Genetic Characteristics of Periophthalmus chrysospilos through COI Gene Analysis in Coastal Provinces of the Mekong Delta
DOI:
https://doi.org/10.48048/tis.2025.9996Keywords:
COI gene sequence, Conservation genetics, Genetic diversity, Mekong Delta, Mudskipper, Periophthalmus chrysospilos, PhylogeneticsAbstract
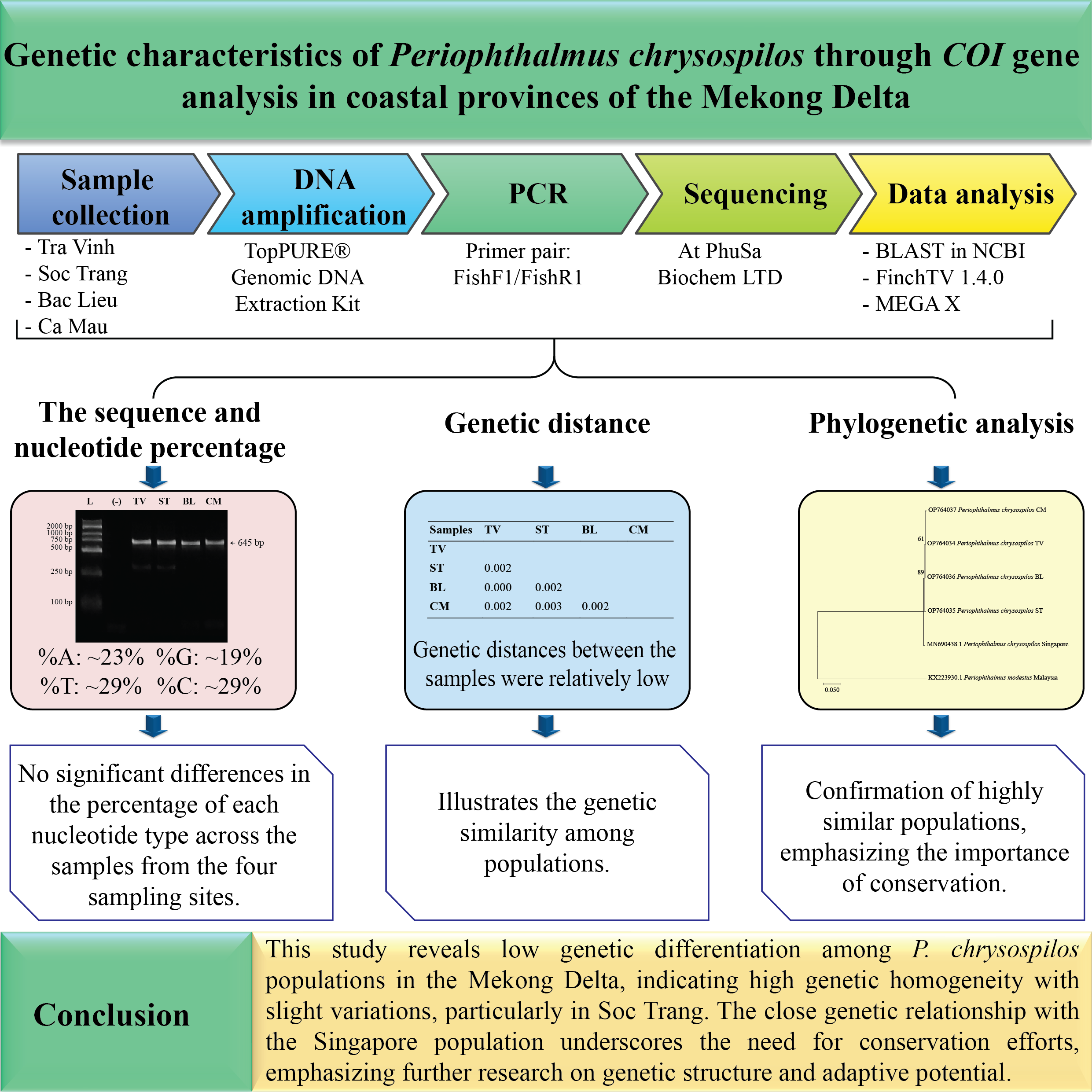
Periophthalmus chrysospilos is an amphibious gobiid species inhabiting intertidal mudflats and mangrove ecosystems across Southeast Asia, with the Mekong Delta as a critical habitat. This study investigates the genetic diversity of P. chrysospilos populations across 4 coastal provinces of the Mekong Delta - Tra Vinh, Soc Trang, Bac Lieu, and Ca Mau - through mitochondrial cytochrome c oxidase subunit I (COI) gene sequence analysis. Genomic DNA was extracted, amplified, and sequenced to assess nucleotide composition, genetic distances, and phylogenetic relationships. The analyzed COI gene sequences were 645 base pairs in length. The results demonstrated a high degree of sequence similarity (99.21 - 99.84 %) between P. chrysospilos populations in the Mekong Delta and those from Singapore. Nucleotide composition analysis revealed a higher proportion of adenine-thymine (AT) content than guanine-cytosine (GC). Furthermore, genetic distances among samples were minimal (0.00 - 0.03), suggesting low levels of genetic differentiation. Phylogenetic analysis clustered all sampled individuals into a single clade, confirming their classification as a single species. These findings indicate limited genetic divergence among populations and highlight the necessity for further investigations into environmental and anthropogenic factors affecting genetic structure. This study contributes to understanding P. chrysospilos genetic diversity and provides insights relevant to conservation and aquaculture management strategies.
HIGHLIGHTS
- Effect of different explant fragmentation and different concentrations of 2,4 Dichlorophenoxyacetic acid (2,4-D) on callus induction of mihanovichii LB2178 Agua Dulce.
- Quantification of anthocyanins from callus of mihanovichii LB2178 Agua Dulce.
- Genetic stability of the mihanovichii LB2178 Agua Dulce using SSR markers.
GRAPHICAL ABSTRACT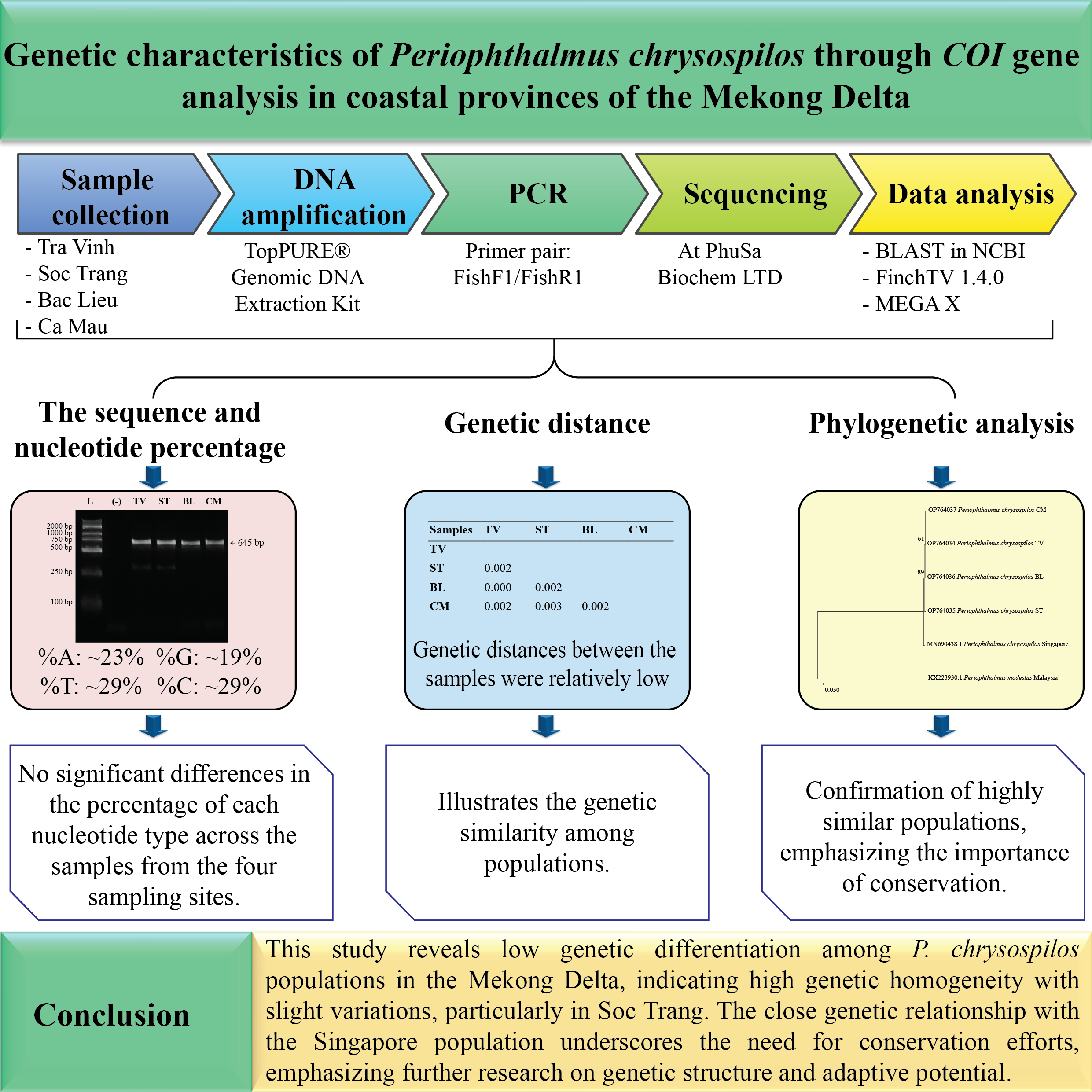
Downloads
References
RVD Laan, WN Eschmeyer and R Fricke. Family-group names of recent fishes. Zootaxa 2014; 3882(1), 1-230.
DD Tran, K Shibukawa, TP Nguyen, PH Ha, XL Tran, VH Mai and K Utsugi. Fishes of Mekong Delta, Vietnam. Can Tho University Publisher, Can Tho, Vietnam, 2013.
QM Dinh and THD Nguyen. Burrow behaviour, structure and utilization of the amphibious mudskipper Periophthalmus chrysospilos Bleeker, 1853 in the Mekong Delta. Saudi Journal of Biological Sciences 2023; 30(2), 103525.
DA Clayton. Mudskippers. Oceanography and Marine Biology: An Annual Review 1993; 31, 507-577.
G Polgar and G Crosa. Multivariate characterisation of the habitats of 7 species of Malayan mudskippers (Gobiidae: Oxudercinae). Marine Biology 2009; 156(7), 1475-1486.
EO Murdy. A taxonomic revision and cladistic analysis of the oxudercine gobies (Gobiidae, Oxudercinae). Australian Museum Journal 1989; 11, 1-93.
M Kottelat. The fishes of the inland waters of Southeast Asia: A catalogue and core bibliography of the fishes known to occur in freshwaters, mangroves and estuaries (Raffles Bulletin of Zoology). Raffles Bulletin of Zoology, Singapore, 2013.
Intergovernmental Panel on Climate Change. Climate change 2021: The physical science basis. Contribution of working group I to the sixth assessment report of the intergovernmental panel on climate change. Intergovernmental Panel on Climate Change, Geneva, Switzerland, 2021.
PDN Hebert, A Cywinska, SL Ball and JR Dewaard. Biological identifications through DNA barcodes. Proceedings of the Royal Society of London. Series B: Biological Sciences 2003; 270(1512), 313-321.
C Saccone, C De Giorgi, C Gissi, G Pesole and A Reyes. Evolutionary genomics in Metazoa: The mitochondrial DNA as a model system. Gene 1999; 238(1), 195-209.
V Sachithanandam, PM Mohan, N Muruganandam, I Chaaithanya, P Dhivya and R Baskaran. DNA barcoding, phylogenetic study of Epinephelus spp. from Andaman coastal region, India. Indian Journal of Geo-Marine Sciences 2012; 42(3), 203-211.
NM Linh, NV Quan, PV Chien, DH Ly, DV Nhan and DT Len. DNA barcoding application of mitochondrial COI gene to identify some fish of family Gobiidae in Vietnam. Vietnam Journal of Marine Science and Technology 2018; 18(4), 443-451.
TY Duong, K Nguyen, SN Bui, VT Nguyen, BL Nguyen and DD Tran. DNA barcodes and morphological characteristics of Pangasius krempfi, P. mekongensis and P. elongatus. Vietnam Journal of Biotechnology 2016; 14(1), 29-37.
SR Zhu, JJ Fu, Q Wang and JL Li. Identification of Channa species using the partial cytochrome c oxidase subunit I (COI) gene as a DNA barcoding marker. Biochemical Systematics and Ecology 2013; 51, 117-122.
T Arai and H Taha. Contrasting patterns of genetic population structure in tropical freshwater eels of genus Anguilla in the Indo-Pacific. Heliyon 2021; 7(5), e07097.
C Zhang, H Liu, X Huang, Z Yuan, S Zhang, S Xu, J Liu, Y Wang, D Wang and J Hu. Comparative analysis of the systematics and evolution of the Pampus genus of fish (Perciformes: Stromateidae) based on osteology, population genetics and complete mitogenomes. Animals 2024; 14(5), 814.
RD Ward, TS Zemlak, BH Innes, PR Last and PD Hebert. DNA barcoding Australia’s fish species. Philosophical Transactions of the Royal Society B: Biological Sciences 2005; 360(1462), 1847-1857.
QM Dinh, THD Nguyen, TTK Nguyen, TTH Lam, NT Truong and DD Tran. Population biological traits of Periophthalmus chrysospilos Bleeker, 1853 in the Vietnamese Mekong Delta. PeerJ 2022; 10, e13289.
QM Dinh, THD Nguyen, TTH Lam, NT Truong, TTK Nguyen and Z Jaafar. Reproduction ecology of an emerging fishery resource, the amphibious mudskipper Periophthalmus chrysospilos, in the Mekong Delta. Ecology and Evolution 2022; 12(1), e8507.
QM Dinh, THD Nguyen, NT Truong, LT Tran and TTK Nguyen. Morphometrics, growth pattern and condition factor of Periophthalmus chrysospilos Bleeker, 1853 (Gobiiformes: Oxudercidae) living in the Mekong Delta. The Egyptian Journal of Aquatic Research 2022; 48(2), 157-161.
TTH Lam, QM Dinh and THD Nguyen. Otolith morphometry and its role in determining the growth of Periophthalmus chrysospilos distributed in some coastal provinces in the Mekong Delta. Veterinary Integrative Sciences 2024; 22(1), 299-313.
G Polgar, A Sacchetti and P Galli. Differentiation and adaptive radiation of amphibious gobies (Gobiidae: Oxudercinae) in semi‐terrestrial habitats. Journal of Fish Biology 2010; 77(7), 1645-1664.
QM Dinh. Aspects of reproductive biology of the red goby Trypauchen vagina (Gobiidae) from the Mekong Delta. Journal of Applied Ichthyology 2018; 34(1), 103-110.
F Sanger, S Nicklen and AR Coulson. DNA sequencing with chain-terminating inhibitors. Proceedings of the National academy of Sciences 1977; 74(12), 5463-5467.
S Kumar, G Stecher and K Tamura. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution 2016; 33(7), 1870-1874.
D Posada and TR Buckley. Model selection and model averaging in phylogenetics: Advantages of Akaike information criterion and Bayesian approaches over likelihood ratio tests. Systematic Biology 2004; 53(5), 793-808.
X Bingpeng, L Heshan, Z Zhilan, W Chunguang, W Yanguo and W Jianjun. DNA barcoding for identification of fish species in the Taiwan Strait. PLoS One 2018; 13(6), e0198109.
JB Zhang and R Hanner. DNA barcoding is a useful tool for the identification of marine fishes from Japan. Biochemical Systematics and Ecology 2011; 39(1), 31-42.
M Chakraborty and SK Ghosh. An assessment of the DNA barcodes of Indian freshwater fishes. Gene 2014; 537(1), 20-28.
A Karahan, J Douek, G Paz, N Stern, AE Kideys, L Shaish, M Goren and B Rinkevich. Employing DNA barcoding as taxonomy and conservation tools for fish species censuses at the southeastern Mediterranean, a hot-spot area for biological invasion. Journal for Nature Conservation 2017; 36, 1-9.
XQ Li and D Du. Variation, evolution, and correlation analysis of C + G content and genome or chromosome size in different kingdoms and phyla. PLoS One 2014; 9(2), e88339.
N Hubert, R Hanner, E Holm, NE Mandrak, E Taylor, M Burridge, D Watkinson, P Dumont, A Curry and P Bentzen. Identifying Canadian freshwater fishes through DNA barcodes. PLoS One 2008; 3(6), e2490.
A Lara, JLPD León, R Rodríguez, D Casane, G Côté, L Bernatchez and E García‐Machado. DNA barcoding of Cuban freshwater fishes: Evidence for cryptic species and taxonomic conflicts. Molecular Ecology Resources 2010; 10(3), 421-430.
MP Tan, HM Gan, MH Nabilsyafiq, AG Mazlan, TNAM Jaafar, MN Siti Azizah, M Danish-Daniel and YY Sung. Genetic diversity of the Pearse’s mudskipper Periophthalmus novemradiatus (Perciformes: Gobiidae) and characterization of its complete mitochondrial genome. Thalassas: An International Journal of Marine Sciences 2020; 36, 103-113.
JG Prunier, M Chevalier, A Raffard, G Loot, N Poulet and S Blanchet. Genetic erosion reduces biomass temporal stability in wild fish populations. Nature Communications 2023; 14(1), 4362.
AC Bockelmann, TBH Reusch, R Bijlsma and JP Bakker. Habitat differentiation vs. isolation‐by‐distance: The genetic population structure of Elymus athericus in European salt marshes. Molecular Ecology 2003; 12(2), 505-515.
AS Martinez, JR Willoughby and MR Christie. Genetic diversity in fishes is influenced by habitat type and life‐history variation. Ecology and Evolution 2018; 8(23), 12022-12031.
G Polgar. Species-area relationship and potential role as a biomonitor of mangrove communities of Malayan mudskippers. Wetlands Ecology and Management 2009; 17(2), 157-164.
KN Shen, BW Jamandre, CC Hsu, WN Tzeng and JD Durand. Plio-Pleistocene sea level and temperature fluctuations in the northwestern Pacific promoted speciation in the globally-distributed flathead mullet Mugil cephalus. BMC Evolutionary Biology 2011; 11(83), 1-17.
G Tesfaye, M Curto, P Meulenbroek, GK Englmaier, PD Tibihika, E Alemayehu, A Getahun and H Meimberg. Genetic diversity of Nile tilapia (Oreochromis niloticus) populations in Ethiopia: Insights from nuclear DNA microsatellites and implications for conservation. BMC Ecology and Evolution 2021; 21, 1-14.
W Zhu, J Fu, M Luo, L Wang, P Wang, Q Liu and Z Dong. Genetic diversity and population structure of bighead carp (Hypophthalmichthys nobilis) from the middle and lower reaches of the Yangtze River revealed using microsatellite markers. Aquaculture Reports 2022; 27, 101377.
CH Faunce and JE Serafy. Mangroves as fish habitat: 50 years of field studies. Marine Ecology Progress Series 2006; 318, 1-18.
PL Munday, RR Warner, K Monro, JM Pandolfi and DJ Marshall. Predicting evolutionary responses to climate change in the sea. Ecology Letters 2013; 16(12), 1488-1500.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






