, 92-105.
SE McGeary, KS Lin, CY Shi, TM Pham, N Bisaria and DP Bartel. The biochemical basis of microRNA targeting efficacy. Science 2018; 366, eaav1741.
TC Dakal. Identification of BAP1-associated MicroRNAs and implications in cancer development. Int. J. Cancer Sci. Ther. 2019; 1, 1-4.
X Fu, W Zhu, L Cai, B Liao, L Peng, Y Chen and J Yang. Improved pre-miRNAs identification through mutual information of pre-miRNA sequences and structures. Front. Genet. 2019; 10, 119.
SC Kwon, C Baek, YG Choi, J Yang, YS Lee, JS Woo and VN Kim. Molecular basis for the single-nucleotide precision of primary microRNA processing. Mol. Cell 2019; 73, 505-18.
W Lan, Q Chen, T Li, C Yuan, S Mann and B Chen. Identification of important positions within miRNAs by integrating sequential and structural features. Curr. Protein Pept. Sci. 2014; 15, 591-7.
F Moosavi, E Giovannetti, L Saso and O Firuzi. HGF/MET pathway aberrations as diagnostic, prognostic, and predictive biomarkers in human cancers. Crit. Rev. Clin. Lab. Sci. 2019; 56, 533-66.
YS Jung and JI Park. Wnt signaling in cancer: Therapeutic targeting of Wnt signaling beyond β-catenin and the destruction complex. Exp. Mol. Med. 2020; 52, 183-91.
GT Chen, DF Tifrea, R Murad, AN Habowski, Y Lyou, MR Duong, L Hosohama, A Mortazavi, RA Edwards and ML Waterman. Disrupting ß-catenin dependent Wnt signaling activates an invasive gene program predictive of colon cancer progression. Mol. Cancer Res. 2022; 20, 468-84.
A Aitchison, C Hakkaart, RC Day, HR Morrin, FA Frizelle and JI Keenan. APC mutations are not confined to hotspot regions in early-onset colorectal cancer. Cancers 2020; 12, 3829.
L Mishra. STRAP: A bridge between mutant APC and Wnt/ß-Catenin signaling in intestinal cancer. Gastroenterology 2022; 162, 44-6.
L Magalhães, LG Quintana, DCF Lopes, AF Vidal, AL Pereira, LCDA Pinto, JJV Pinheiro, AS Khayat, LR Goulart, R Burbano, PPD Assumpção and A Ribeiro-Dos-Santos. APC gene is modulated by hsa-miR-135b-5p in both diffuse and intestinal gastric cancer subtypes. BMC Cancer 2018; 18, 1055.
A Wezel, SMJ Welten, W Razawy, HM Lagraauw, MRD Vries, EAC Goossens, MC Boonstra, JF Hamming, ER Kandimalla, J Kuiper, PHA Quax, AY Nossent and I Bot. Inhibition of MicroRNA-494 reduces carotid artery atherosclerotic lesion development and increases plaque stability. Ann. Surg. 2015; 262, 841-7.

Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2022 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






