In Silico Study of Compounds from Nanoherbal Jopan (Clibadium surinamense L.) Leaves as Inhibitors AKT1 Interaction
DOI:
https://doi.org/10.48048/tis.2024.8729Keywords:
AKT1 inhibitors, Clibadium surinamense L, Breast cancer, Bioactive compounds, Molecular docking, Nanoherbal, Drug-likeness, GCMS, AKT1 inhibitors, Clibadium surinamense L., Breast cancer, Bioactive compounds, Nanoherbal, Molecular docking, Drug-likeness, GCMSAbstract
Breast cancer is the most prevalent cause of cancer-related deaths worldwide, and mortality rates are on the increase. This study aims to evaluate the potential of bioactive compounds from Clibadium surinamense L. leaves as inhibitors of AKT1 protein interactions, which play a crucial role in tumor growth mechanisms. Leaves of Clibadium surinamense L., obtained from Padangsidimpuan, North Sumatra, were extracted using methanol at a ratio of 1:20 for 48 h. The resulting extract was analyzed using Gas Chromatography-Mass Spectrometry (GC-MS) to identify the bioactive compounds. Drug likeness was then assessed according to Lipinski’s rules, and molecular docking analysis was performed using Autodock Vina. GC-MS results identified 11 bioactive compounds, including 2-Cyclopenten-1-one and Hexadecanoic acid, methyl ester. Of the 8 compounds evaluated, 6 met the criteria for drug candidates. Molecular docking analysis revealed significant interactions between the bioactive compounds and the AKT1 protein. These findings suggest that bioactive compounds from Clibadium surinamense L. have potential as AKT1 interaction inhibitors, which could enhance cancer treatment mechanisms. This research paves the way for further studies on the therapeutic applications of these compounds in breast cancer treatment.
HIGHLIGHTS
- Bioactive compounds were identified from Clibadium surinamense leaf extract using the Gas Chromatography-Mass Spectrometry (GC-MS) technique.
- Molecular docking analysis showed significant interactions between bioactive compounds and AKT1 protein.
- Bioactive compounds from Clibadium surinamense have the potential to function as AKT1 interaction inhibitors to increase the effectiveness of breast cancer treatment.
- These findings pave the way for further studies regarding the therapeutic applications of these compounds in cancer treatment.
GRAPHICAL ABSTRACT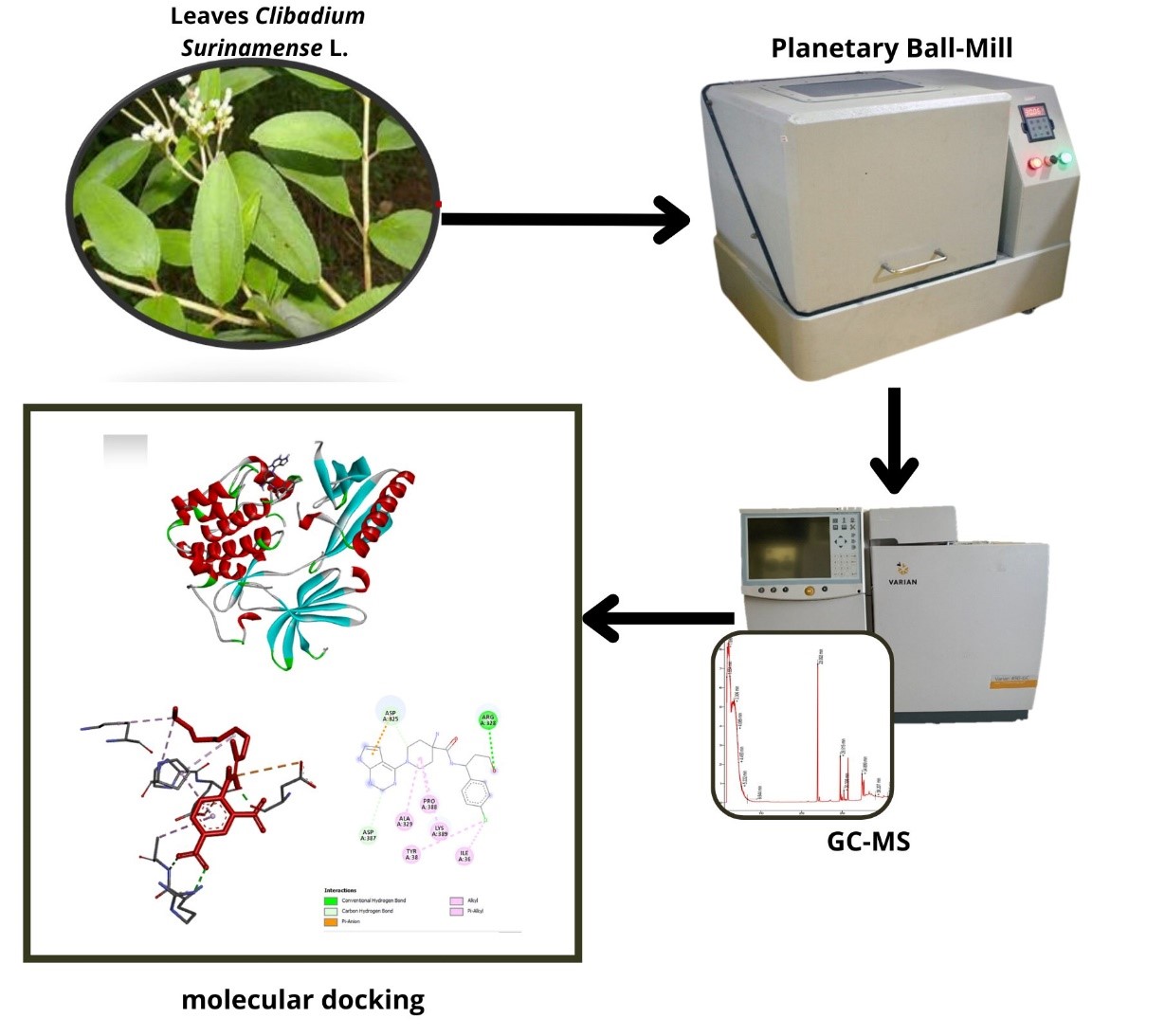
Downloads
References
F Bray, M Laversanne, H Sung, J Ferlay, RL Siegel, I Soerjomataram and A Jemal. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians 2024; 74(3), 229-263.
SK Kim and SW Cho. The evasion mechanisms of cancer immunity and drug intervention in the tumor microenvironment. Frontiers in Pharmacology 2022; 13, 868695.
R Tang, J Xu, B Zhang, J Liu, C Liang, J Hua, Q Meng, X Yu and S Shi. Ferroptosis, necroptosis, and pyroptosis in anticancer immunity. Journal of Hematology & Oncology 2020; 13, 110.
MR Paul, TC Pan, DK Pant, NNC Shih, Y Chen, KL Harvey, A Solomon, D Lieberman, JJD Morrissette, D Soucier-Ernst, NG Goodman, SW Stavropoulos, KN Maxwell, C Clark, GK Belka, M Feldman, A DeMichele and LA Chodosh. Genomic landscape of metastatic breast cancer identifies preferentially dysregulated pathways and targets. The Journal of Clinical Investigation 2020; 130(8), 4252-4265.
MM Kenna, S McGarrigle and GP Pidgeon. The next generation of PI3K-Akt-MTOR pathway inhibitors in breast cancer cohorts. Biochimica et Biophysica Acta (BBA) - Reviews on Cancer 2018; 1870(2), 185-197.
MK Fath, RA Masouleh, N Afifi, S Loghmani, P Tamimi, A Fazeli, SA Mousavian, MM Falsafi and G Barati. PI3K/AKT/MTOR signaling pathway modulation by circular RNAs in breast cancer progression. Pathology - Research and Practice 2023; 241, 154279.
MA Ortega, O Fraile-Martínez, Á Asúnsolo, J Buján, N García-Honduvilla and S Coca. Signal transduction pathways in breast cancer: The important role of PI3K/Akt/MTOR. Journal of Oncology 2020; 2020, 9258396.
N Hinz and M Jücker. Distinct functions of AKT isoforms in breast cancer: A comprehensive review. Cell Communication and Signaling 2019; 17, 154.
M Song, AM Bode, Z Dong and MH Lee. AKt as a therapeutic target for cancer. Cancer Research 2019; 79(6), 1019-1031.
A Basu and CB Lambring. Akt isoforms: A family affair in breast cancer. Cancers 2021; 13, 3445.
D Miricescu, A Totan, II Stanescu-Spinu, SC Badoiu, C Stefani and M Greabu. PI3K/AKT/MTOR signaling pathway in breast cancer: From molecular landscape to clinical aspects. International Journal of Molecular Sciences 2021; 22(1), 173.
A Andrikopoulou, S Chatzinikolaou, E Panourgias, M Kaparelou, M Liontos, MA Dimopoulos and F Zagouri. The emerging role of capivasertib in breast cancer. Breast 2022; 63, 157-167.
LM Smyth, K Tamura, M Oliveira, E Ciruelos, IA Mayer, MP Sablin, L Biganzoli, H Ambrose, J Ashton, A Barnicle, D Cashell, C Corcoran, EC de Bruin, A Foxley, J Hauser, JPO Lindemann, R Maudsley, R McEwen, M Moschetta, …, DM Hyman. Capivasertib, an AKT kinase inhibitor, as monotherapy or in combination with fulvestrant in patients with AKT1E17K-mutant, ER-positive metastatic breast cancer. Clinical Cancer Research 2020; 26(15), 3947-3957.
Y Yuan, H Long, Z Zhou, Y Fu and B Jiang. PI3K-AKT-targeting breast cancer treatments: Natural products and synthetic compounds. Biomolecules 2023; 13(1), 93.
MK Dewi, AY Chaerunisaa, M Muhaimin and IM Joni. Improved activity of herbal medicines through nanotechnology. Nanomaterials 2022; 12(22), 4073.
Y Guo, Y Zhao, X Zhao, S Song and B Qian. Exploring the anticancer effects of tin oxide nanoparticles synthesized by pulsed laser ablation technique against breast cancer cell line through downregulation of PI3K/AKT/MTOR signaling pathway. Arabian Journal of Chemistry 2021; 14(7), 103212.
G Baskar, T Palaniyandi, M Ravi, S Viswanathan, MRA Wahab, H Surendran, M Govindaraj, A Sugumaran, MH Almutairi and BO Almutairi. Biosynthesis of iron oxide nanoparticles from red seaweed hypnea valentiae and evaluation of their antioxidant and antitumor potential via the AKT/PI3K pathway. Process Biochemistry 2024; 141, 155-169.
M Ali, S Ud, D Wani, SN Manjula, K Mruthunjaya, T Dey, M Iqbal and J Singh. Recent advance of herbal medicines in cancer - a molecular approach. Heliyon 2023; 9(2), e13684.
DA Mcgrowder, FG Miller, CR Nwokocha, MS Anderson, C Wilson-clarke, K Vaz, L Anderson-jackson and J Brown. Medicinal herbs used in traditional management of breast cancer: Mechanisms of action. Medicines 2020; 7(8), 47.
RV Ulia, Suryati and A Santoni. Cytotoxic potential of essential oil isolated from Semambu (Clibadium Surinamese L) leaves against T47D breast and HeLa cervical cancer cells. Molekul 2023; 18(2), 289-299.
MK Swamy, UR Sinniah and MS Akhtar. In vitro pharmacological activities and GC-Ms analysis of different solvent extracts of Lantana camara leaves collected from tropical region of Malaysia. Evidence-Based Complementary and Alternative Medicine 2015; 2015, 506413.
S Dallakyan and AJ Olson. Small-molecule library screening by docking with PyRx. In: J Hempel, C Williams and C Hong (Eds.). Chemical biology. Methods in molecular biology. Vol 1263. Humana Press, New York, p. 243-250.
Y Peng, Y Wang, C Zhou, W Mei and C Zeng. PI3K/Akt/MTOR pathway and its role in cancer therapeutics: Are we making headway? Frontiers in Oncology 2022; 12, 819128.
H Hua, H Zhang, J Chen, J Wang, J Liu and Y Jiang. Targeting Akt in cancer for precision therapy. Journal of Hematology & Oncology 2021; 14, 128.
D Das, M Nanda, P Banjare and S Lanjhiyana. Exploration of multitargeted antialzheimer’s activity of safflower leaves phytoconstituents: In silico molecular docking approach. European Journal of Medicinal Chemistry Reports 2024; 10, 100119.
S Rosignoli and A Paiardini. DockingPie: A consensus docking plugin for PyMOL. Bioinformatics 2022; 38(17), 4233-4234.
P Santoso, S Ilyas, YH Midoen, R Maliza and DF Belahusna. Predictive bioactivity of compounds from Vitis gracilis leaf extract to counteract doxorubicin-induced cardiotoxicity via sirtuin 1 and adenosine monophosphate-activated protein kinase: An in-silico study. Journal of Applied Pharmaceutical Science 2024; 14(4), 99-114.
A Daina, O Michielin and V Zoete. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Scientific Reports 2017; 7, 42717.
A Allouche. Software news and update autodock vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of Computational Chemistry 2010; 31(2), 174-182.
M Azmal, MS Hossen, MNH Shohan, R Taqui, A Malik and A Ghosh. A computational approach to identify phytochemicals as potential inhibitor of acetylcholinesterase: Molecular docking, ADME profiling and molecular dynamics simulations. PLoS One 2024; 19(6), e0304490.
JF Ayala-Cabrera, L Montero, SW Meckelmann, F Uteschil and OJ Schmitz. Review on atmospheric pressure ionization sources for gas chromatography-mass spectrometry. Part II: Current applications. Analytica Chimica Acta 2023; 1238, 340379.
C Abadie, J Lalande and G Tcherkez. Exact mass GC-MS analysis: Protocol, database, advantages and application to plant metabolic profiling. Plant, Cell & Environment 2022; 45, 3171-3183.
Y Marcus. Extraction by subcritical and supercriticalwater, methanol, ethanol and their mixtures. Separations 2018; 5(1), 4.
DA Filimonov, AA Lagunin, TA Gloriozova, AV Rudik, DS Druzhilovskii, PV Pogodin and VV Poroikov. Prediction of the biological activity spectra of organic compounds using the pass online web resource. Chemistry of Heterocyclic Compounds 2014; 50, 444-457.
P Vijaya and G Sundaraselvan. Synthesis, characterization, PASS analysis and ADMET properties of oxazolone ring containing hydrazone derivatives. Materials Today: Proceedings 2019; 48, 502-507.
SM Ivanov, AA Lagunin, DA Filimonov and VV Poroikov. Relationships between the structure and severe drug-induced liver injury for low, medium, and high doses of drugs. Chemical Research in Toxicology 2022; 35(3), 402-411.
A Tsantili-kakoulidou and VJ Demopoulos. Drug-like properties and fraction lipophilicity index as a combined metric. ADMET DMPK 2021; 9(3), 177-190.
GR Bickerton, GV Paolini, J Besnard, S Muresan and AL Hopkins. Quantifying the chemical beauty of drugs. Nature Chemistry 2012; 4, 90-98.
AL Whitworth, NH Mann and AWD Larkum. Inhibition assays for AKT1, ∆PH-AKT1, AKT2, ∆PH-AKT2 and AKT3 activity, AKT1 activation, and binding kinetics. Ultrasound in Obstetrics & Gynecology 2006; 50, 776-780.
Z Qureshi, F Altaf, M Khanzada, Z Zaheer, E Fatima and M Bakhtiar. Capivasertib in hormone receptor-positive, human epidermal growth factor receptor 2-negative advanced breast cancer. Current Problems in Cancer 2024; 51, 101114.
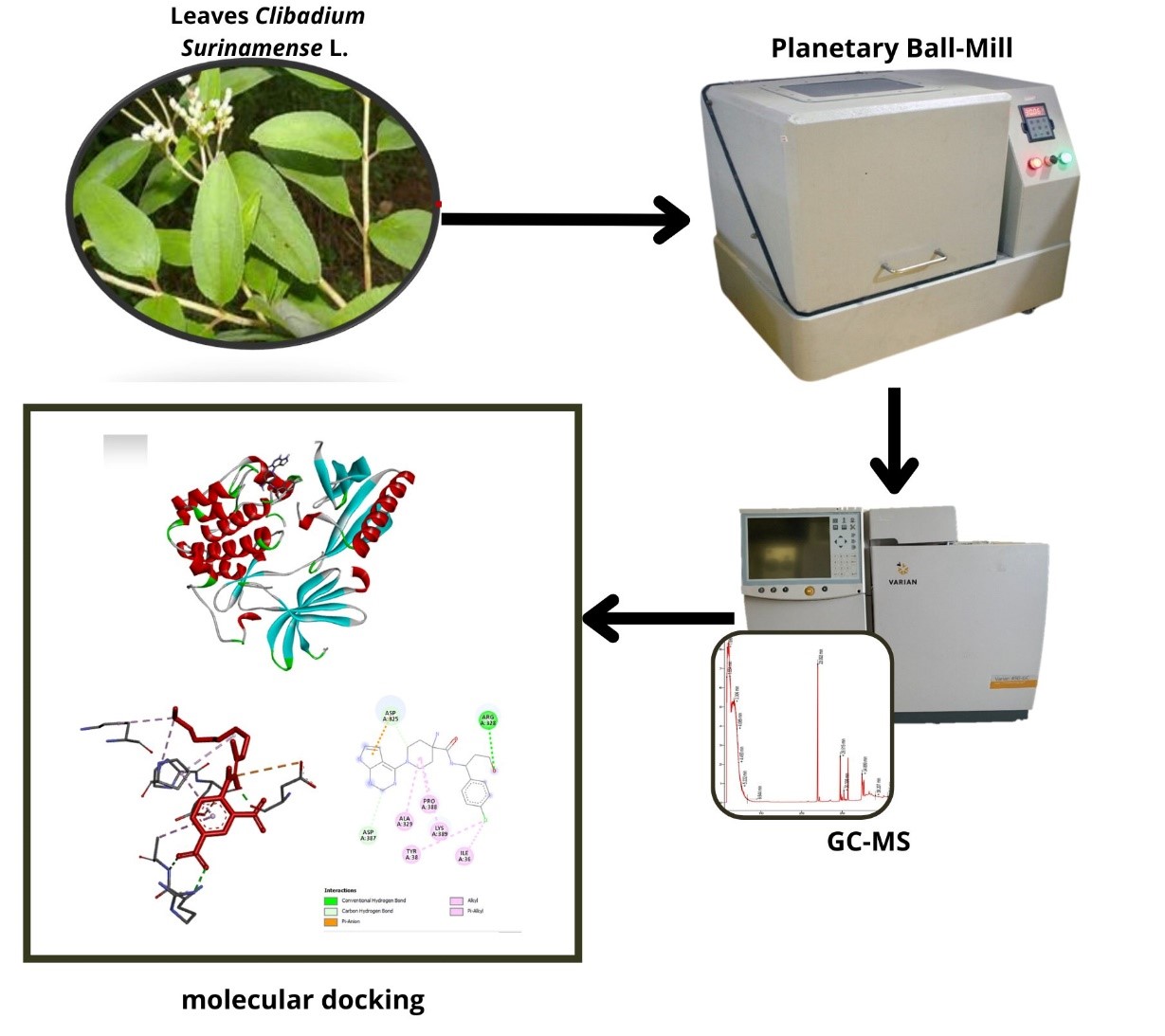
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2024 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






