Quinazolinone Derivatives as Targeting pfDHFR and pfDHODH Inhibitor: In Silico Studies Using Molecular Docking, Molecular Dynamics Simulations, MM-PBSA, and ADMET Analysis
DOI:
https://doi.org/10.48048/tis.2026.11331Keywords:
Antimalarial, Quinazolinones, Molecular docking, Molecular dynamics, ADMET, Antimalarial, pfDHFR, pfDHODH, Quinazolinone, Molecular docking, Molecular dynamics, MM-PBSA, RAM, ADMETAbstract
Malaria remains a significant global health concern, with rising resistance to current antimalarial drugs. pfDHFR (Plasmodium falciparum Dihydrofolate Reductase) and pfDHODH (Plasmodium falciparum Dihydroorotate Dehydrogenase) are critical enzymes for parasite survival and have emerged as promising targets for drug development. Quinazolinones have shown potential as antimalarial agents due to their diverse pharmacological activities. This study aims to evaluate quinazolinones as potential inhibitors of pfDHFR and pfDHODH using molecular docking, molecular dynamics simulations, and ADMET (Absorption, Distribution, Metabolism, Excretion and Toxicity) profiling to identify optimal candidates with improved efficacy and pharmacokinetics. Thirty quinazolinone derivatives were subjected to molecular docking against pfDHFR and pfDHODH to assess binding affinity and interaction modes. Promising compounds were further analyzed through molecular dynamics simulations to evaluate complex stability. Additionally, ADMET profiling was conducted to predict pharmacokinetic properties and toxicity. Molecular docking identified compounds 3, 5, and 12 as promising pfDHFR inhibitors, with compound 3 exhibiting the most favorable binding energy. For pfDHODH, compounds 2, 11, and 20 showed strong interactions. Molecular dynamics simulations confirmed the stability of these complexes, with compounds 3 and 5 being stable against pfDHFR and compounds 20 and 11 against pfDHODH. ADMET analysis revealed favorable drug-like properties for these compounds, although some toxicity concerns were noted. This study demonstrates the potential of quinazolinone derivatives as next-generation antimalarial agents targeting pfDHFR and pfDHODH. Compounds 3, 5, 11, and 12 are identified as promising candidates due to their strong binding affinities and stable pharmacokinetic profiles. Further optimization and experimental validation are necessary to develop these compounds into effective therapeutic agents against malaria.
HIGHLIGHTS
- Thirty quinazolinone derivatives were designed and examined as potential inhibitors of pfDHFR and pfDHODH through molecular docking and dynamic simulations.
- C-Arylquinazolinones bearing hydroxyl groups were the most potential candidates.
- The ADMET results showed that the quinazolinones exhibited no potential toxicity.
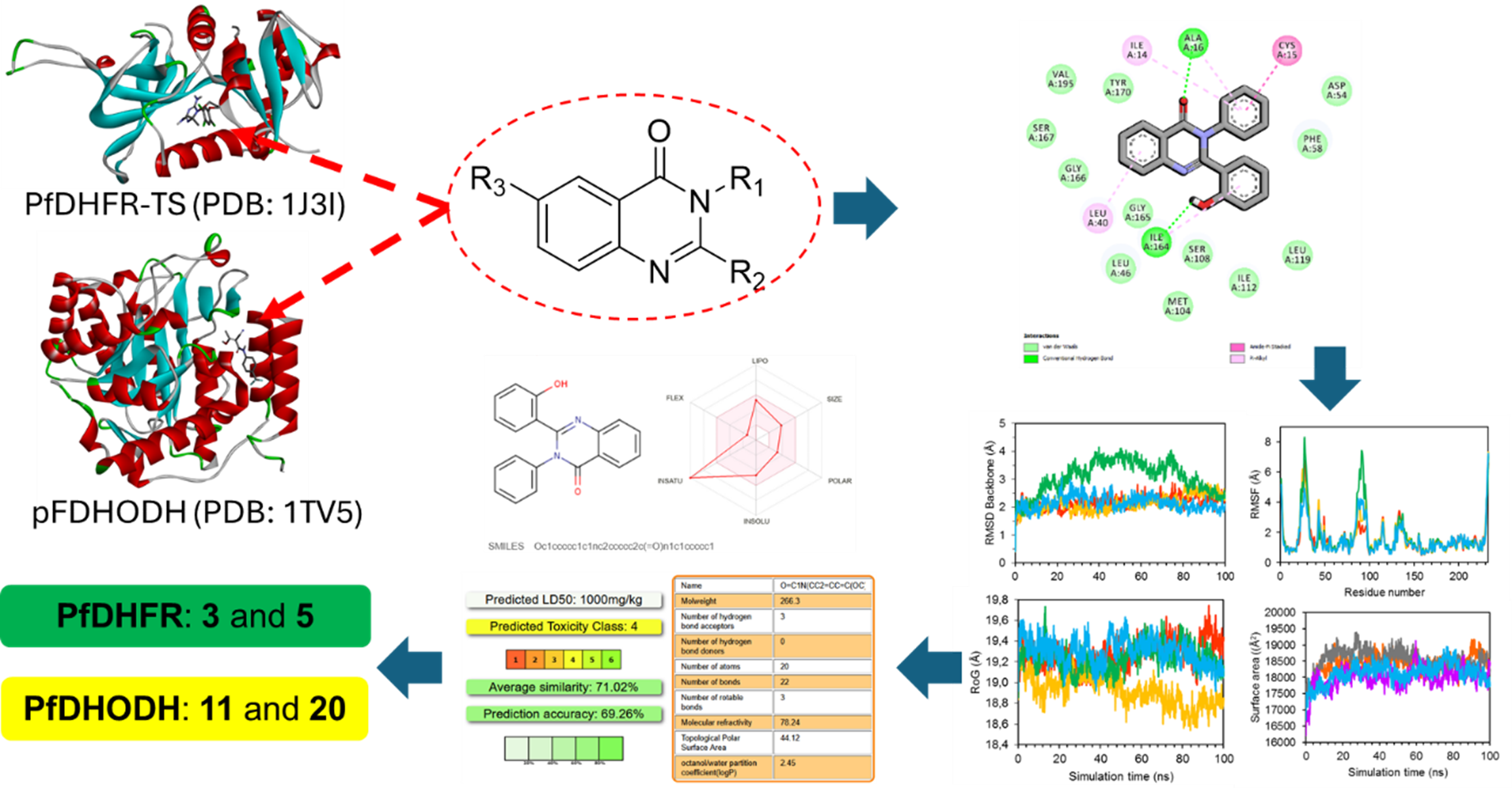
GRAPHICAL ABSTRACT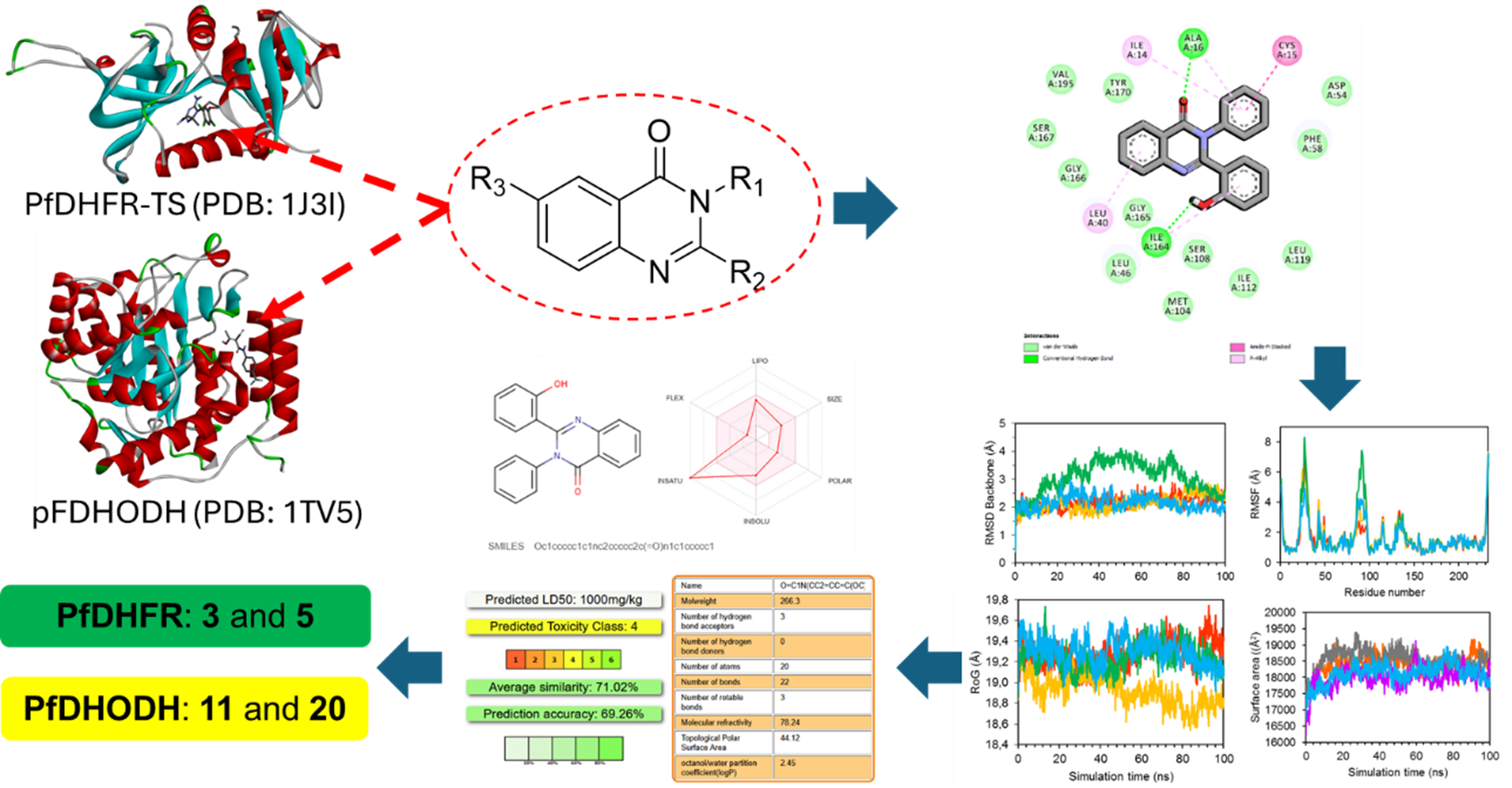
Downloads
References
A Alemu, B Lemma, T Bekele, G Geshere, EA Simma, CT Deressa and T Ketema. Malaria burden and associated risk factors among malaria suspected patients attending health facilities in Kaffa zone, Southwest Ethiopia. Malaria Journal 2024; 23(1), 397.
J Li, HJ Docile, D Fisher, K Pronyuk and L Zhao. Current status of malaria control and elimination in Africa: Epidemiology, diagnosis, treatment, progress and challenges. Journal of Epidemiology and Global Health 2024; 14, 561-579.
W Haileselassie, R Adam, M Habtemichael, RE David, N Solomon, S Workineh, J Haider, A Belachew, W Deressa, G Yan, N Kassaw and D Parker. Socio-demographic and economic inequity in the use of insecticide-treated bed nets during pregnancy: A survey-based case study of 4 sub-Saharan African countries with a high burden of malaria. Archives of Public Health 2023; 81(1), 64.
Y Wang, N Chitnis and E Fairbanks. Optimizing malaria vector control: A systematic review and mathematical modelling study to identify desirable characteristics of novel tools in different settings. Parasites & Vectors 2023. https://doi.org/10.21203/rs.3.rs-3332552/v1
LP Hastuti, F Hermawan, MR Iresha and T Ernawati. In-silico studies of hydroxyxanthone derivatives as potential pfDHFR and pfDHODH inhibitor by molecular docking, molecular dynamics simulation, MM-PBSA calculation and pharmacokinetics prediction. Informatics in Medicine Unlocked 2024; 47, 101485.
S Johannsen, RM Gierse, A Krüger, RL Edwards, V Nanna, A Fontana, D Zhu, T Masini, E de Souza, M Droge, B van Vliet, J den Hartog, M Hutter, J Held, Odom, A John, C Wrenger and A Hirsch. High target homology does not guarantee inhibition: Aminothiazoles emerge as inhibitors of Plasmodium falciparum. ACS Infectious Diseases 2024; 10(3), 1000-1022.
OB Akide Ndunge, N Kilian and MM Salman. Cerebral malaria and neuronal implications of Plasmodium Falciparum infection: From mechanisms to advanced models. Advanced Science 2022; 9(36), 202202944.
TM Schäfer, L Pessanha de Carvalho, J Inoue, A Kreidenweiss and J Held. The problem of antimalarial resistance and its implications for drug discovery. Expert Opinion on Drug Discovery 2024; 19(2), 209-224.
A Amusan, O Akinola, K Akano, M Hernández-Castañeda, JK Dick, A Sowunmi, G Hart and G Gbotosho. Frequency of chloroquine-resistant haplotype of Plasmodium falciparum (CVIET) in Ibadan, Southwest Nigeria 17 years post-chloroquine withdrawal. Acta Tropica 2024; 260, 107435.
G Matambisso, N Brokhattingen, S Maculuve, P Cístero, H Mbeve, A Escoda, G Bambo, B Cuna, C Melembe, N Ndimande, K Tetteh, C Drakeley, B Gamain, C Chitnis, V Chauhan, L Quinto, E Macete and A Mayor. Sustained clinical benefit of malaria chemoprevention with sulfadoxine-pyrimethamine (SP) in pregnant women in a region with high SP resistance markers. Journal of Infection 2024; 88, 106144.
E Kokori, G Olatunji, A Akinboade, A Akinoso, E Egbunu, SA Aremu, CE Okafor, O Oluwole and N Aderinto. Triple artemisinin-based combination therapy (TACT): advancing malaria control and eradication efforts. Malaria Journal 2024; 23, 25.
P Bravo, L Bizzarri, D Steinbrunn, J Lohse, AKH Hirsch, P Maser, M Rottmann and H Hahne. Integral solvent-induced protein precipitation for target-engagement studies in Plasmodium falciparum. ACS Infectious Diseases 2024; 10(12), 4073-4086.
M Sharma, V Pandey, G Poli, T Tuccinardi, ML Lolli and VK Vyas. A Comprehensive review of synthetic strategies and sar studies for the discovery of pfDHODH inhibitors as antimalarial agents. Part 1: Triazolopyrimidine, Isoxazolopyrimidine and pyrrole-based (DSM) compounds. Bioorganic Chemistry 2024; 146, 107249.
PM Cheuka, P Njaria, G Mayoka and E Funjika. Emerging drug targets for antimalarial drug discovery: Validation and insights into molecular mechanisms of function. Journal of Medicinal Chemistry 2024; 67(2), 838-863.
N Suwanakitti, Y Talawanich, J Vanichtanankul, S Taweechai, Y Yuthavong, S Kamchonwongpaisan and D Kongkasuriyachai. folA thyA knockout E. coli as a suitable surrogate model for evaluation of antifolate sensitivity against PfDHFR-TS. Acta Tropica 2024; 258, 107360.
H Shamshad, R Bakri and AZ Mirza. Dihydrofolate reductase, thymidylate synthase, and serine hydroxy methyltransferase: Successful targets against some infectious diseases. Molecular Biology Reports 2022; 49, 6659-6691.
X Zhang, Q Yang, X Zeng, Y Fu, Q Ding and Y Peng. Highly selective synthesis of selenium-containing (E)-N-propenolquinazolinones via FeCl 3-mediated cascade reaction of propargyl quinazoline-4-yl ethers with diselenides. Organic & Biomolecular Chemistry 2024; 22(47), 9231-9341.
SS Acharya, S Patra, LM Barad, A Roul and BB Parida. Recent advances in iodine-DMSO mediated C(sp3)-H functionalization of methyl-azaarenes via Kornblum oxidation. New Journal of Chemistry 2024; 48(17), 7614-7638.
D Sengupta, D Sharma, RK Das, P Das, M Halder, P Rai and O Chakrabarti. Pioneering the photoactive relevance of quinazolinone-fullereropyrrolidine nanohybrids to address chemotherapeutic resistance in cancer. ACS Medicinal Chemistry Letters 2024; 15(7), 1118-1126.
B Laleu, Y Akao, A Ochida, S Duffy, L Lucantoni, DM Shackleford, G Chen, K Katneni, F Chiu, K White, X Chen, A Sturm, K Dechering, B Crespo, L Sanz, B Wang, S Wittlin, S Charman, V Avery, N Cho and M Kamaura. Discovery and structure-activity relationships of quinazolinone-2-carboxamide derivatives as novel orally efficacious antimalarials. Journal of Medicinal Chemistry 2021; 64(17), 12582-12602.
UV Mhetre, NB Haval, GM Bondle, SS Rathod, PB Choudhari, J Kumari, D Sriram and KP Haval. Design, synthesis and molecular docking study of novel triazole-quinazolinone hybrids as antimalarial and antitubercular agents. Bioorganic & Medicinal Chemistry Letters 2024; 108, 129800.
JL Siqueira-Neto, KJ Wich, K Chibale, JN Burrows, DA Fidock and EA Winzeler. Author correction: Antimalarial drug discovery: Progress and approaches. Nature Reviews Drug Discovery 2024; 23, 880.
D Gheidari, M Mehrdad and Z Karimelahi. Virtual screening, ADMET prediction, molecular docking, and dynamic simulation studies of natural products as BACE1 inhibitors for the management of Alzheimer’s disease. Scientific Reports 2024; 14(1), 26431.
X Huang and J Hu. In silico discovery of novel androgen receptor inhibitors for prostate cancer therapy using virtual screening, molecular docking, and molecular dynamics simulations. Scientific Reports 2025; 15, 29404.
A Trezza, A Visibelli, B Roncaglia, R Barletta, S Iannielli, L Mahboob, O Spiga and A Santucci. Unveiling dynamic hotspots in protein-ligand binding: Accelerating target and drug discovery approaches. International Journal of Molecular Sciences 2025; 26(9), 3971.
S Ghani, N Khan, H Sable, F Yao and M Shafiq. Computational techniques for enhancing PK/PD modeling and simulation and ADMET prediction. In: M Ishfaq (Ed.). Computational methods in medicinal chemistry, pharmacology, and toxicology. Academic Press, New York, 2025, p. 153-174.
AB Reddy, TR Allaka, VSR Avuthu, K Chepuri, MZ Ahmed and H Nagarajaiah. New quinazolinone-1,2,4-triazole analogues: Synthesis, anticancer evaluation, molecular docking, and in silico ADMET prediction. Journal of Molecular Structure 2025; 1334, 141850.
MA Alamri, O Afzal, MJ Akhtar, S Karim, M Husain, MA Alossaimi and Y Riadi. Synthesis, in silico and in vitro studies of novel quinazolinone derivatives as potential SARS-CoV-2 3CLpro inhibitors. Arabian Journal of Chemistry 2024; 17(1), 105384.
AA Alsfouk, IMM Othman, MM Anwar, A Saleh, NY Tashkandi and ES Nossier. New quinazolinone-based heterocyclic compounds as promising antimicrobial agents: Development, DNA gyrase B/topoisomerase IV inhibition activity, and in silico computational studies. Journal of Molecular Structure 2025; 1344, 142953.
YS Kurniawan, E Yudha, G Nugraha, N Fatmasari, HD Pranowo, J Jumina and EN Sholikhah. Molecular docking and molecular dynamic investigations of xanthone-chalcone derivatives against epidermal growth factor receptor for preliminary discovery of novel anticancer agent. Indonesian Journal of Chemistry 2024; 24(1), 250-266.
MO Miranda, DJR Duarte and VM Rayón. The influence of halogen-mediated interactions on halogen abstraction reactions by formyl radicals. Physical Chemistry Chemical Physics 2025; 27(6), 3330-3340.
PR Varadwaj, HM Marques and A Varadwaj. π-hole halogen bonds are sister interactions to σ-hole halogen bonds. Crystal Growth & Design 2024; 24(19), 7789-7807.
R Canabal and C González-Bello. Chemical sensors for the early diagnosis of bacterial resistance to β-lactam antibiotics. Bioorganic Chemistry 2024; 150, 107528.
RA Laskowski, J Jabłońska, L Pravda, RS Vařeková and JM Thornton. PDBsum: Structural summaries of PDB entries. Protein Science 2018; 27(1), 129-134.
K Saritha, M Alivelu and M Mohammad. Drug-likeness analysis, in silico ADMET profiling of compounds in Kedrostis foetidissima (Jacq.) Cogn, and antibacterial activity of the plant extract. In Silico Pharmacology 2024; 12(2), 67.
MY Alsedfy, AA Ebnalwaled, M Moustafa and AH Said. Investigating the binding affinity, molecular dynamics, and ADMET properties of curcumin-IONPs as a mucoadhesive bioavailable oral treatment for iron deficiency anemia. Scientific Reports 2024; 14, 22027.
H Guterres and W Im. CHARMM-GUI-based induced fit docking workflow to generate reliable protein-ligand binding modes. Journal of Chemical Information and Modeling 2023; 63(15), 4772-4779.
R Touir, N Errahmany, M Rbaa, F Benhiba, M Doubi, EH EL Kafssaoui and B Lakhrissi. Experimental and computational chemistry investigation of the molecular structures of new synthetic quinazolinone derivatives as acid corrosion inhibitors for mild steel. Journal of Molecular Structure 2024; 1303, 137499.
GO Oduselu, OF Elebiju, TA Ogunnupebi, S Akash, OO Ajani and E Adebiyi. Employing hexahydroquinolines as PfCDPK4 inhibitors to combat malaria transmission: An advanced computational approach. Advances and Applications in Bioinformatics and Chemistry 2024; 17, 83-105.
MA Ojong, N Mujafarkani, FAK Khazaal, AS Hussam, OC Godfrey, K Muzammil, A Ahamed, R Edadi, I Anyambula, E Moses and I Benjamin. Investigating the impact of solvation on p-Phenylenediamine-2-Amino pyrimidine-Formaldehyde Terpolymer (P2APF) ligand’s reactivity and drug suitability for malaria treatment: Insights from experimental and quantum calculations. Journal of Molecular Structure 2024; 1310, 138113.
S Jaiswal, K Verma, A Srivastva, N Arya, J Dwivedi and S Sharma. Green synthetic and pharmacological developments in the hybrid quinazolinone moiety: An updated review. Current Topics in Medicinal Chemistry 2025; 25(5), 493-532.
MM Mansouri, L Emami, Z Rezaei and S Khabnadideh. Design, synthesis, biological assessments and computational studies of 3-substituted phenyl quinazolinone derivatives as promising anti-cancer agents. BMC Chemistry 2025; 19(1), 125.
RM Hassan, H Yehia, MF El-Behairy, AAS El-Azzouny and MN Aboul-Enein. Design and synthesis of new quinazolinone derivatives: investigation of antimicrobial and biofilm inhibition effects. Molecular Diversity 2025; 29(1), 21-42.
S Moghadam Farid, S Moradi Dehaghi, A Iraji, M Mahdavi and M Saeedi. Synthesis, biological evaluations, and in silico assessments of phenylamino quinazolinones as tyrosinase inhibitors. Scientific Reports 2025; 15(1), 846.
M Alruwaili, HH Alhassan, H Almutary and M Tahir ul Qamar. Computational identification of aspartic protease inhibitors for antimalarial drug development against Plasmodium Vivax. Scientific Reports 2025; 15, 14824.
I Sama-ae, P Muengthongon, A Tohlaeh, W Rukhachan, P Kiattikul, F Samaeng, A Mitklin, P Kwankaew, M Kotepui and A Kepan. Penicillium‐derived inhibitors of Plasmodium falciparum lactate dehydrogenase (PfLDH): A computational approach for novel antimalarial therapy development. Scientifica 2025; 2025(1), 8838031.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






