Diagnostic and Prognostic Potential of Let-7 miRNAs in Breast Cancer: A Meta-Analysis
DOI:
https://doi.org/10.48048/tis.2025.9407Keywords:
Let-7 miRNAs, Breast cancer, Diagnosis, Prognosis, Meta-analysis, miR-202, BiomarkerAbstract
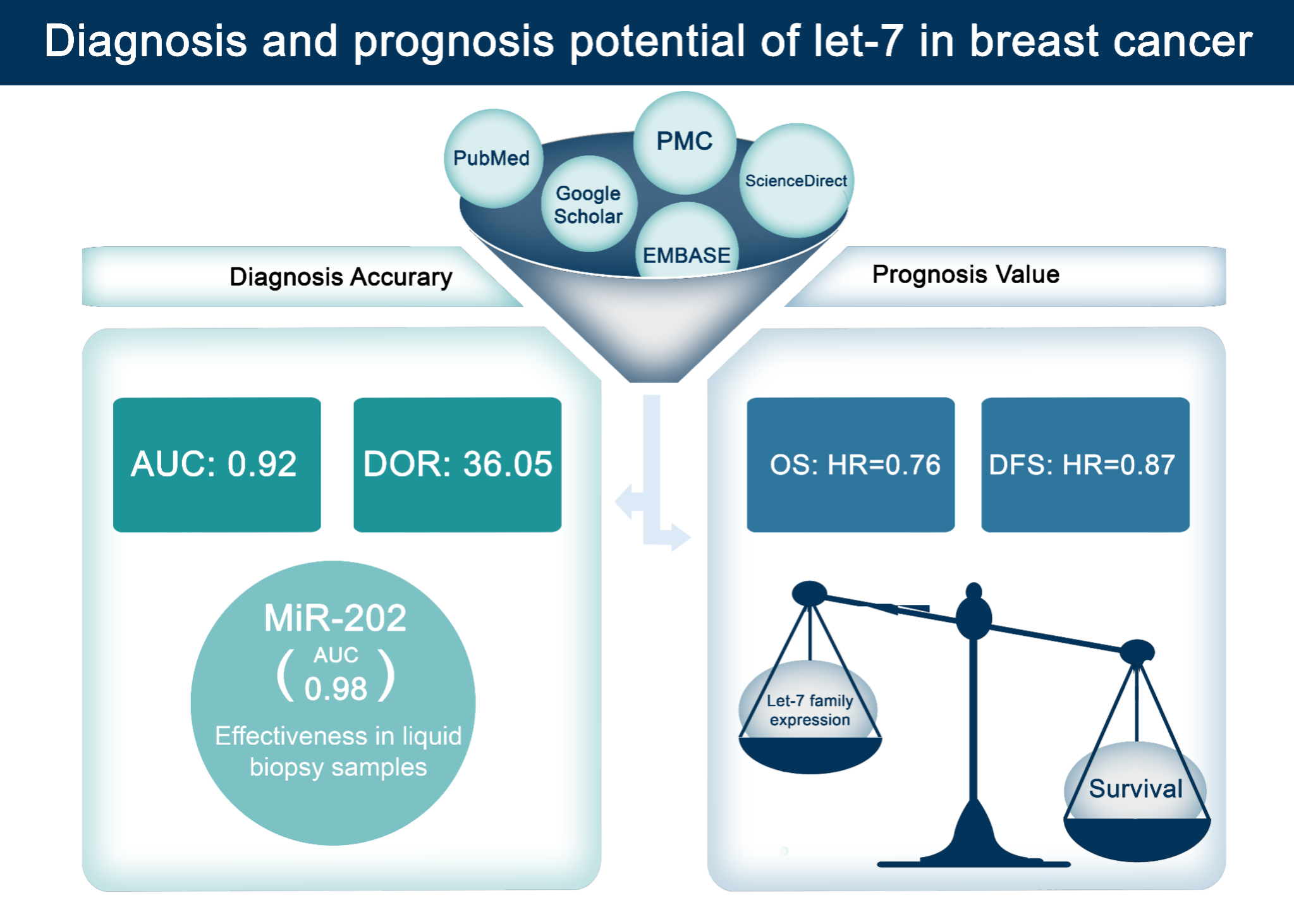
MicroRNAs (miRNAs), particularly those in the let-7 family, show significant promise as diagnostic and prognostic biomarkers in breast cancer (BC). This meta-analysis systematically evaluates their diagnostic accuracy and prognostic value, addressing inconsistencies across previous studies. We searched PubMed, PMC, ScienceDirect and EMBASE databases for studies assessing let-7 miRNAs in BC. A total of 23 studies involving 8,249 participants were included. For diagnosis, the let-7 miRNA family demonstrated a pooled sensitivity of 0.87 (95 % CI: 0.80 - 0.92) and specificity of 0.76 (95 % CI: 0.62 - 0.86), with a diagnostic odds ratio (DOR) of 36.05 (95 % CI: 17.26 - 75.26) and an area under the curve (AUC) of 0.92 (95 % CI: 0.86 - 0.98). Subgroup analysis revealed that miR-202 in liquid biopsy samples from Asian populations showed the highest diagnostic accuracy (AUC: 0.98). For prognosis, low expression of let-7 miRNAs correlated with poorer survival outcomes, with a pooled hazard ratio (HR) for overall survival (OS) of 0.76 (95 % CI: 0.59 - 0.99). Notably, let-7a and let-7c were associated with aggressive BC subtypes. These findings underscore the let-7 family’s potential for BC diagnosis and prognosis, particularly miR-202 for non-invasive detection. However, high heterogeneity across studies was noted due to variations in methods and populations. Future research should validate these findings in diverse populations and standardize detection methods to explore their clinical utility.
HIGHLIGHTS
- Let-7 miRNAs demonstrate high diagnostic accuracy for breast cancer, with a sensitivity (87 %), specificity (76 %), and an AUC of 0.92.
- miR-202 shows the highest diagnostic potential, particularly in liquid biopsy samples from Asian populations (AUC: 0.98).
- Let-7a and let-7c are associated with poorer survival outcomes, highlighting their prognostic value in aggressive breast cancer subtypes (HR: 0.76 for OS).
- Meta-regression identifies RT-PCR chemistries (TaqMan > SYBR Green) as a major source of heterogeneity in diagnostic performance.
- These findings support let-7 miRNAs as promising non-invasive biomarkers for early detection and the personalized management of breast cancer.
GRAPHICAL ABSTRACT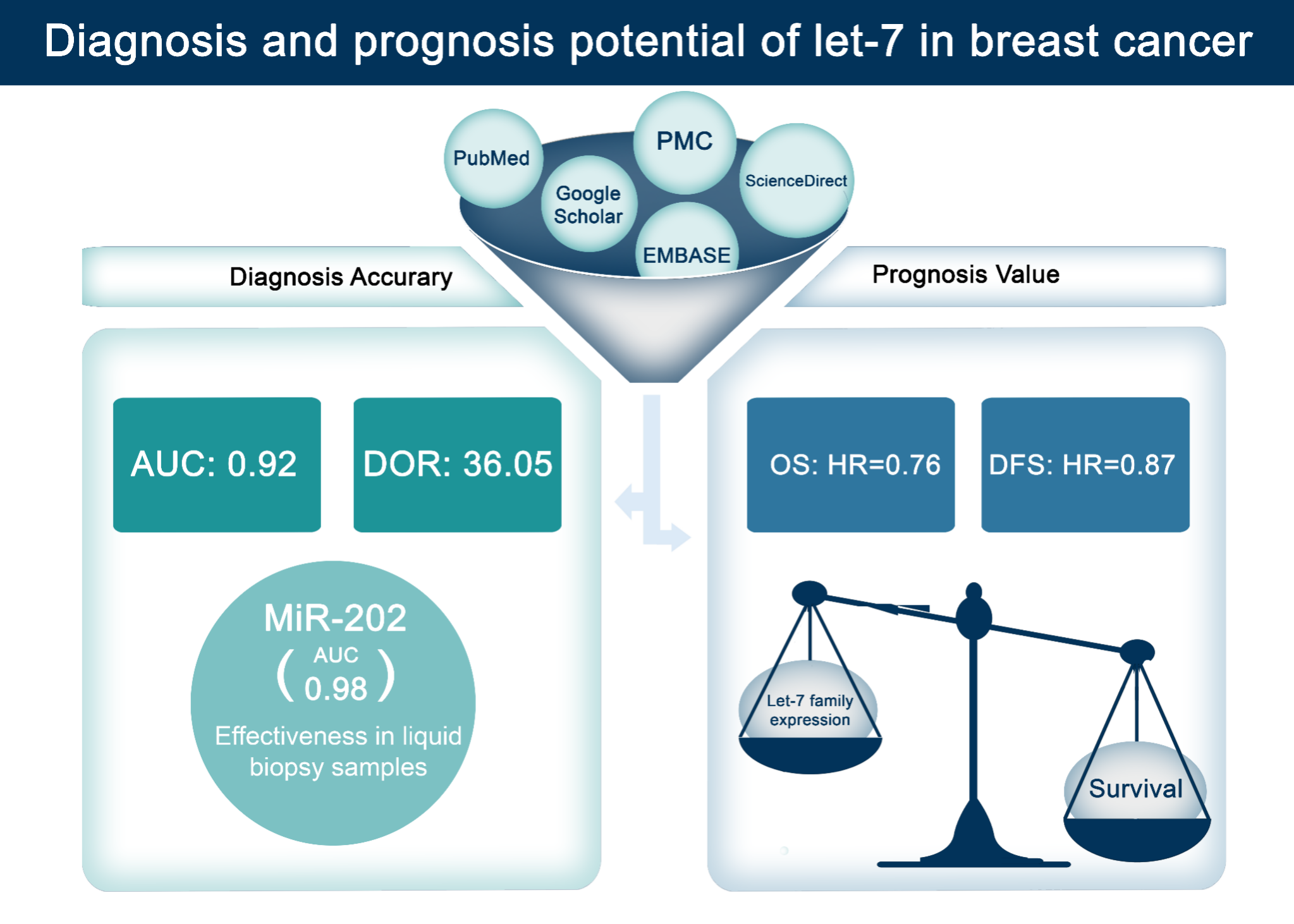
Downloads
References
F Bray, M Laversanne, H Sung, J Ferlay, R L Siegel, I Soerjomataram and A Jemal. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians 2024; 74(3), 205-313.
S Eslamizadeh, M Heidari, S Agah, E Faghihloo, H Ghazi, A Mirzaei and A Akbari. The role of microRNA signature as diagnostic biomarkers in different clinical stages of colorectal cancer. Cell Journal (Yakhteh) 2018; 20(2), 220-230.
W Wang and Y Luo. MicroRNAs in breast cancer: Oncogene and tumor suppressors with clinical potential. Journal of Zhejiang University-Science B 2015; 16(1), 18-31.
CK Thammaiah and S Jayaram. Role of let-7 family microRNA in breast cancer. Non-coding RNA Research 2016; 1(1), 77-82.
X Song, Y Liang, Y Sang, Y Li, H Zhang, B Chen, L Du, Y Liu, L Wang, W Zhao, T Ma, C Wang and Q Yang. CircHMCU promotes the proliferation and metastasis of breast cancer by sponging the let-7 family. Molecular Therapy-Nucleic Acids 2020; 20, 518-533.
G Mu, H Liu, F Zhou, X Xu, H Jiang, Y Wang and Y Qu. Correlation of overexpression of HMGA1 and HMGA2 with poor tumor differentiation, invasion, and proliferation associated with let-7 down-regulation in retinoblastomas. Human Pathology 2010; 41(4), 493-502.
L Guo, X Cheng, H Chen, C Chen, S Xie, M Zhao, D Liu, Q Deng, Y Liu, X Wang, X Chen, J Wang, Z Yin, S Qi, J Gao, Y Ma, N Guo and M Shi. Induction of breast cancer stem cells by M1 macrophages through Lin-28B-let-7-HMGA2 axis. Cancer Letters 2019; 452, 213-225.
T Gunel, B Dogan, E Gumusoglu, MK Hosseini, S Topuz and K Aydinli. Regulation of HMGA2 and KRAS genes in epithelial ovarian cancer by miRNA hsa-let-7d-3p. Journal of Cancer Research and Therapeutics 2019; 15(6), 1321-1327.
TT Liao, WH Hsu, CH Ho, WL Hwang, HY Lan, T Lo, CC Chang, SK Tai and MH Yang. Let-7 modulates chromatin configuration and target gene repression through regulation of the ARID3B complex. Cell Reports 2016; 14(3), 520-533.
X Wang, L Cao, Y Wang, X Wang, N Liu and Y You. Regulation of let-7 and its target oncogenes. Oncology Letters 2012; 3, 955-960.
U Gezer, S Keskin, A Iğci, M Tükenmez, D Tiryakioğlu, M Cetinkaya, R Dişci, N Dalay and Y Eralp. Abundant circulating microRNAs in breast cancer patients fluctuate considerably during neoadjuvant chemotherapy. Oncology Letters 2014; 8(2), 845-848.
M Li, X Zou, T Xia, T Wang, P Liu, X Zhou, S Wang and W Zhu. A 5‐miRNA panel in plasma was identified for breast cancer diagnosis. Cancer Medicine 2019; 8(16), 7006-7017.
MM Marques, AF Evangelista, T Macedo, RAC Vieira, CS Neto, RM Reis and AL Carvalho. P041 Serum and tissue expression of tumor suppressors miR-195 and let-7A in breast cancer. The Breast 2015; 24, S40.
A Qattan, H Intabli, W Alkhayal, C Eltabache, T Tweigieri and SB Amer. Robust expression of tumor suppressor miRNA’s let-7 and miR-195 detected in plasma of Saudi female breast cancer patients. BMC Cancer 2017; 17, 1-10.
EA Ahmed, P Rajendran and H Scherthan. The microRNA-202 is a diagnostic biomarker and a potential tumor suppressor. International Journal of Molecular Sciences 2022; 23(11), 5870.
HM Heneghan, N Miller, AJ Lowery, KJ Sweeney, J Newell and MJ Kerin. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Annals of Surgery 2010; 251(3), 499-505.
J Kim, S Park, D Hwang, SI Kim and H Lee. Diagnostic value of circulating miR-202 in early-stage breast cancer in South Korea. Medicina (B Aires) 2020; 56(7), 340.
S Gao, C Cao, Q Dai, J Chen and J Tu. MiR-202 acts as a potential tumor suppressor in breast cancer. Oncology Letters 2018; 16(1), 1155-1162.
PF Whiting, AWS Rutjes, ME Westwood, S Mallett, JJ Deeks, JB Reitsma, MMG Leeflang, JAC Sterne, PMM Bossuyt and QUADAS-2 Group. QUADAS-2: A revised tool for the quality assessment of diagnostic accuracy studies. Annals of Internal Medicine 2011; 155(8), 529-536.
WJA Grooten, E Tseli, BO Äng, K Boersma, BM Stålnacke, B Gerdle and P Enthoven. Elaborating on the assessment of the risk of bias in prognostic studies in pain rehabilitation using QUIPS aspects of interrater agreement. Diagnostic and Prognostic Research 2019; 3, 1-11.
AM Šimundić. Measures of diagnostic accuracy: Basic definitions. Journal of the International Federation of Clinical Chemistry and Laboratory Medicine 2009; 19, 203.
CM Jones and T Athanasiou. Summary receiver operating characteristic curve analysis techniques in the evaluation of diagnostic tests. The Annals of Thoracic Surgery 2005; 79, 16-20.
JPT Higgins, SG Thompson, JJ Deeks and DG Altman. Measuring inconsistency in meta-analyses. Thebmj 2003; 327, 557-560.
D Jackson, IR White and SG Thompson. Extending DerSimonian and Laird’s methodology to perform multivariate random effects meta‐analyses. Statistics in Medicine 2010; 29(12), 1282-1297.
SR Shim, SJ Kim and J Lee. Diagnostic test accuracy: Application and practice using R software. Epidemiol Health 2019; 41, e2019007.
CH Lee, WH Kuo, CC Lin, YJ Oyang, HC Huang and HF Juan. MicroRNA-regulated protein-protein interaction networks and their functions in breast cancer. International Journal of Molecular Sciences 2013; 14(6), 11560-11606.
SA Joosse, V Müller, B Steinbach, K Pantel and H Schwarzenbach. Circulating cell-free cancer-testis MAGE-A RNA, BORIS RNA, let-7b and miR-202 in the blood of patients with breast cancer and benign breast diseases. British Journal of Cancer 2014; 111(5), 909-917.
ZQ Deng, JY Yin, Qin Tang, FQ Liu, J Qian, J Lin, R Shao, M Zhang and L He. Over-expression of miR-98 in FFPE tissues might serve as a valuable source for biomarker discovery in breast cancer patients. International Journal of Clinical and Experimental Pathology 2014; 7(3), 1166.
XX Li, SY Gao, PY Wang, X Zhou, YJ Li, Y Yu, YF Yan, HH Zhang, CJ Lv, HH Zhou and SY Xie. Reduced expression levels of let 7c in human breast cancer patients. Oncology Letter 2015; 9(3), 1207-1212.
S Huang, Q Luo, H Peng, J Li, M Zhao, J Wang, Y Gu, Y Li, P Yuan, G Zhao and C Huang. A panel of serum noncoding RNAs for the diagnosis and monitoring of response to therapy in patients with breast cancer. Medical Science Monitor: International Medical Journal of Experimental and Clinical Research 2018; 24, 2476.
AM Ibrahim, MM Said, AM Hilal, AM Medhat and IMA Elsalam. Candidate circulating microRNAs as potential diagnostic and predictive biomarkers for the monitoring of locally advanced breast cancer patients. Tumor Biology 2020; 42(10), 1-13.
H Aksan, BP Kundaktepe, U Sayili, M Velidedeoglu, G Simsek, S Koksal, R Gelisgen, I Yaylim and H Uzun. Circulating miR‐155, let‐7c, miR‐21, and PTEN levels in differential diagnosis and prognosis of idiopathic granulomatous mastitis and breast cancer. Biofactors 2020; 46(6), 955-962.
J Kim. Identification of MicroRNAs as diagnostic biomarkers for breast cancer based on the cancer genome atlas. Diagnostics 2021; 11(1), 107.
X Zou, T Xia, M Li, T Wang, P Liu, X Zhou, Z Huang and W Zhu. MicroRNA profiling in serum: Potential signatures for breast cancer diagnosis. Cancer Biomarkers 2021; 30(1), 41-53.
JL Quesne, J Jones, J Warren, SJ Dawson, HR Ali, H Bardwell, F Blows, P Pharoah and C Caldas. Biological and prognostic associations of miR‐205 and let‐7b in breast cancer revealed by in situ hybridization analysis of micro‐RNA expression in arrays of archival tumour tissue. The Journal of Pathology 2012; 227(3), 306-314.
S Volinia, M Galasso, ME Sana, TF Wise, J Palatini, K Huebner and CM Croce. Breast cancer signatures for invasiveness and prognosis defined by deep sequencing of microRNA. Proceedings of the National Academy of Science 2012; 109(8), 3024-3029.
K Jonsdottir, SR Janssen, FCD Rosa, E Gudlaugsson, I Skaland, JPA Baak and EAM Janssen. Validation of expression patterns for 9 miRNAs in 204 lymph-node negative breast cancers. PLoS One 2012; 7(11), e48692.
A Markou, GM Yousef, E Stathopoulos, V Georgoulias and E Lianidou. Prognostic significance of metastasis-related microRNAs in early breast cancer patients with a long follow-up. Clinical Chemistry 2014; 60(1), 197-205.
G Turashvili, ED Lightbody, K Tyryshkin, SK Sengupta, BE Elliott, Y Madarnas, A Ghaffari, A Day and CJB Nicol. Novel prognostic and predictive microRNA targets for triple-negative breast cancer. The FASEB Journal 2018; 32(11), 5937-5954.
Q Guo, R Wen, B Shao, Y Li, X Jin, H Deng, J Wu, F Su and F Yu. Combined let-7a and H19 signature: A prognostic index of progression-free survival in primary breast cancer patients. Journal of Breast Cancer 2018; 21(2), 142-149.
M Geng, JK Pan, Y Luo, L Tian and JH Wang. Prognostic values of expression of Let-7 family in breast cancer. Journal of Regional Anatomy and Operative Surgery 2019; 6, 1-5.
SA Fahim, MS Abdullah, NAE Sánchez, H Hassan, AM Ibrahim, SH Ahmed, G Shakir, MA Badawy, NI Zakhary, B Greve, ME Shinawi, M Götte and SA Ibrahim. Inflammatory breast carcinoma: Elevated microRNA miR-181b-5p and reduced miR-200b-3p, miR-200c-3p, and miR-203a-3p expression as potential biomarkers with diagnostic value. Biomolecules 2020; 10(7), 1059.
M Zaka, CW Sutton, Y Peng and S Konur. Model-based integration analysis revealed the presence of novel prognostic miRNA targets and important cancer driver genes in triple-negative breast cancers. Cancers 2020; 12(3), 632.
M Zhang, Z Li and X Liu. MiR-98-5p/IGF2 axis influence herceptin sensitivity through igf1r/her2 heterodimer formation and AKT/mTOR signal pathway in HER2 positive breast cancer. Asian Pacific Journal of Cancer Prevention 2021; 22 (11), 3693-3703.
P Fuso, M D Salvatore, C Santonocito, D Guarino, C Autilio, A Mulè, D Arciuolo, A Rinninella, F Mignone, M Ramundo, B Di Stefano, A Orlandi, E Capoluongo, N Nicolotti, G Franceschini, AM Sanchez, G Tortora, G Scambia, C Barone and A Cassano. Let-7a-5p, miR-100-5p, miR-101-3p, and miR-199a-3p hyperexpression as potential predictive biomarkers in early breast cancer patients. Journal of Personalized Medicine 2021; 11(8), 816.
JX Hing, CW Mok, PT Tan, SS Sudhakar, CM Seah, WP Lee and SM Tan. Clinical utility of tumour marker velocity of cancer antigen 15-3 (CA 15-3) and carcinoembryonic antigen (CEA) in breast cancer surveillance. The Breast 2020; 52, 95-101.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2024 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






