Analytical Validation of A 101 Germline miR-SNP Ion AmpliSeq Panel for Breast Cancer-Related Genetic Studies
DOI:
https://doi.org/10.48048/tis.2026.12823Keywords:
miRNA-SNP, Breast cancer, Ion AmpliSeq, MPS, Genotyping, ValidationAbstract
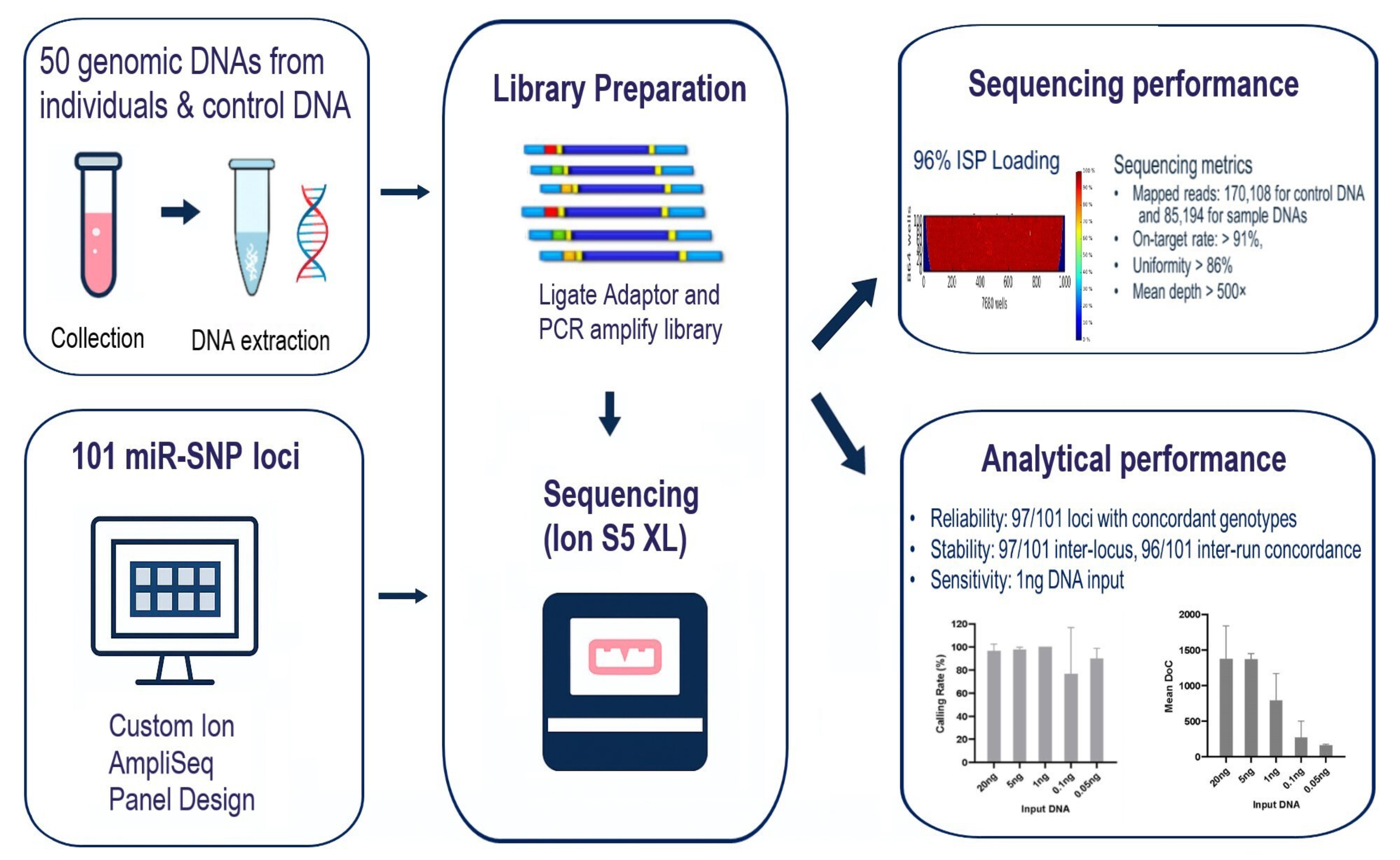
Massively parallel sequencing (MPS) technology enables simultaneous genotyping of many SNPs across multiple samples with high depth and accuracy. Such technology has become increasingly valuable for studies investigating genetic variants, including miRNA-related SNPs (miR-SNPs), that may contribute to cancer susceptibility. In this study, we designed a customized Ion AmpliSeq panel targeting 101 candidate breast cancer-associated miR-SNPs and evaluated its sequencing performance and analytical reliability on the Ion S5 XL system. Sequencing metrics, including depth of coverage (DoC), locus coverage balance (LCB), frequency of major allele reads (FMAR), and locus strand balance (LSB), were assessed across all loci. Across 50 DNA samples, the sequencing achieved a mean of 85,194 (95% CI: 79,904 - 101,503), with 91.44% (95% CI: 90.68 - 94.45) on-target reads, an average depth of 515.1× (95% CI: 488.9 - 592.9), and uniformity of 88.77% (95% CI: 88.76 - 89.64), indicating high-quality and consistent performance across libraries. Among 101 miR-SNPs, 97 loci yielded valid genotypes, while four loci resulted in no-calls due to low coverage. Genotypes obtained from the customized miR-SNP panel were fully concordant with those verified by Sanger sequencing. The panel demonstrated stable performance and high reproducibility, with consistent variant calling across replicate runs. Furthermore, reliable and complete genotyping profiles were obtained from as little as 1 ng of input DNA. Collectively, these results indicate that the customized miR-SNP MPS panel provides robust analytical performance and high accuracy, supporting its applicability for large-scale genetic and breast cancer association studies.
HIGHLIGHTS
- A 101 germline miR-SNP panel was designed using the Ion AmpliSeq platform.
- The panel targets polymorphisms in miRNA-encoding regions associated with breast cancer.
- Analytical validation demonstrated high coverage uniformity and calling accuracy.
- The panel achieved consistent performance across serial DNA dilutions and replicates.
- This validated assay supports large-scale studies of germline miR-SNPs in breast cancer risk.
GRAPHICAL ABSTRACT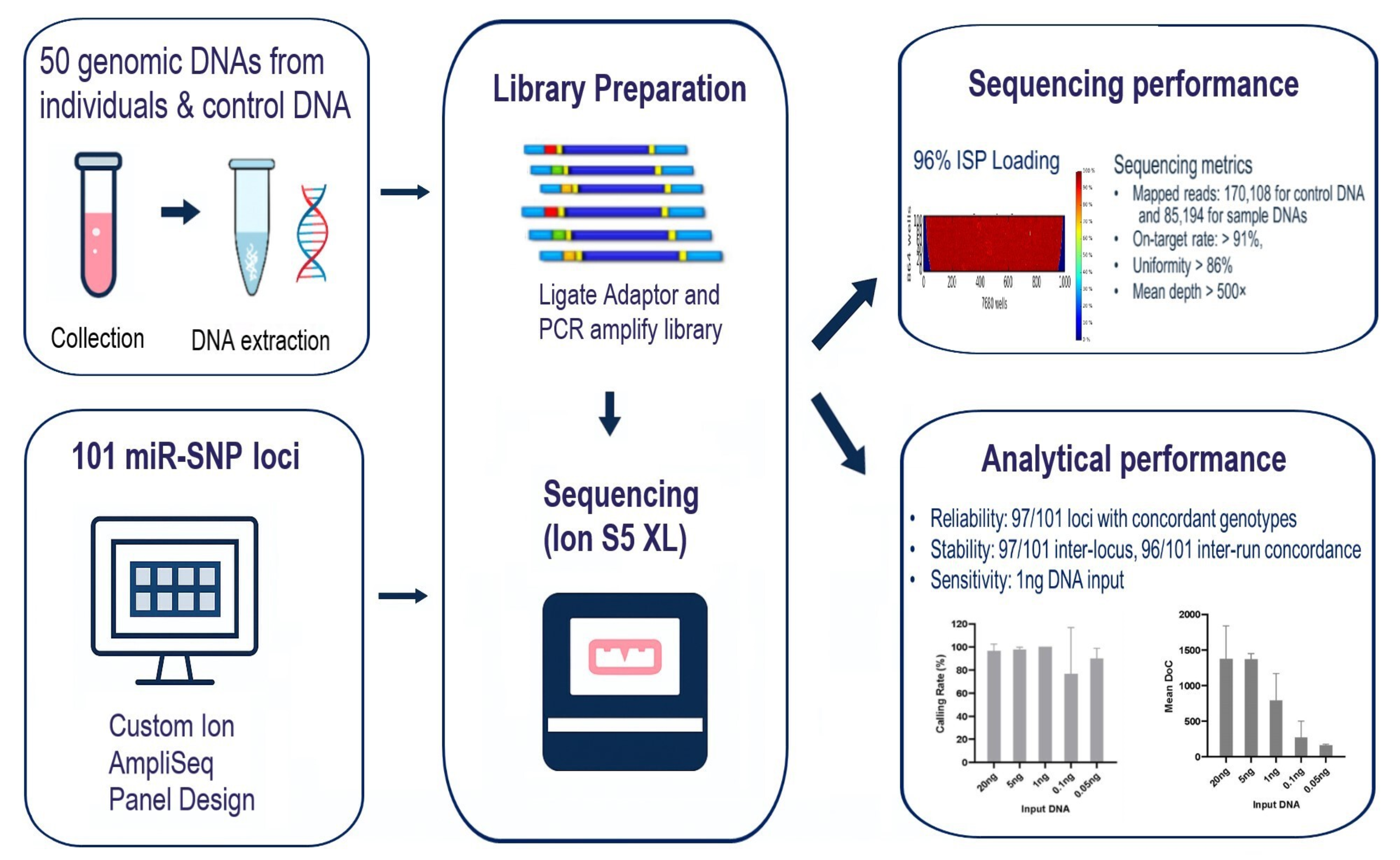
Downloads
References
S Ding and P Wang. The life of microRNAs: Biogenesis, function and decay in cancer. Biomolecules 2025; 15(10), 1393.
A Syeda, SSS Langden, C Munkhzul, M Lee and SJ Song. Regulatory mechanism of microRNA expression in cancer. International Journal of Molecular Sciences 2020; 21(5), 1723.
HY Loh, BP Norman, KS Lai, N Rahman, NBM Alitheen and MA Osman. The regulatory role of microRNAs in breast cancer. International Journal of Molecular Sciences 2019; 20(19), 4940.
W Jelski, S Okrasinska and B Mroczko. MicroRNAs as biomarkers of breast cancer. International Journal of Molecular Sciences 2025; 26(9), 4395.
S Chauhan, R Mathur and AK Jha. The impact of microRNA SNPs on breast cancer: Potential biomarkers for disease detection. Molecular Biotechnology 2024; 67(3), 845-861.
Y Chhichholiya, AK Suryan, P Suman, A Munshi and S Singh. SNPs in miRNAs and target sequences: Role in cancer and diabetes. Frontiers in Genetics 2021; 12, 793523.
Y Wang and GC Hon. Towards functional maps of non-coding variants in cancer. Frontiers in Genome Editing 2024; 6, 1481443.
T Arancibia, S Morales-Pison, E Maldonado and L Jara. Association between single-nucleotide polymorphisms in miRNA and breast cancer risk: An updated review. Biological Research 2021; 54(1), 26.
T Kongton, S Sanguansin, P Saelee, A Thongmee and W Pongstaporn. The association of pri-miR34 b/c gene polymorphism and clinicopathologic data in breast cancer patients. Asian Pacific Journal of Cancer Prevention 2024; 25(7), 2415-2420.
G Hina, N Khan, ST Muntaha, A Ullah, S Khan, A Alanzi, I Ali and T Chen. Association analysis of miRNA-146a and miRNA-196a polymorphism with breast cancer risk in Khyber Pakhtunkhwa, Pakistan: A preliminary study. Molecular Biology Reports 2024; 51(1), 827.
AM McInerney-Leo and EL Duncan. Massively parallel sequencing for rare genetic disorders: Potential and pitfalls. Frontiers in Endocrinology 2020; 11, 628946.
F Guo, J Yu, L Zhang and J Li. Massively parallel sequencing of forensic STRs and SNPs using the Illumina® ForenSeq™ DNA Signature Prep kit on the MiSeq FGx™ Forensic Genomics system. Forensic Science International: Genetics 2017; 31, 135-148.
CKY Ng, AM Schultheis, FC Bidard, B Weigelt and JS Reis-Filho. Breast cancer genomics from microarrays to massively parallel sequencing: Paradigms and new insights. Journal of the National Cancer Institute 2015; 107(5), djv015.
C Kraus, J Hoyer, G Vasileiou, M Wunderle, MP Lux, PA Fasching, M Krumbiegel, S Uebe, M Reuter, MW Beckmann and A Reis. Gene panel sequencing in familial breast/ovarian cancer patients identifies multiple novel mutations also in genes others than BRCA1/2. International Journal of Cancer 2017; 140(1), 95-102.
JR Bradford, Y Hey, T Yates, Y Li, SD Pepper and CJ Miller. A comparison of massively parallel nucleotide sequencing with oligonucleotide microarrays for global transcription profiling. BMC Genomics 2010; 11(1), 282.
LJ Semenuk, BA Kartolo, HE Feilotter, SM Lee, CA Savage, AH Boag, GC Digby and M Mates. Implementing next-generation sequencing process changes to increase capacity and improve timeliness of molecular biomarker profiling for lung cancer patients. The Journal of Applied Laboratory Medicine 2023; 9(2), 284-294.
S Shin, Y Kim, SC Oh, N Yu, ST Lee, JR Choi and KA Lee. Validation and optimization of the Ion Torrent S5 XL sequencer and Oncomine workflow for BRCA1 and BRCA2 genetic testing. Oncotarget 2017; 8(21), 34858-34866.
M Mehrotra, DY Duose, RR Singh, BA Barkoh, J Manekia, MA Harmon, KP Patel, MJ Routbort, LJ Medeiros, II Wistuba and R Luthra. Versatile ion S5XL sequencer for targeted next generation sequencing of solid tumors in a clinical laboratory. PLoS One 2017; 12(8), e0181968.
M Ivanov, K Laktionov, V Breder, P Chernenko, E Novikova, E Telysheva, S Musienko, A Baranova and V Mileyko. Towards standardization of next-generation sequencing of FFPE samples for clinical oncology: Intrinsic obstacles and possible solutions. Journal of Translational Medicine 2017; 15(1), 22.
T Huebner, M Steffens and C Scholl. Current status of the analytical validation of next generation sequencing applications for pharmacogenetic profiling. Molecular Biology Reports 2023; 50(11), 9587-9599.
LJ Jennings, ME Arcila, C Corless, S Kamel-Reid, IM Lubin, J Pfeifer, RL Temple-Smolkin, KV Voelkerding and MN Nikiforova. Guidelines for validation of next-generation sequencing-based oncology panels: A joint consensus recommendation of the Association for Molecular Pathology and College of American Pathologists. The Journal of Molecular Diagnostics 2017; 19(3), 341-365.
JT Bensen, CK Tse, SJ Nyante, JS Barnholtz-Sloan, SR Cole and RC Millikan. Association of germline microRNA SNPs in pre-miRNA flanking region and breast cancer risk and survival: The carolina breast cancer study. Cancer Causes & Control 2013; 24(6), 1099-1109.
D Chacon-Cortes, RA Smith, LM Haupt, RA Lea, PH Youl and LR Griffiths. Genetic association analysis of miRNA SNPs implicates MIR145 in breast cancer susceptibility. BMC Medical Genetics 2015; 16, 107.
J Gong, Y Tong, HM Zhang, K Wang, T Hu, G Shan, J Sun and AY Guo. Genome-wide identification of SNPs in microRNA genes and the SNP effects on microRNA target binding and biogenesis. Human Mutation 2012; 33(1), 254-263.
LH Huu, T Nguyen and N Hue. Association between single nucleotide polymorphism rs4919510 (c > g) in miRNA-608 and breast cancer in Vietnamese population. International Research Journal of Obstetrics and Gynecology 2019; 2, 11.
TTH Minh, NTN Thanh, TV Thiep and NT Hue. Association between single nucleotide polymorphism rs11614913 (C>T) on miR-196a2 and breast cancer in Vietnamese population. In: TV Van, TAN Le and TN Duc (Eds.). 6th international conference on the development of biomedical engineering in Vietnam (BME6). Springer, Singapore, 2018, p. 381-386.
TTN Thi Ngoc Nguyen, THN Nguyen, LH Huynh, HN Phan and HT Nguyen. Predicting SNPs in mature microRNAs dysregulated in breast cancer. In: L Tutar (Ed.). Recent advances in noncoding RNAs. IntechOpen, London, 2022.
NT Hue, NDH Chan, PT Phong, NTT Linh and NDT Giang. Extraction of human genomic DNA from dried blood spots and hair roots. International Journal of Bioscience, Biochemistry and Bioinformatics 2012; 2(1), 21-26.
S Roy, C Coldren, A Karunamurthy, NS Kip, EW Klee, SE Lincoln, A Leon, M Pullambhatla, RL Temple-Smolkin, KV Voelkerding, C Wang and AB Carter. Standards and guidelines for validating next-generation sequencing bioinformatics pipelines: A joint recommendation of the Association for Molecular Pathology and the College of American Pathologists. The Journal of Molecular Diagnostics 2018; 20(1), 4-27.
TF Scientific. Ion AmpliSeq™ designer performance specifications for custom DNA panel, Available at: https://www.ampliseq.com/otherContent/help-content/help_html/GUID-5193AC77-0A80-488E-B26A-1CD138BE1D21.html?utm_source=chatgpt.com, accessed October 2025.
F Muyas, M Bosio, A Puig, H Susak, L Domènech, G Escaramis, L Zapata, G Demidov, X Estivill, R Rabionet and S Ossowski. Allele balance bias identifies systematic genotyping errors and false disease associations. Human Mutation 2019; 40(1), 115-126.
H Thorvaldsdóttir, JT Robinson and JP Mesirov. Integrative Genomics Viewer (IGV): High-performance genomics data visualization and exploration. Briefings in Bioinformatics 2012; 14(2), 178-192.
W Huber, VJ Carey, R Gentleman, S Anders, M Carlson, BS Carvalho, HC Bravo, S Davis, L Gatto, T Girke, R Gottardo, F Hahne, KD Hansen, RA Irizarry, M Lawrence, MI Love, J MacDonald, V Obenchain, AK Oleś, H Pagès, A Reyes, …, M Morgan. Orchestrating high-throughput genomic analysis with Bioconductor. Nature Methods 2015; 12(2), 115-121.
LM Bragg, G Stone, MK Butler, P Hugenholtz and GW Tyson. Shining a light on dark sequencing: Characterising errors in Ion Torrent PGM data. PLoS Computational Biology 2013; 9(4), e1003031.
D Ballard, J Winkler-Galicki and J Wesoły. Massive parallel sequencing in forensics: Advantages, issues, technicalities, and prospects. International Journal of Legal Medicine 2020; 134(4), 1291-1303.
I Avent, AG Kinnane, N Jones, I Petermann, R Daniel, ME Gahan and D McNevin. The QIAGEN 140-locus single-nucleotide polymorphism (SNP) panel for forensic identification using massively parallel sequencing (MPS): An evaluation and a direct-to-PCR trial. International Journal of Legal Medicine 2019; 133(3), 677-688.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






