Hybrid Cassava Identification from Morphometric Analysis to Deep Convolutional Neural Networks and Confirmation Strategies
DOI:
https://doi.org/10.48048/tis.2025.9475Keywords:
Cassava, CNN, Deep learning, Fine-tuning, Hybrid identification, LIME, PCAAbstract
The correct identification of cassava varieties is critical for crop management, such as for developing high-value products or against agricultural pests. In this study, plant characteristic regions used for classification were verified by principal component analysis (PCA) techniques. A deep learning method was applied using well-known pretrained models to identify hybrid cassava through image classification. The models employed—ResNet-18, VGG-16, AlexNet, and GoogLeNet—yielded impressive accuracies in three-fold cross-validation experiments, achieving 100, 99.06, 99.06, and 98.59 % averaged accuracy, respectively. The fine-tuned ResNet-18 model had the highest accuracy and was selected for identifying hybrid cassava. A confusion matrix revealed 3 misidentifications. Cultivar variety (cv) R72 was mistakenly classified as R5 in both the 1st and 2nd folds and as R1 in the 2nd fold. Additionally, we utilized Local Interpretable Model-agnostic Explanations (LIME) to ensure that our models offered insightful explanations for their decision-making processes. The outcomes from Principal Component Analysis (PCA) and Local Interpretable Model-agnostic Explanations (LIME) exhibited close resemblance, particularly within the region encompassing the stem, branch, petiole, and stipule of cassava. These findings were leveraged for the identification of the 3 distinct cultivated cassava varieties. The results demonstrated the efficacy of deep learning as a potent technique for discerning hybrid cassava varieties, presenting promising prospects for its practical deployment in on-site testing and widespread adoption due to its time-saving capabilities.
HIGHLIGHTS
A deep learning method was applied to identify cassava with similar genetic varieties through image classification. High performance was achieved using a fine-tuned ResNet-18 model achieving 100 % accuracy for identifying hybrid cassava. The LIME technique was employed to generate interpretable explanations for individual predictions.
GRAPHICAL ABSTRACT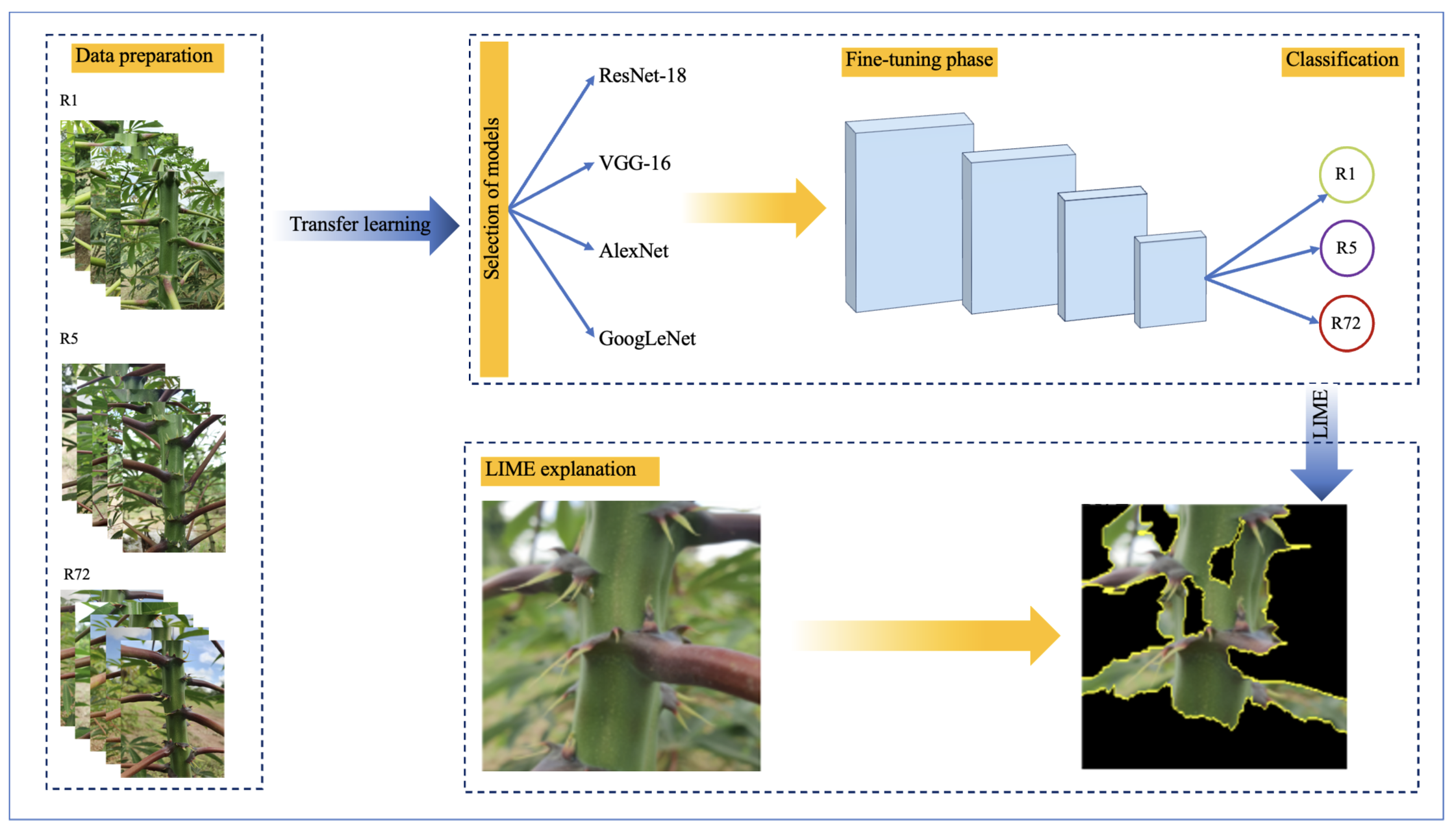
Downloads
References
A Bellotti, BVH Campo and G Hyman. Cassava production and pest management: Present and potential threats in a changing environment. Tropical Plant Biology 2012; 5, 39-72.
P Kongsil, H Ceballos, W Siriwan, S Vuttipongchaikij, P Kittipadakul, C Phumichai, W Wannarat, W Kositratana, V Vchukit, E Sarobol and C Rojanaridpiched. Cassava breeding and cultivation challenges in thailand: Past, present, and future perspectives. Plants 2024; 13(14), 1899.
EJD Oliveira, SASD Oliveira, SAV Boas, CS Hohenfeld and VDS Santos. Selection of cassava accessions with multiple resistance to pathogens associated with root rot disease. Euphytica 2017; 213, 185.
P Changtor, W Jaroenpol, K Buddhachat, W Wattanachaiyingcharoen and N Yimtragool. Rapid detection of Sclerotium rolfsii causing dry stem and root rot disease in cassava by recombinase polymerase amplification technique (RPA) combined with CRISPR/Cas12a. Crop Protection 2023; 172, 106340.
REC Mba, P Stephenson, K Edwards, S Melzer, J Nkumbira, U Gullberg, K Apel, M Gale, J Tohme and M Fregene. Simple sequence repeat (SSR) markers survey of the cassava (Manihot esculenta Crantz) genome: Towards an SSR-based molecular genetic map of cassava. Theoretical and Applied Genetics 2001; 102, 21-31.
J Adjebeng-Danquah, J Manu-Aduening, IK Asante, RY Agyare, V Gracen and SK Offei. Genetic diversity and population structure analysis of Ghanaian and exotic cassava accessions using simple sequence repeat (SSR) markers. Heliyon 2020; 6(1), e03154.
D Nadjiam, P Sarr, M Naitormbaide, M Mbailao and A Guisse. Agro-morphological characterization of cassava (Manihot esculenta Crantz) cultivars from chad. Agricultural Sciences 2016; 7(7), 479-492.
Z Li, R Guo, M Li, Y Chen and G Li. A review of computer vision technologies for plant phenotyping. Computers and Electronics in Agriculture 2020; 176, 105672.
K Suzuki. Overview of deep learning in medical imaging. Radiological Physics and Technology 2017; 10(3), 257-273.
Z Zhao, P Zheng, S Xu and X Wu. Object detection with deep learning: A review. IEEE Transactions on Neural Networks and Learning Systems 2019; 30(11), 3212-3232.
I Adjabi, A Ouahabi, A Benzaoui and A Taleb-Ahmed. Past, present, and future of face Recognition: A review. Electronics 2020; 9(8), 1188.
EC Too, L Yujian, S Njuki and L Yingchun. A comparative study of fine-tuning deep learning models for plant disease identification. Computers and Electronics in Agriculture 2019; 161, 272-279.
M Torky, G Dahy and AE Hassanien. Recognizing sounds of Red Palm Weevils (RPW) based on the VGGish model: Transfer learning methodology. Computers and Electronics in Agriculture 2023; 212, 108079.
X Jin, J Xiong, Y Rao, T Zhang, W Ba, S Gu, X Zhang and J Lu. TranNas-NirCR: A method for improving the diagnosis of asymptomatic wheat scab with transfer learning and neural architecture search. Computers and Electronics in Agriculture 2023; 213, 108271.
I Khan, SS Sohail, DO Madsen and BK Khare. Deep transfer learning for fine-grained maize leaf disease classification. Journal of Agriculture and Food Research 2024; 16, 101148.
KK Saha, A Rahman, M Moniruzzaman, M Syduzzaman, MZ Uddin, MM Rahman, MA Ali, DFA Riza and MMH Olivar. Classification of starfruit maturity using smartphone-image and multivariate analysis. Journal of Agriculture and Food Research 2023; 11, 100473.
MM Ghazi, B Yanikoglu and E Aptoula. Plant identification using deep neural networks via optimization of transfer learning parameters. Neurocomputing 2017; 235, 228-235.
T Nguyen, TL Le, H Vu, H Nguyen and HV Sam. A combination of deep learning and hand-designed feature for plant identification based on leaf and flower images. Springer Nature, London, 2017.
P Jasitha, MR Dileep and M Divya. Venation based plant leaves classification using GoogLeNet and VGG. In: Proceedings of the 4th International Conference on Recent Trends on Electronics, Information, Communication & Technology (RTEICT), Bangalore, India. 2019.
O J V Ramirez, JECDl Cruz and WAM Machaca. Agroindustrial plant for the classification of hass avocados in real-time with ResNet-18 architecture. In: Proceedings of the 5th International Conference on Robotics and Automation Sciences (ICRAS), Wuhan, China. 2021.
SY Arafat, N Arshad and R Khan. Holistic based plant identification using deep learning. In: Proceedings of the 16th International Conference on Emerging Technologies (ICET), Islamabad, Pakistan. 2021.
MT Ribeiro, S Singh and C Guestrin. “Why should i trust you?”: Explaining the predictions of any classifier. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, California. 2016.
K Sharma, R Bhattacharjee, A Sartie and P Kumar. An improved method of DNA extraction from plants for pathogen detection and genotyping by polymerase chain reaction. African Journal of Biotechnology 2013; 12(15), 1894-1901.
A Paszke, S Gross, F Massa, A Lerer, J Bradbury, G Chanan, T Killeen, Z Lin, N Gimelshein, L Antiga, A Desmaison, A Köpf, E Yang, Z DeVito, M Raison, A Tejani, S Chilamkurthy, B Steiner, L Fang, ..., S Chintala. In: Proceedings of the 33rd International Conference on Neural Information Processing Systems, New York. 2019, p. 8026-8037.
K He, X Zhang, S Ren and J Sun. Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, Nevada. 2016.
A Krizhevsky, I Sutskever and GE Hinton. ImageNet classification with deep convolutional neural networks. Communications of the ACM 2017; 60(6), 84-90.
K Simonyan and A Zisserman. Very deep convolutional networks for large-scale image recognition. International Conference on Learning Representations, California, 2015.
C Szegedy, W Liu, Y Jia, P Sermanet, S Reed, D Anguelov, D Erhan, V Vanhoucke and A Rabinovich. Going deeper with convolutions. In: Proceedings of the IEEE conference on computer vision and pattern recognition 2014, Ohio. 2014.
SK O’Hair. Tropical root and tuber crop. Horticultural Reviews 1990; 12, 157-196.
S Sivan, K Arya, MN Sheela, BS Revathi, BSP Krishnan and SK Muthusamy. Genetic diversity analysis of Indian Cassava (Manihot esculenta Crantz) accessions using morphological and molecular markers. South African Journal of Botany 2023; 161, 347-357.
S Ngorian, J Kansup, B Ruangwised, V Chanroj, S Amawan and P Wongtiem, The study of genetic diversity of cassava (Manihot esculenta) using SSR markers. Thai Agricultural Research Journal 2019; 37(1), 2-13.
A Kaya, AS Keceli, C Catal, HY Yalic, H Temucin and B Tekinerdogan. Analysis of transfer learning for deep neural network-based plant classification models. Computers and Electronics in Agriculture 2019; 158, 20-29.
Y Chen, Y Huang, Z Zhang, Z Wang, B Liu, C Liu, C Huang, S Dong, X Pu, F Wan, X Qiao and W Qian. Plant image recognition with deep learning: A review. Computers and Electronics in Agriculture 2023; 212, 108072.
F Jia, EB Martin and A J Morris. Non-linear principal components analysis for process fault detection. Computers & Chemical Engineering 1998; 229(1), 851-854.
H Wang, G Li, Z Ma and X. Li. Image recognition of plant diseases based on principal component analysis and neural networks. In: Proceedings of the 8th International Conference on Natural Computation, Sichuan. 2012.
Y Sun, Y Liu, G Wang and H Zhang. Deep learning for plant identification in natural environment. Computational Intelligence and Neuroscience 2017; 2017(1), 7361042.
GL Grinblat, LC Uzal, MG Larese and PM Granitto. Deep learning for plant identification using vein morphological patterns. Computers and Electronics in Agriculture 2016; 127, 418-424.
MS Hossain, M Al-Hammadi and G Muhammad. Automatic fruit classification using deep learning for industrial applications. IEEE Transactions on Industrial Informatics 2019; 15(2), 1027-1034.
A Rocha, DC Hauagge, J Wainer and S Goldenstein. Automatic fruit and vegetable classification from images. Computers and Electronics in Agriculture 2010; 70(1), 96-104.
M Togaçar, N Muzoglu, B Ergen, BSB Yarman and AM Halefoglu. Detection of COVID-19 findings by the local interpretable model-agnostic explanations method of types-based activations extracted from CNNs. Biomedical Signal Processing and Control 2022; 71, 103128.
X Zhao, L Zhang, G Zhu, C Cheng, J He, S Traore and VP Singh. Exploring interpretable and non-interpretable machine learning models for estimating winter wheat evapotranspiration using particle swarm optimization with limited climatic data. Computers and Electronics in Agriculture 2023; 212, 108140.

Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






