Lagerstroemin from Lagerstroemia speciosa as Antibreast Cancer Candidate Targeting AURKA, EGFR and SRC Protein: A Comprehensive Computational Study
DOI:
https://doi.org/10.48048/tis.2024.8205Keywords:
Apoptosis, AURKA, Breast cancer, EGFR, L. speciosa, SRCAbstract
Breast cancer is a type of cancer that has a high rate of diagnosis and mortality in the world. Lagerestroemia speciosa is a herb that has anticancer activity. This study aims to analyze the compounds in L. speciosa that most act as anticancer of the breast. The compounds in L. speciosa were selected based on drug-likeness, physicochemical properties, ability to penetrate the lipid bilayer and toxicity. The Lagerstroemin target related to Lagestroemin breast cancer was predicted using DisGeNET, SWISS Target Prediction and cBioportal. Molecular docking between Lagerstroemin and AURKA, EGFR and SRC was performed using AutoDock Vina. The interaction stability of each complex was analyzed by molecular dynamic simulation using YASARA with parameters of RMSD protein, RMSD ligand, number of hydrogen bonds and molecular dynamic binding energy. Of the 22 compounds, Quercetin, Caffeic acid, Lagerstroemin and Dypirone were predicted to have good ADME properties and can easily penetrate lipid membranes. Therefore, Quercetin, Caffeic acid and Lagerstroemin were predicted to have anti-breast cancer bioactivity. Of the 3 compounds, Lagerstroemin had the lowest toxicity. Lagerstroemin was predicted to interact with breast cancer-related proteins AURKA, EGFR and SRC. Molecular docking and dynamics showed that Lagerestroemin interacted stably at the ATP binding site of the 3 proteins, so it was very potential as an inhibitor of these 3 proteins. Therefore, Lagerstroemin was predicted to be the compound in L. speciosa with the most potential breast anticancer agent by targeting AURKA, EGFR and SRC.
HIGHLIGHTS
- Quercetin, Caffeic acid and Lagerstroemin from Lagerstroemia speciosa show promising anticancer properties for breast cancer treatment.
- Lagerstroemin stands out with low toxicity and potent anticancer activity, suggesting its safety and effectiveness as a therapeutic agent.
- Lagerstroemin interacts stably with breast cancer-related proteins AURKA, EGFR and SRC, indicating its potential as a powerful inhibitor targeting key pathways in breast cancer.
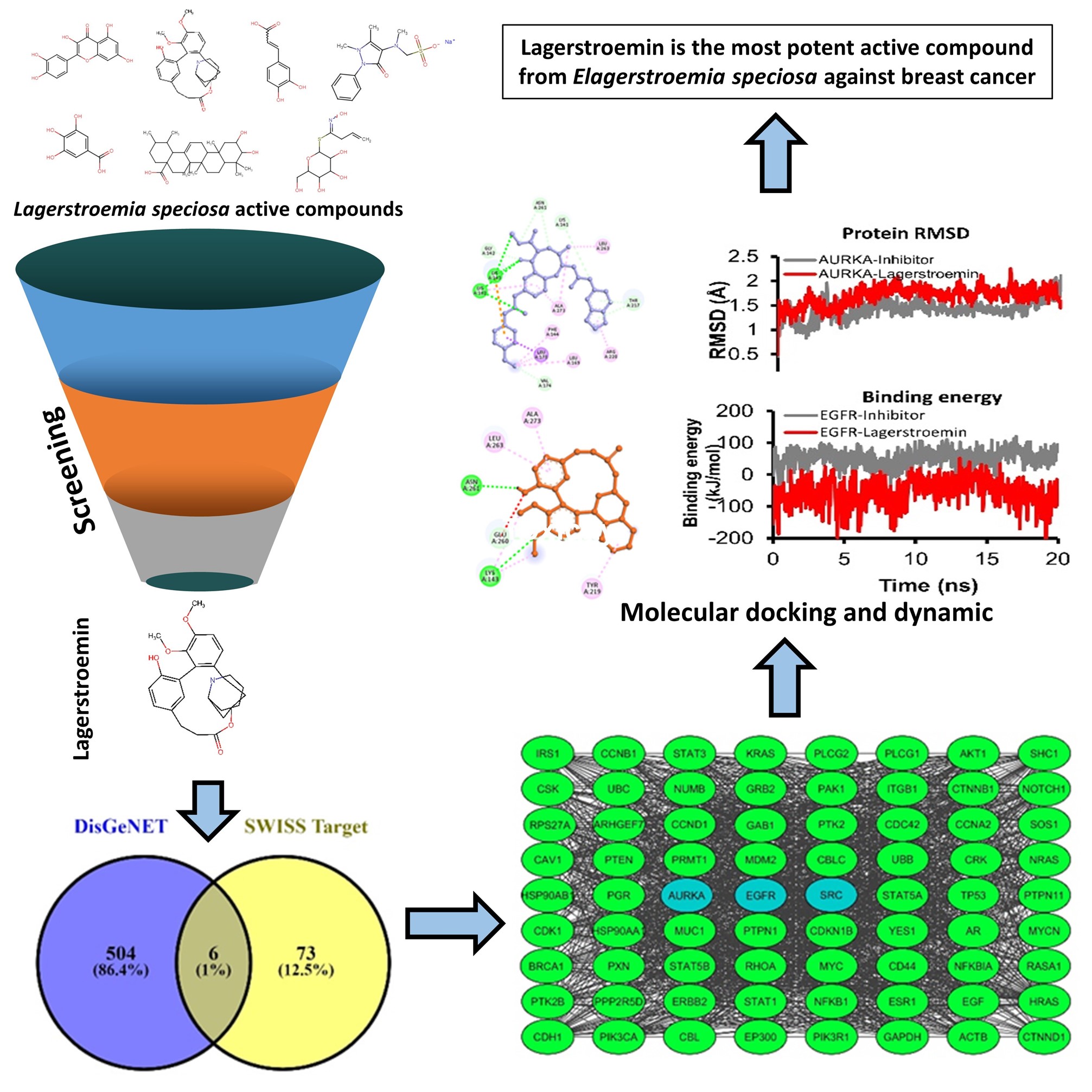
GRAPHICAL ABSTRACT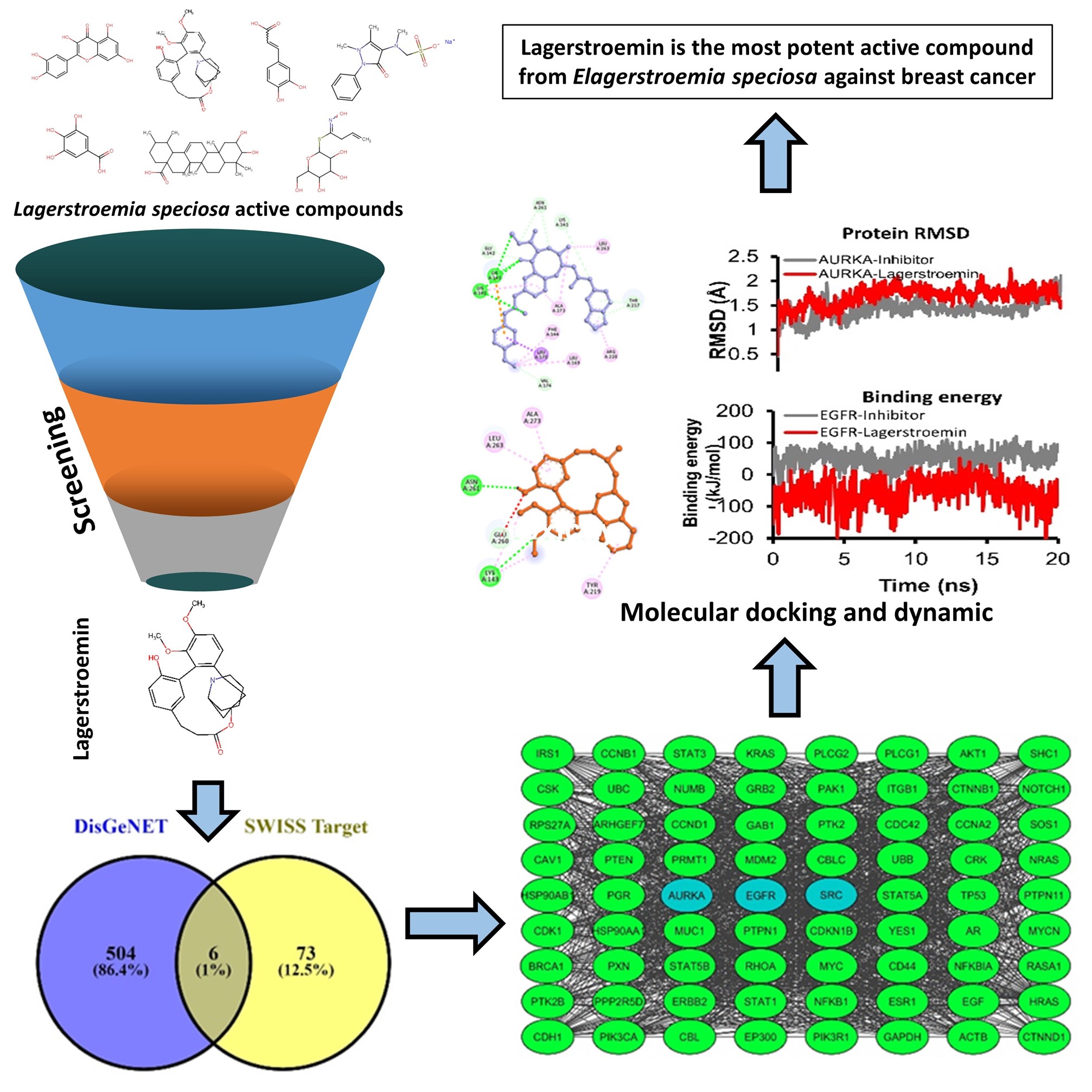
Downloads
References
BS Chhikara and K Parang. Global cancer statistics 2022: The trends projection analysis. Chem. Biol. Lett. 2023; 10, 451.
M Akram, M Iqbal, M Daniyal and AU Khan. Awareness and current knowledge of breast cancer. Biol. Res. 2017; 50, 33.
EA Nigg. Mitotic kinases as regulators of cell division and its checkpoints. Nat. Rev. Mol. Cell Biol. 2001; 2, 21-32.
Y Cirak, Y Furuncuoglu, O Yapicier, A Aksu and E Cubukcu. Aurora A overexpression in breast cancer patients induces taxane resistance and results in worse prognosis. J. Balkan Union Oncol. 2015; 20, 1414-9.
A Katsha, A Belkhiri, L Goff and W El-Rifai. Aurora kinase A in gastrointestinal cancers: Time to target. Mol. Cancer 2015; 14, 106.
TV Do, F Xiao, LE Bickel, AJ Klein-Szanto, HB Pathak, X Hua, C Howe, SW O’Brien, M Maglaty, JA Ecsedy, S Litwin, EA Golemis, RJ Schilder, AK Godwin and DC Connolly. Aurora kinase A mediates epithelial ovarian cancer cell migration and adhesion. Oncogene 2014; 33, 539-49.
S Sankaran, DE Crone, RE Palazzo and JD Parvin. Aurora-A kinase regulates breast cancer-associated gene 1 inhibition of centrosome-dependent microtubule nucleation. Cancer Res. 2007; 67, 11186-94.
S Sigismund, D Avanzato and L Lanzetti. Emerging functions of the EGFR in cancer. Mol. Oncol. 2018; 12, 3-20.
A Oprita, SC Baloi, GA Staicu, O Alexandru, DE Tache, S Danoiu, ES Micu and AS Sevastre. Updated insights on EGFR signaling pathways in glioma. Int. J. Mol. Sci. 2021; 22, 587.
AE Maennling, MK Tur, M Niebert, T Klockenbring, F Zeppernick, S Gattenlöhner, I Meinhold-Heerlein and AF Hussain. Molecular targeting therapy against EGFR family in breast cancer: Progress and future potentials. Cancers 2019; 11, 1826.
S Belli, D Esposito, A Servetto, A Pesapane, L Formisano and R Bianco. c-Src and EGFR inhibition in molecular cancer therapy: What else can we improve? Cancers 2020; 12, 1489.
SA Courtneidge. Isolation of novel Src substrates. Biochem. Soc. Trans. 2003; 31, 25-8.
TP Wall, PD Crowley, A Sherwin, AG Foley and DJ Buggy. Effects of lidocaine and Src inhibition on metastasis in a murine model of breast cancer surgery. Cancers 2019; 11, 1414.
T Amaral and C Garbe. Non-melanoma skin cancer: New and future synthetic drug treatments. Expert. Opin. Pharmacother. 2017; 18, 689-99.
A Karimi, M Majlesi and M Rafieian-Kopaei. Herbal versus synthetic drugs; beliefs and facts. J. Nephropharmacol. 2015; 4, 27-30.
AE Al-Snafi. Medicinal value of lagerstroemia speciosa: An updated review. Int. J. Curr. Pharm. Sci. 2019; 11, 18-26.
T Sharmin, MS Rahman and H Mohammadi. Investigation of biological activities of the flowers of Lagerstroemia speciosa, the Jarul flower of Bangladesh. BMC Compl. Altern. Med. 2018; 18, 231.
VS Saraswathi and K Santhakuma. Photocatalytic activity against azo dye and cytotoxicity on MCF-7 cell lines of zirconium oxide nanoparticle mediated using leaves of Lagerstroemia speciosa. J. Photochem. Photobiol. B 2017; 169, 47-55.
B Shaker, S Ahmad, J Lee, C Jung and D Na. In silico methods and tools for drug discovery. Comput. Biol. Med. 2021; 137, 104851.
Y Tang, W Zhu, K Chen and H Jiang. New technologies in computer-aided drug design: Toward target identification and new chemical entity discovery. Drug Discovery Today Tech. 2006; 3, 307-13.
MH Widyananda, ST Wicaksono, K Rahmawati, S Puspitarini, SM Ulfa, YD Jatmiko, M Masruri and N Widodo. A potential anticancer mechanism of finger root (Boesenbergia rotunda) extracts against a breast cancer cell line. Scientifica 2022; 2022, 9130252.
S Guo, X Ren, K He, X Chen, S Zhang, M Roller, B Zheng, Q Zheng, CT Ho and N Bai. The anti-diabetic effect of eight Lagerstroemia speciosa leaf extracts based on the contents of ellagitannins and ellagic acid derivatives. Food Funct. 2020; 11, 1560-71.
P Kolakul and B Sripanidkulchai. Phytochemicals and anti-aging potentials of the extracts from Lagerstroemia speciosa and Lagerstroemia floribunda. Ind. Crops Prod. 2017; 109, 707-16.
BK Tiwary, S Dutta, P Dey, M Hossain, A Kumar, S Bihani, AK Nanda, TK Chaudhuri and R Chakraborty. Radical scavenging activities of Lagerstroemia speciosa (L.) Pers. Petal extracts and its hepato-protection in CCl4-intoxicated mice. BMC Compl. Altern. Med. 2017; 17, 55.
W Hou, Y Li, Q Zhang, X Wei, A Peng, L Chen and Y Wei. Triterpene acids isolated from Lagerstroemia speciosa leaves as α-glucosidase inhibitors. Phytother. Res. 2009; 23, 614-8.
CA Lipinski. Lead-and drug-like compounds: The rule-of-five revolution. Drug Discov. Today Tech. 2004; 1, 337-41.
WJ Egan, KM Merz and JJ Baldwin. Prediction of drug absorption using multivariate statistics. J. Med. Chem. 2000; 43, 3867-77.
DF Veber, SR Johnson, HY Cheng, BR Smith, KW Ward and KD Kopple. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 2002; 45, 2615-23.
AK Ghose, VN Viswanadhan and JJ Wendoloski. A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. A qualitative and quantitative characterization of known drug databases. J. Comb. Chem. 1999; 1, 55-68.
AL Lomize, JM Hage, K Schnitzer, K Golobokov, MB LaFaive, AC Forsyth and ID Pogozheva. PerMM: A web tool and database for analysis of passive membrane permeability and translocation pathways of bioactive molecules. J. Chem. Inform. Model. 2019; 59, 3094-9.
AA Lagunin, VI Dubovskaja, AV Rudik, PV Pogodin, DS Druzhilovskiy, TA Gloriozova, DA Filimonov, NG Sastry and VV Poroikov. CLC-Pred: A freely available web-service for in silico prediction of human cell line cytotoxicity for drug-like compounds. PLoS One 2018; 13, e0191838.
A Lagunin, A Rudik, D Druzhilovsky, D Filimonov, V Poroikov and J Wren. ROSC-Pred: Web-service for rodent organ-specific carcinogenicity prediction. Bioinformatics 2018; 34, 710-2.
P Banerjee, AO Eckert, AK Schrey and R Preissner. ProTox-II: A webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2018; 46, W257-W263.
P Razavi, MN Dickler, PD Shah, W Toy, DN Brown, HH Won, BT Li, R Shen, N Vasan, S Modi, K Jhaveri, BA Caravella, S Patil, P Selenica, S Zamora, AM Cowan, E Comen, A Singh, A Covey, … S Chandarlapaty. Alterations in PTEN and ESR1 promote clinical resistance to alpelisib plus aromatase inhibitors. Nat. Cancer 2020; 1, 382-93.
P Razavi, MT Chang, G Xu, C Bandlamudi, DS Ross, N Vasan, Y Cai, CM Bielski, MTA Donoghue, P Jonsson, A Penson, R Shen, F Pareja, R Kundra, S Middha, ML Cheng, A Zehir, C Kandoth, R Patel, … J Baselga. The genomic landscape of endocrine-resistant advanced breast cancers. Cancer Cell 2018; 34, 427-38.
Q Li, B Jiang, J Guo, H Shao, ISD Priore, Q Chang, R Kudo, Z Li, P Razavi, B Liu, AS Boghossian, MG Rees, MM Ronan, JA Roth, KA Donovan, M Palafox, JS Reis-Filho, ED Stanchina, ES Fischer, N Rosen, V Serra, … and S Chandarlapaty. INK4 tumor suppressor proteins mediate resistance to CDK4/6 kinase inhibitors. Cancer Discovery 2022; 12, 356-71.
BT Sherman, M Hao, J Qiu, X Jiao, MW Baseler, HC Lane, T Imamichi and W Chang. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022; 50, W216-W221.
NM O’Boyle, M Banck, CA James, C Morley, T Vandermeersch and GR Hutchison. Open babel: An open chemical toolbox. J. Cheminform. 2011; 3, 33.
O Trott and AJ Olson. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2009; 31, 455-61.
MH Widyananda, SK Pratama, RS Samoedra, FN Sari, VD Kharisma, ANM Ansori and Y Antonius. Molecular docking study of sea urchin (Arbacia lixula) peptides as multi-target inhibitor for non-small cell lung cancer (NSCLC) associated proteins. J. Pharm. Pharmacogn. Res. 2021; 9, 484-96.
E Krieger and G Vriend. New ways to boost molecular dynamics simulations. J. Comput. Chem. 2015; 36, 996-1007.
CY Jia, JY Li, GF Hao and GF Yang. A drug-likeness toolbox facilitates ADMET study in drug discovery. Drug Discovery Today 2020; 25, 248-58.
TL Wargasetia, H Ratnawati, N Widodo and MH Widyananda. Antioxidant and anti-inflammatory activity of sea cucumber (Holothuria scabra) active compounds against KEAP1 and iNOS protein. Bioinform. Biol. Insights 2023; 17, 1-10.
DA Filimonov, AA Lagunin, TA Gloriozova, AV Rudik, DS Druzhilovskii, PV Pogodin and VV Poroikov. Prediction of the biological activity spectra of organic compounds using the pass online web resource. Chem. Heterocycl. Comp. 2014; 50, 444-57.
M Janeček, M Rossmann, P Sharma, A Emery, DJ Huggins, SR Stockwell, JE Stokes, YS Tan, EG Almeida, B Hardwick, AJ Narvaez, M Hyvönen, DR Spring, GJ McKenzie and AR Venkitaraman. Allosteric modulation of AURKA kinase activity by a small-molecule inhibitor of its protein-protein interaction with TPX. Sci. Rep. 2016; 6, 28528.
AS Nikonova, I Astsaturov, IG Serebriiskii, RL Dunbrack and EA Golemis. Aurora A kinase (AURKA) in normal and pathological cell division. Cell. Mol. Life Sci. 2013; 70, 661-87.
TT Komoto, TM Bernardes, TB Mesquita, LFB Bortolotto, G Silva, TA Bitencourt, SJ Baek, M Marins and AL Fachin. Chalcones repressed the AURKA and MDR proteins involved in metastasis and multiple drug resistance in breast cancer cell lines. Molecules 2018; 23, 2018.
F Wang, H Zhang, H Wang, T Qiu, B He and Q Yang. Combination of AURKA inhibitor and HSP90 inhibitor to treat breast cancer with AURKA overexpression and TP53 mutations. Med. Oncol. 2022; 39, 180.
Z Zou, Z Yuan, Q Zhang, Z Long, J Chen, Z Tang, Y Zhu, S Chen, J Xu, M Yan, J Wang and Q Liu. Aurora kinase A inhibition-induced autophagy triggers drug resistance in breast cancer cells. Autophagy 2012; 8, 1798-810.
L Xia, Z Zheng, JY Liu, YJ Chen, J Ding, GS Hu, YH Hu, S Liu, WX Luo, NS Xia and W Liu. Targeting triple-negative breast cancer with combination therapy of EGFR CAR T cells and CDK7 inhibition. Cancer Immunol. Res. 2021; 9, 707-22.
L Song, Z Liu, HH Hu, Y Yang, TY Li, ZZ Lin, J Ye, J Chen, X Huang, DT Liu, J Zhou, Y Shi, H Zhao, C Xie, L Chen, E Song, SY Lin and SC Lin. Proto-oncogene Src links lipogenesis via lipin-1 to breast cancer malignancy. Nat. Commun. 2020; 11, 5842.
PC Hsu, CT Yang, DM Jablons and L You. The crosstalk between Src and hippo/YAP signaling pathways in Non-Small Cell Lung Cancer (NSCLC). Cancers 2020; 12, 1361.
S Jain, X Wang, CC Chang, C Ibarra-Drendall, H Wang, Q Zhang, SW Brady, P Li, H Zhao, J Dobbs, M Kyrish, TS Tkaczyk, A Ambrose, C Sistrunk, BK Arun, R Richards-Kortum, W Jia, VL Seewaldt and D Yu. Src inhibition blocks c-Myc translation and glucose metabolism to prevent the development of breast cancer. Cancer Res. 2015; 75, 4863-75.
KRA Azeez, S Chatterjee, C Yu, TR Golub, F Sobott and JM Elkins. Structural mechanism of synergistic activation of Aurora kinase B/C by phosphorylated INCENP. Nat. Commun. 2019; 10, 3166.
KS Gajiwala, J Feng, R Ferre, K Ryan, O Brodsky, S Weinrich, JC Kath and A Stewart. Insights into the aberrant activity of mutant EGFR kinase domain and drug recognition. Structure 2013; 21, 209-19.
SW Cowan-Jacob, G Fendrich, PW Manley, W Jahnke, D Fabbro, J Liebetanz and T Meyer. The crystal structure of a c-Src complex in an active conformation suggests possible steps in c-Src activation. Structure 2005; 13, 861-71.
L Martínez. Automatic identification of mobile and rigid substructures in molecular dynamics simulations and fractional structural fluctuation analysis. PLoS One 2015; 10, e0119264.
M Majewski, S Ruiz-Carmona and X Barril. An investigation of structural stability in protein-ligand complexes reveals the balance between order and disorder. Commun. Chem. 2019; 2, 110.
DE Chen, DL Willick, JB Ruckel and WB Floriano. Principal component analysis of binding energies for single-point mutants of hT2R16 bound to an agonist correlate with experimental mutant cell response. J. Comput. Biol. 2015; 22, 37-53.
NH Nguyen, QTH Ta, QT Pham, TNH Luong, VT Phung, TH Duong and VG Vo. Anticancer activity of novel plant extracts and compounds from Adenosma bracteosum (Bonati) in human lung and liver cancer cells. Molecules 2020; 25, 2912.
RA Ward, S Fawell, N Floc’h, V Flemington, D McKerrecher and PD Smith. Challenges and opportunities in cancer drug resistance. Chem. Rev. 2021; 121, 3297-351.
NN Chen, XD Ma, Z Miao, XM Zhang, BY Han, AA Almaamari, JM Huang, XY Chen, YJ Liu and SW Su. Doxorubicin resistance in breast cancer is mediated via the activation of FABP5/PPARγ and CaMKII signaling pathway. Front. Pharmacol. 2023; 14, 1150861.
DW Shen, LM Pouliot, MD Hall and MM Gottesman. Cisplatin resistance: A cellular self-defense mechanism resulting from multiple epigenetic and genetic changes. Pharmacol. Rev. 2012; 64, 706-21.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2024 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






