Characterization of Staphylococcus aureus from Bovine Mastitis: Genetic Markers and Antibiotic Resistance
DOI:
https://doi.org/10.48048/tis.2025.9505Keywords:
Bovine milk, Subclinical mastitis, Antibiotic resistance, Molecular characterization, Beta-lactamaseAbstract
Staphylococcus aureus is a significant contributor to subclinical bovine mastitis, causing economic losses in the dairy sector. This study aimed to isolate and characterize S. aureus from 51 subclinical bovine milk samples. Bacterial colonies were cultured on Mannitol Salt Agar (MSA) using ten-fold serial dilutions, and isolates were identified based on catalase and coagulase tests. Antibiotic resistance screening was conducted on MSA supplemented with 16 µg/mL penicillin G. Molecular characterization employed PCR to detect the nuc, coa, clfa, mecA, blaZ, and se genes. Antibiotic susceptibility testing followed the disc diffusion method, and penicillin G minimum inhibitory concentrations (MICs) were determined using broth microdilution. Sequence and phylogenetic analyses of the blaZ gene were performed to investigate genetic variations and their association with resistance. Results showed that 14 isolates harbored resistance genes from milk samples with high somatic cell counts, though the se gene was absent. All isolates demonstrated 100 % susceptibility to gentamicin, streptomycin, rifampin, ampicillin, and chloramphenicol, while exhibiting complete resistance to penicillin, cefotetan, and nalidixic acid. The minimum inhibitory concentration (MIC) of penicillin G varied widely, ranging from 16 to 512 µg/mL. Phylogenetic analysis identified three distinct groups; however, no significant correlation was found between blaZ variations and resistance levels. These findings emphasizing the importance of robust surveillance and responsible antibiotic use to control bovine mastitis in Thailand.
HIGHLIGHTS
- Only 293 colonies (6.87 %) were identified as S. aureus, with 14 penicillin-resistant S. aureus were able to grow on MSA supplemented with 16 µg/mL penicillin G, highlighting regional differences and antibiotic resistance concerns.
- Five distinct patterns were identified with complete resistance to penicillin, cefotetan, and nalidixic acid, emphasizing the challenge of treating bovine mastitis caused by these isolates.
- All the isolates possessed the nuc and coa genes, with subsets carrying the clfa and mecA genes, suggesting varying pathogenicity and methicillin resistance.
- Minimum inhibitory concentration (MIC) for penicillin ranged from 16 to 512 µg/mL, with widespread presence of the blaZ gene supporting β-lactamase-mediated resistance.
- Phylogenetic analysis revealed 3 distinct clusters of blaZ gene sequences, offering new insights into genetic variability and its limited correlation with phenotypic resistance levels.
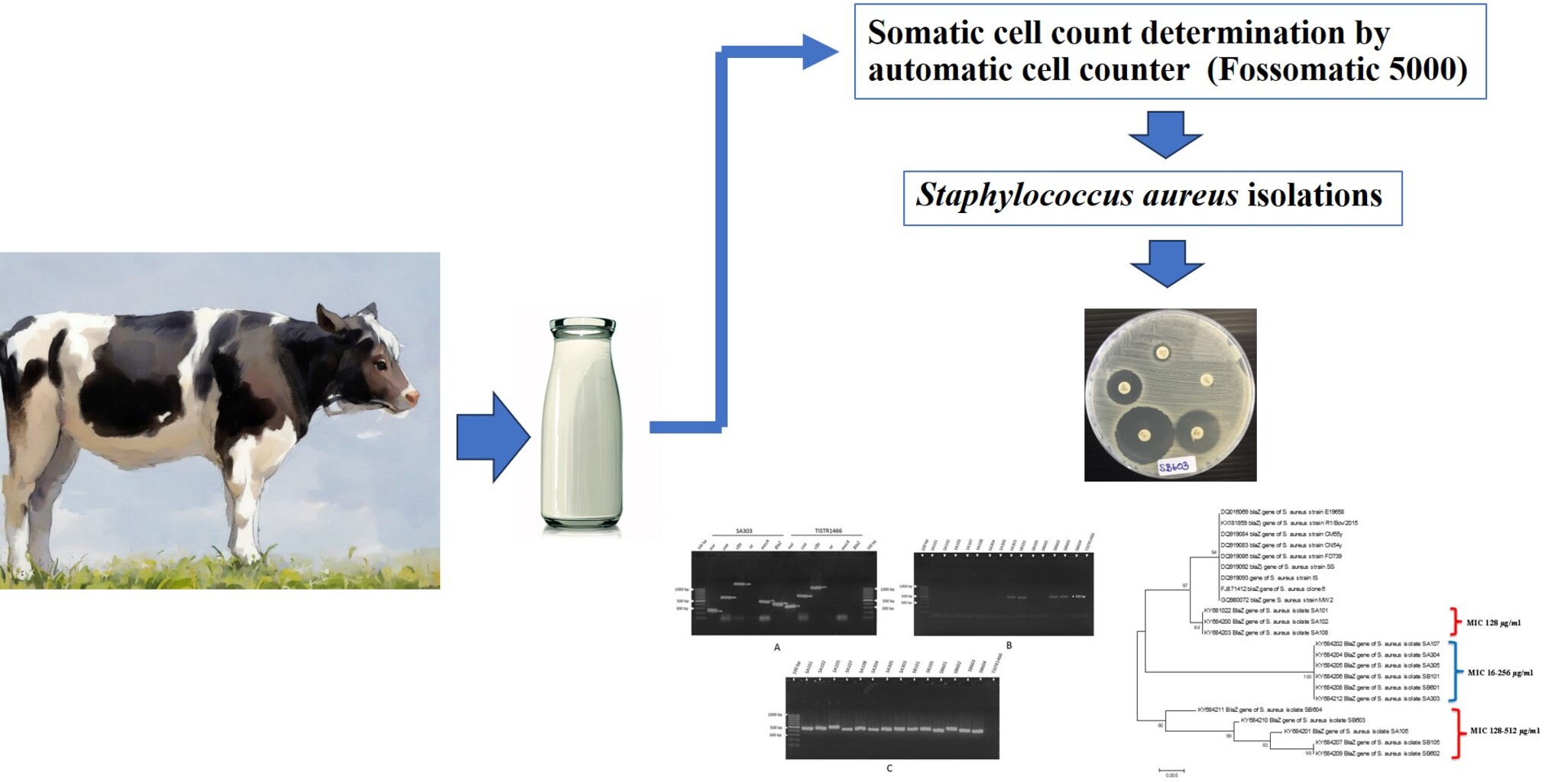
GRAPHICAL ABSTRACT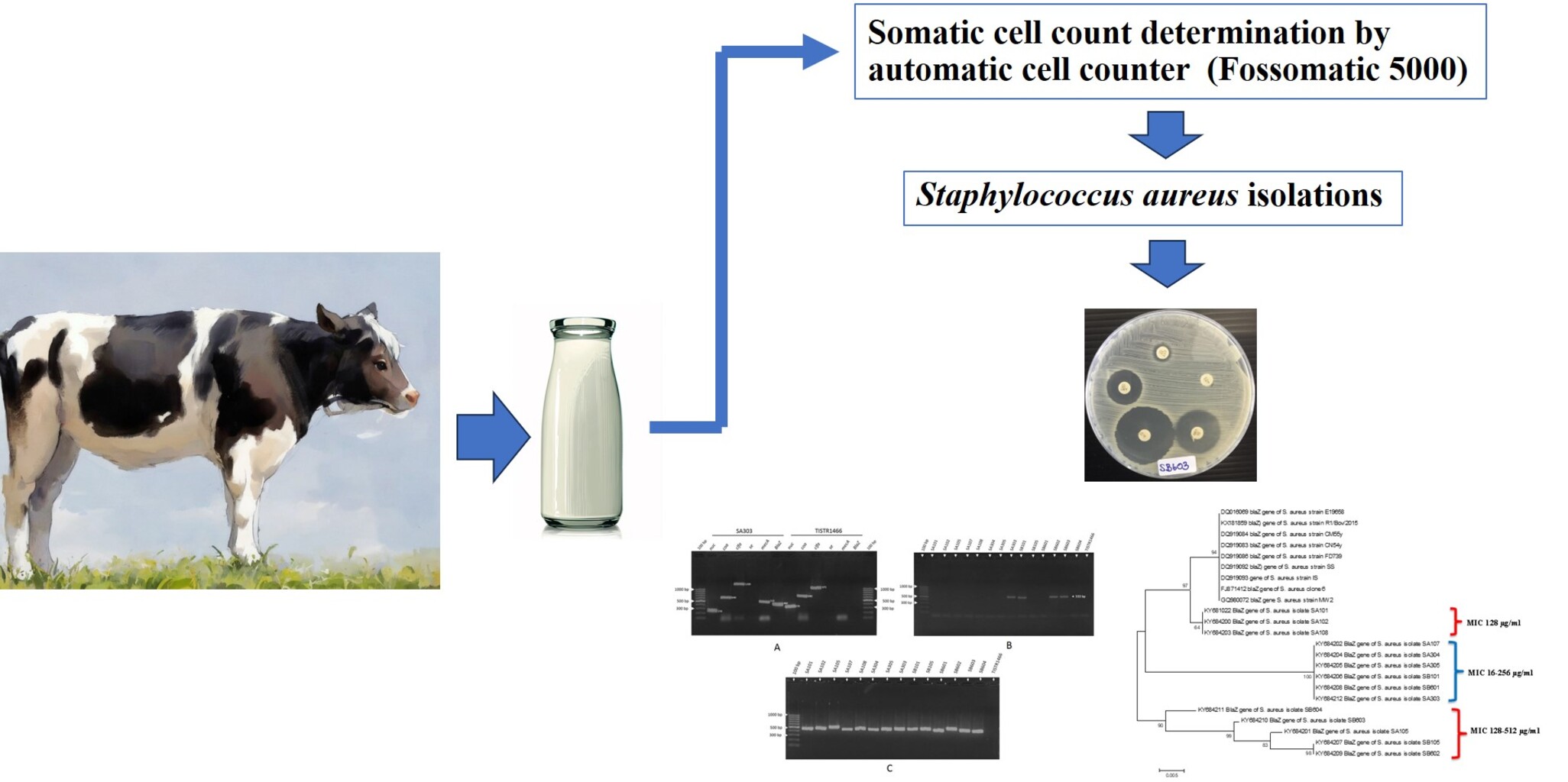
Downloads
References
R Paramasivam, DR Gopal, R Dhandapani, R Subbarayalu, MP Elangovan, B Prabhu, V Veerapan, A Nandheeswaran, S Paramasivam and S Muthupandian. Is AMR in dairy products a threat to human health? An updated review on the origin, prevention, treatment, and economic impacts of subclinical mastitis. Infection and Drug Resistance 2023; 16,155-178.
R Soundhararajan, M Veerasami, S Khan, A Islam and H Srinivasan. Polyherbal nano formulation: A potent antibiotic resistance breaker in bovine mastitis causing MDR pathogens. Journal of Molecular Liquids 2023; 392, 123477.
BL Crippa, LG de Matos, FN Souza and NCC Silva. Non-aureus staphylococci and mammaliicocci (NASM): Their role in bovine mastitis and One Health. Journal of Dairy Research 2024; 91(1), 44-56.
A Pesce, F Garofalo, P Coppa, GD Marco, B Cioffi, C Salzano and A Guarino. Antimicrobial resistance in pathogens causing mastitis. Large Animal Review 2013; 19(4), 162-164.
W Santiago-Neto, G Machado, DS Paim, TD Campos, MA Brito, MR Cardoso and LG Corbellini. Age related to the presence of antimicrobial resistant bacteria in twenty-one dairy herds in Rio Grande do Sul, Brazil. Pesquisa Veterinária Brasileira 2014; 34, 613-620.
WN Cheng and SG Han. Bovine mastitis: Risk factors, therapeutic strategies, and alternative treatments: A review. Asian-Australasian Journal of Animal Sciences 2020; 33(11), 1699.
J Kümmel, B Stessl, M Gonano, G Walcher, O Bereuter, M Fricker, T Grunert and M Wagner. Staphylococcus aureus entrance into the dairy chain: Tracking S. aureus from dairy cow to cheese. Frontiers in Microbiology 2016; 7, 1603.
BG Botaro, CS Cortinhas, AG Dibbern, LFPE Silva, NR Benites and MV dos Santos. Staphylococcus aureus intramammary infection affects milk yield and SCC of dairy cows. Tropical Animal Health and Production 2015; 47, 61-66.
KD Buchan, SJ Foster and SA Renshaw. Staphylococcus aureus: Setting its sights on the human innate immune system. Microbiology 2019; 165(4), 367-385.
M Greenlee-Wacker, FR DeLeo and WM Nauseef. How methicillin-resistant Staphylococcus aureus evade neutrophil killing. Current Opinion in Hematology 2015; 22(1), 30-35.
CL Martini, CC Lange, MA Brito, JB Ribeiro, LC Mendonça and EK Vaz. Characterisation of penicillin and tetracycline resistance in Staphylococcus aureus isolated from bovine milk samples in Minas Gerais Brazil. Journal of Dairy Research 2017; 84(2), 202-205.
E Kaczorek-Łukowska, J Małaczewska, P Sowińska, M Szymańska, EA Wójcik and AK Siwicki. Staphylococcus aureus from subclinical cases of mastitis in dairy cattle in Poland, what are they hiding? Antibiotic resistance and virulence profile. Pathogens 2022; 11(12), 1404.
N Tawil, F Mouawad, S Lévesque, E Sacher, R Mandeville and M Meunier. The differential detection of methicillin-resistant, methicillin-susceptible and borderline oxacillin-resistant Staphylococcus aureus by surface plasmon resonance. Biosensors and Bioelectronics 2013; 49, 334-340.
SJ Peacock and GK Paterson. Mechanisms of methicillin resistance in Staphylococcus aureus. Annual Review of Biochemistry 2015; 84, 577-601.
VV Gostev, OE Punchenko and VS Sergey. The current view on betalactam resistance in Staphylococcus aureus. Clinical Microbiology and Antimicrobial Chemotherapy 2022; 23(4), 375-387.
C Milheiriço, H de Lencastre and A Tomasz. Full-genome sequencing identifies in the genetic background several determinants that modulate the resistance phenotype in methicillin-resistant Staphylococcus aureus strains carrying the novel mecC gene. Antimicrobial Agents and Chemotherapy 2017; 61(3), e02500-16.
GD Rocha, JF Nogueira, MVG Dos Santos, JA Boaventura, RAN Soares, JJ de Simoni Gouveia, MM da Costa and GV Gouveia. Impact of polymorphisms in blaZ, blaR1 and blaI genes and their relationship with β-lactam resistance in S. aureus strains isolated from bovine mastitis. Microbial Pathogenesis 2022; 165, 105453.
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. Clinical and Laboratory Standards Institute, Wayne, 2024.
TA Hall. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 1999; 41, 95-98.
K Tamura, G Stecher and S Kumar. MEGA11: Molecular evolutionary genetics analysis version 11. Molecular Biology and Evolution 2021; 38(7), 3022-3027.
J Kampa, V Sukolapong, A Buttasri and A Charoenchai. Prevalence of Mycoplasma bovis and other contagious bovine mastitis pathogens in bulk tank milk of dairy cattle herds in Khon Kaen Province, Thailand. The Thai Journal of Veterinary Medicine 2009; 39(3), 275-280.
W Suriyasathaporn. Epidemiology of subclinical mastitis and their antibacterial susceptibility in smallholder dairy farms, Chiang Mai province, Thailand. Journal of Animal and Veterinary Advances 2011; 10(3), 316-321.
N Pumipuntu, W Tunyong, N Chantratita, P Diraphat, P Pumirat, N Sookrung, W Chaicumpa and N Indrawattana. Staphylococcus spp. associated with subclinical bovine mastitis in central and northeast provinces of Thailand. PeerJ 2019; 7, e6587.
F Javid, A Taku, MA Bhat, GA Badroo, M Mudasir and TA Sofi. Molecular typing of Staphylococcus aureus based on coagulase gene. Veterinary World 2018; 11, 423.
Y Hu, J Meng, C Shi, K Hervin, PM Fratamico and X Shi. Characterization and comparative analysis of a second thermonuclease from Staphylococcus aureus. Microbiological Research 2013; 168(3), 174-182.
MR Kiedrowski, HA Crosby, FJ Hernandez, CL Malone, JO McNamara and AR Horswill. Staphylococcus aureus Nuc2 is a functional, surface-attached extracellular nuclease. PloS One 2014; 9(4), e95574.
S Ashraf, J Cheng and X Zhao. Clumping factor A of Staphylococcus aureus interacts with Annexin A2 on mammary epithelial cells. Scientific Reports 2017; 7(1), 40608.
B Sinha and M Fraunholz. Staphylococcus aureus host cell invasion and post-invasion events. International journal of medical microbiology. International Journal of Medical Microbiology 2010; 300(2-3), 170-175.
C Heilmann. Adhesion mechanisms of staphylococci. Bacterial Adhesion: Chemistry, Biology and Physics 2011; 715, 105-123.
SW Kerrigan, N Clarke, A Loughman, G Meade, TJ Foster and D Cox. Molecular basis for Staphylococcus aureus-mediated platelet aggregate formation under arterial shear in vitro. Arteriosclerosis, Thrombosis, and Vascular Biology 2008; 28(2), 335-340.
CJ Wintersdorff, J Penders, JM Niekerk, ND Mills, S Majumder, LB Alphen, HM Savelkoul and PF Wolffs. Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Frontiers in Microbiology 2016; 7, 173.
S Tao, H Chen, N Li, T Wang and W Liang. The spread of antibiotic resistance genes in vivo model. Canadian Journal of Infectious Diseases and Medical Microbiology 2022; 1, 3348695.
P Patchanee, P Tadee, O Arjkumpa, D Love, K Chanachai, T Alter, S Hinjoy and P Tharavichitkul. Occurrence and characterisation of livestock-associated methicillin-resistant Staphylococcus aureus in pig industries of Northern Thailand. Journal of Veterinary Science 2014; 15(4), 529-536.
P Fungwithaya, K Boonchuay, R Narinthorn, N Sontigun, C Sansamur, Y Petcharat, T Thomrongsuwannakij and T Wongtawan. First study on diversity and antimicrobial-resistant profile of staphylococci in sports animals of Southern Thailand. Veterinary World 2022; 15(3), 765.
EY Klein, TP Van Boeckel, EM Martinez, S Pant, S Gandra, SA Levin, H Goossens and R Laxminarayan. Global increase and geographic convergence in anti-biotic consumption between 2000 and 2015. Proceedings of the National Academy of Sciences 2018; 115(15), E3463-E3470.
R Nomura, H Nakaminami, K Takasao, S Muramatsu, Y Kato, T Wajima and N Noguchi. A class A β-lactamase produced by borderline oxacillin-resistant Staphylococcus aureus hydrolyses oxacillin. Journal of Global Antimicrobial Resistance 2020; 22, 244-247.
S Rawat, N Shrivastava, A Shrivastav, S Singh, PK Singh, AK Niranjan and R Ranjan. Isolation and Characterization of Staphylococcus aureus in Bovine Milk from Rewa, India. Indian Journal of Microbiology 2024; 64, 1835-1845.
P Arêde, J Ministro and DC Oliveira. Redefining the role of the β-lactamase locus in methicillin-resistant Staphylococcus aureus: β-lactamase regulators disrupt the MecI-mediated strong repression on mecA and optimize the phenotypic expression of resistance in strains with constitutive mecA expression. Antimicrobial Agents and Chemotherapy 2013; 57(7), 3037-3045.
SM Cascarina and ED Ross. Natural and pathogenic protein sequence variation affecting prion-like domains within and across human proteomes. BMC Genomics 2020; 21, 23.
PJ Gómez-González, AD Grabowska, LD Tientcheu, AG Tsolaki, ML Hibberd, S Campino,JE Phelan and TG Clark. Functional genetic variation in pe/ppe genes contributes to diversity in Mycobacterium tuberculosis lineages and potential interactions with the human host. Frontiers in Microbiology 2023; 14, 1244319.
H Tahmasebi, B Zeiyni, S Dehbashi, H Motamedi, M Vafaeifar, F Keramat and MR Arabestani. The study of blaZ and mecA gene expression in methicillin-resistant Staphylococcus aureus strains and the relationship between the gene expression patterns. Journal of Isfahan Medical School 2017; 35(443), 1062-1067.
C Milheiriço, A Tomasz and HD Lencastre. Impact of the stringent stress response on the expression of methicillin resistance in Staphylococcaceae strains carrying mecA, mecA1 and mecC. Antibiotics 2022; 11(2), 255.
OG Brakstad, K Aasbakk and JA Maeland. Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. Journal of Clinical Microbiology 1992; 30(7), 1654-1660.
I Montesinos, E Salido, T Delgado, M Cuervo and A Sierra. Epidemiologic genotyping of methicillin-resistant Staphylococcus aureus by pulsed-field gel electrophoresis at a university hospital and comparison with antibiotyping and protein A and coagulase gene polymorphisms. Journal of Clinical Microbiology 2002; 40(6), 2119-2125.
PC Proietti, G Coppola, A Bietta, ML Marenzoni, DR Hyatt, M Coletti and F Passamonti. Characterization of genes encoding virulence determinants and toxins in Staphylococcus aureus from bovine milk in Central Italy. Journal of Veterinary Medical Science 2010; 72(11), 1443-1448.
R Stephan, C Annemüller, AA Hassan and C Lämmler. Characterization of enterotoxigenic Staphylococcus aureus strains isolated from bovine mastitis in north-east Switzerland. Veterinary Microbiology 2001; 78(4), 373-382.
M Kaase, S Lenga, S Friedrich, F Szabados, T Sakinc, B Kleine and G Gatermann. Comparison of phenotypic methods for penicillinase detection in Staphylococcus aureus. Clinical Microbiology and Infection 2008; 14(6), 614-616.
NL Dias, DCB Silva, DCBS Oliveira, FAA Junior, ML Sales and N Silva. Detection of genes of Staphylococcus aureus, enterotoxins and methicillin resistance in milk. Arquivo Brasileiro de Medicina Veterinária e Zootecnia 2011; 63, 1547-1552.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






