Advancing Marine Genomics: The Role of Deep Learning in Deciphering Chelonia mydas Genetic Data
DOI:
https://doi.org/10.48048/tis.2025.9149Abstract
The urgent advancement of marine genomics is essential for the conservation of endangered species like Chelonia mydas (green sea turtle), and deep learning plays a pivotal role in deciphering their complex genetic data. Marine genomics as a field seems to be shifting more and more to the realm of big data especially with the introduction of new technologies in producing vast amounts of data. Such advancements have made it possible to manage huge datasets within genomics and this has provided artificial intelligence and particularly deep learning as a crucial method of acquiring insightful patterns. Review of the subject aims at focusing on the subject of deep learning methods and their usefulness in appearance and utilization in sub disciplinary areas of genomics of Chelonia mydas (green sea turtle). We introduce deep learning into marine genomics research by pointing out the existing gaps, as well as well-PSYCH detailed study fields. Moreover, we can only briefly mention the rather late incorporation of deep learning tools into marine genomics and the eminently discussed consequences for conservation and ecological science. By writing this review the authors envisage to let the biotechnology and genomic scientists to know the importance and applicability of using deep learning methods in Chelonia mydas genomics, the difficulty and prospects of this field.
HIGHLIGHTS
- Emphasize the role of meta-omic integration and the large-scale DL model the power of conservation genetics, population monitoring and disease surveillance, as well as the assessment of habitat quality.
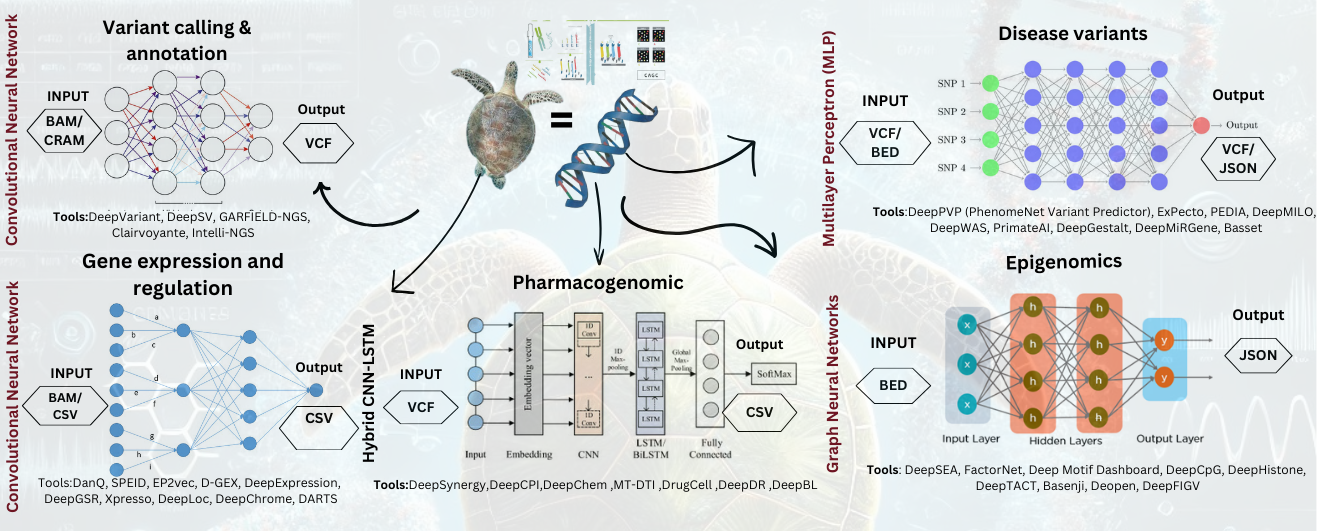
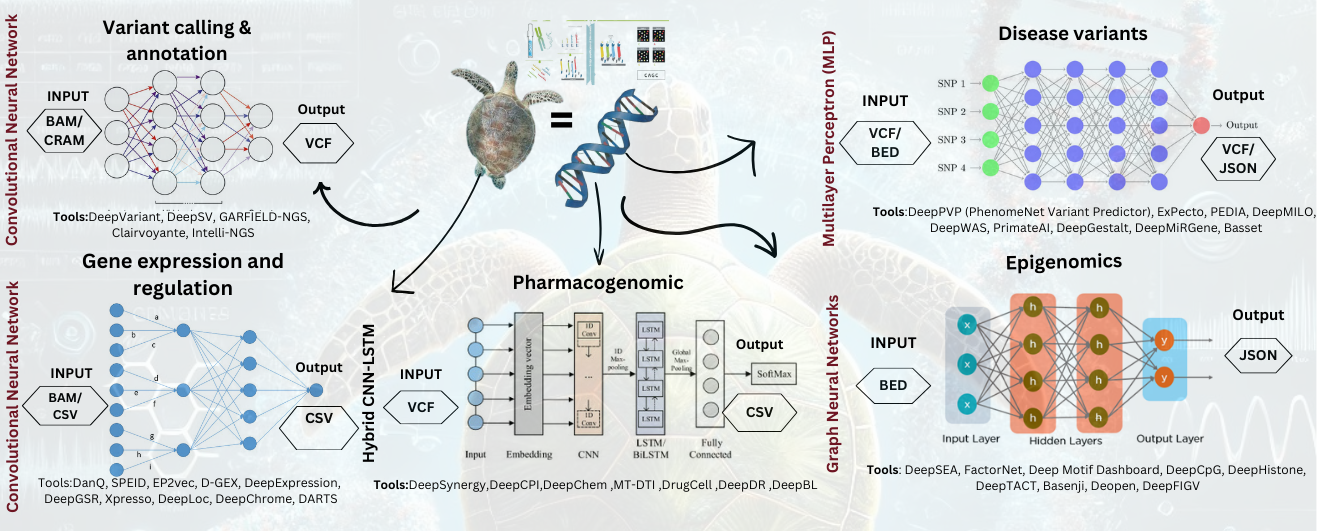
- Introduces Real Implementation of DL tools including DeepVariant, DeepSEA, DeepCpG and DeepSynergy that have enhanced the accuracy by the thirty percent and the computational speed of over forty percent.
- Introduces some of the most important frameworks based on DL which help researchers investigate the pattern of gene expression and epigenomic changes in Chelonia mydas such as DanQ, DeepChrome, and DeepHistone.
- Explains the possible application of DL in genomics and pharmacogenomics of Chelonia mydas, variant calling and epigenetics, gene expression analysis.
- Stresses DL approaches for identifying and predicting genetic variations, regulatory sites and drugs using more accuracy and capability.
GRAPHICAL ABSTRACT
Downloads
References
ELV Dijk, H Auger, Y Jaszczyszyn and C Thermes. Ten years of next-generation sequencing technology. Trends in Genetics 2014; 30(9), 418-426.
MW Libbrecht and WS Noble. Machine learning applications in genetics and genomics. Nature Reviews Genetics 2015; 16(6), 321-332.
B Alipanahi, A Delong, MT Weirauch and BJ Frey. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nature Biotechnology 2015; 33(8), 831-838.
Y Lecun, Y Bengio and G Hinton. Deep learning. Nature 2015; 521(7553), 436-444.
D Chicco and G Jurman. The advantages of the Matthews correlation coefficient (MCC) over F1 score and accuracy in binary classification evaluation. BMC Genomics 2020; 21(6), 6-14.
G Mao, Z Pang, K Zuo, Q Wang, X Pei, X Chen, and J Liu. Predicting gene regulatory links from single-cell RNA-seq data using graph neural networks. Briefings in Bioinformatics 2023; 24(6), bbad414.
I Goodfellow, Y Bengio and A Courville. Deep learning. The MIT Press, Cambridge, Massachusetts, 2018, p. 800.
J Schmidhuber. Deep learning in neural networks: An overview. Neural Networks 2015; 61, 85-117.
J Zhou, CL Theesfeld, K Yao, KM Chen, A Wong, and O Troyanskaya. Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk. Nature Genetics 2018; 50(8), 1171-1179.
V Boža, B Brejová and T Vinař. DeepNano: Deep recurrent neural networks for base calling in MinION nanopore reads. PLoS One 2017; 12(6), e0178751.
H Zeng, MD Edwards, G Liu and DK Gifford. Convolutional neural network architectures for large-scale DNA sequence classification and motif discovery. Bioinformatics 2018; 34(12), 1014-1020.
R Luo, FJ Sedlazeck, TW Lam and MC Schatz. Clairvoyante: A multi-task convolutional deep neural network for variant calling in single molecule sequencing. Nature Communications 2019; 10(1), 1-11.
TC Hsieh, MA Mensah, JT Pantel, D Aguilar, O Bar, A Bayat, L Becerra-Solano, HB Bentzen, S Biskup, O Borisov, O Braaten, C Ciaccio, M Coutelier, K Cremer, M Danyel, S Daschkey, HD Eden, K Devriendt, S Wilson, S Douzgou, ... PM Krawitz. PEDIA: prioritization of exome data by image analysis. Genetics in Medicine 2019, 21(12), 2807-2814.
R Singh, J Lanchantin, G Robins and Y Qi. DeepChrome: Deep-learning for predicting gene expression from histone modifications. Bioinformatics 2016; 32(17), i639-i648.
A Singh and P Bhatia. Intelli-NGS: Intelligent NGS, a deep neural network-based artificial intelligence to delineate good and bad variant calls from IonTorrent sequencer data. bioRxiv 2019, https://doi.org/10.1101/2019.12.17.879403
C Angermueller, T Pärnamaa, L Parts and O Stegle. Deep learning for computational biology. Molecular Systems Biology 2016; 12(7), 878.
S Min, B Lee and S Yoon. Deep learning in bioinformatics. Briefings in Bioinformatics 2017; 18(5), 851-869.
J Zou, M Huss, A Abid, P Mohammadi, A Torkamani and A Telenti. A primer on deep learning in genomics. Nature Genetics 2018; 51(1), 12-18.
AD Ewing and RE Green. Finding elusive structural variants in 1000 Genomes Project data. Genome Research 2015; 25(10), 1516-1523.
R Nielsen, JS Paul, A Albrechtsen and YS Song. Genotype and SNP calling from next-generation sequencing data. Nature Reviews Genetics. 2011; 12(6), 443-451.
R Poplin, PC Chang, D Alexander, S Schwartz, T Colthurst, A Ku, D Newburger, J Dijamco, N Nguyen, PT Afshar, SS Gross, L Dorfman, CY McLean and MA DePristo. A universal SNP and small-indel variant caller using deep neural networks. Nature Biotechnology 2018; 36(10), 983-987.
L Cai, Y Wu and J Gao. DeepSV: Accurate calling of genomic deletions from high-throughput sequencing data using deep convolutional neural network. BMC Bioinformatics 2019, 20, 665.
L Sundaram, H Gao, SR Padigepati, JF McRae, Y Li, JA Kosmicki, N Fritzilas, J Hakenberg, A Dutta, J Shon, J Xu, S Batzoglou, X Li and KH Farh. Predicting the clinical impact of human mutation with deep neural networks. Nature Genetics 2018, 50(8), 1161-1170.
DR Kelley, J Snoek, and JL Rinn. Basset: Learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Research 2016; 26(7), 990-999.
I Boudellioua, M Kulmanov, PN Schofield, GV Gkoutos and R Hoehndorf. DeepPVP: phenotype-based prioritization of causative variants using deep learning. BMC Bioinformatics 2019; 20(1), 65.
J Zhou, CL Theesfeld, K Yao, KM Chen, AK Wong and OG Troyanskaya. Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk. Nature Genetics 2018; 50, 1171-1179.
J Arloth, G Eraslan, TFM Andlauer, J Martins, S Iurato, B Kühnel, M Waldenberger, J Frank, R Gold, B Hemmer, BB Ebert, H Akil, E Binder, M Hrabě de Angelis, K-A Nave, MJ Bamberg, and FJ Theis. DeepWAS: Multivariate genotype-phenotype associations by directly integrating regulatory information using deep learning. PLOS Computational Biology 2020; 16(2), e1007616.
Y Gurovich, Y Hanani, O Bar, G Nadav, N Fleischer, D Gelbman, L Basel-Salmon, PM Krawitz, SB Kamphausen, M Zenker, LM Bird, and KW Gripp. Identifying facial phenotypes of genetic disorders using deep learning. Nature Medicine 2019; 25(1), 60-64.
A Vaswani, N Shazeer, N Parmar, J Uszkoreit, L Jones, AN Gomez, L Kaiser and I Polosukhin. Attention is all you need. Advances in Neural Information Processing Systems 2017; 30, 5998-6008.
DL Black. Mechanisms of alternative pre-messenger RNA splicing. Annual Review of Biochemistry 2003; 72(1), 291-336.
C Angermueller, HJ Lee, W Reik and O Stegle. DeepCpG: Accurate prediction of single-cell DNA methylation states using deep learning. Genome Biology 2017; 18, 67.
T Steijger, JF Abril, PG Engström, F Kokocinski, The RGASP Consortium, TJ Hubbard, R Guigó, J Harrow, and P Bertone. Assessment of transcript reconstruction methods for RNA-seq. Nature Methods 2013; 10(12), 1177-1184.
X Li, K Wang, Y Lyu, H Pan, J Zhang, D Stambolian, K Susztak and MP Reilly. Deep learning enables accurate clustering with batch effect removal in single-cell RNA-seq analysis. Nature Communications 2020; 11(1), 1-14.
VY Kiselev, K Kirschner, MT Schaub, T Andrews, A Yiu, T Chandra, KN Natarajan, W Reik, M Barahona, AR Green, and M Hemberg. SC3: Consensus clustering of single-cell RNA-seq data. Nature Methods 2019; 14(5), 483-486.
D Chicco and G Jurman. The advantages of the Matthews correlation coefficient (MCC) over F1 score and accuracy in binary classification evaluation. BMC Genomics 2020; 21, 6.
D Quang and X Xie. DanQ: A hybrid convolutional and recurrent deep neural network for quantifying the function of DNA sequences. Nucleic Acids Research 2016; 44(11), e107.
S Singh, Y Yang, B Póczos, and J Ma. Predicting enhancer-promoter interactions with neural networks. bioRxiv 2016, https://doi.org/10.1101/085241
W Zeng, M Wu and R Jiang. Prediction of enhancer–promoter interactions via natural language processing. BMC Genomics 2018; 19(S2), 84.
Y Chen, Y Li, R Narayan, A Subramanian and X Xie. Gene expression inference with deep learning. Bioinformatics 2016; 32(12), 1832-1839.
J Zrimec, CS Börlin, F Buric, AS Muhammad, R Chen, V Siewers, V Verendel, J Nielsen, M Töpel and A Zelezniak. Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure. Nature Communications 2020; 11, 6141.
M Kalkatawi, A Magana-Mora, B Jankovic and VB Bajic. DeepGSR: An optimized deep-learning structure for the recognition of genomic signals and regions. Bioinformatics 2019, 35(7), 1125-1132.
V Agarwal and J Shendure. Predicting mRNA abundance directly from genomic sequence using deep convolutional neural networks. Cell Reports 2020; 31(7), 107663.
JJA Armenteros, CK Sønderby, SK Sønderby, H Nielsen and O Winther. DeepLoc: Prediction of protein subcellular localization using deep learning. Bioinformatics 2017; 33(21), 3387-3395.
Z Zhang, Z Pan, Y Ying, Z Xie, S Adhikari, J Phillips, RP Carstens, DL Black, Y Wu and Y Xing. (2019). Deep-learning augmented RNA-seq analysis of transcript splicing. Nature Methods 2019; 16(4), 307-310.
Q Liu, F Xia, Q Yin and R Jiang. Chromatin accessibility prediction via a hybrid deep convolutional neural network. Bioinformatics 2018; 34(5), 732-738.
Q Yin, M Wu, Q Liu and R Jiang. DeepHistone: A deep learning approach to predicting histone modifications. BMC Genomics 2019; 20(S2), 193.
J Zhou and OG Troyanskaya. Predicting effects of noncoding variants with deep learning-based sequence model. Nature Methods 2015; 12(10), 931-934.
D Quang and X Xie. FactorNet: A deep learning framework for predicting cell type-specific transcription factor binding from nucleotide-resolution sequential data. Methods 2019; 166, 40-47.
W Li, WH Wong and R Jiang. DeepTACT: Predicting 3D chromatin contacts via bootstrapping deep learning. Nucleic Acids Research 2019; 47(10), e60.
DR Kelley, YA Reshef, M Bileschi, D Belanger, CY McLean and J Snoek. Sequential regulatory activity prediction across chromosomes with convolutional neural networks. Genome Research 2018; 28(5), 739-750.
GE Hoffman, J Bendl, K Girdhar, EE Schadt and P Roussos. Functional interpretation of genetic variants using deep learning predicts impact on chromatin accessibility and histone modification. Nucleic Acids Research 2019; 47(20), 10597-10611.
J Lanchantin, R Singh, B Wang and Y Qi. Deep Motif Dashboard: Visualizing and understanding genomic sequences using deep neural networks. Pacific Symposium on Biocomputing 2017; 22, 254-265.
K Preuer, RPI Lewis, S Hochreiter, A Bender, KC Bulusu and G Klambauer. DeepSynergy: Predicting anticancer drug synergy with deep learning. Bioinformatics 2018; 34(8), 1538-1546.
F Wan, Y Zhu, H Hu, A Dai, X Cai, L Chen, H Gong, T Xia, D Yang, MW Wang and J Zeng. DeepCPI: A deep learning-based framework for large-scale in silico drug screening. Genomics, Proteomics & Bioinformatics 2019, 17(5), 478-495.
B Ramsundar, P Eastman, P Walters, V Pande, K Leswing and Z Wu. (2019). Deep learning for the life sciences. O'Reilly Media. Available at: https://www.amazon.com/Deep-Learning-Life-Sciences-Microscopy/dp/1492039837, accessed July 2024.
B Shin, S Park, K Kang and JC Ho. Self-attention based molecule representation for predicting drug-target interaction. In: Proceedings of the 4th Machine Learning for Healthcare Conference, Proceedings of Machine Learning Research, Ann Arbor, Michigan. 2019, p. 230-248.
BM Kuenzi, J Park, SH Fong, KS Sanchez, J Lee, JF Kreisberg, J Ma and T Ideker, T. Predicting drug response and synergy using a deep learning model of human cancer cells. Cancer Cell 2020; 38(5), 672-684.
X Zeng, S Zhu, X Liu, Y Zhou, R Nussinov and F Cheng. deepDR: A network-based deep learning approach to in silico drug repositioning. Bioinformatics 2019; 35(24), 5191-5198.
Y Wang, F Li, M Bharathwaj, NC Rosas, A Leier, T Akutsu, GI Webb, TT Marquez-Lago, J Li, T Lithgow and J Song. DeepBL: A deep learning-based approach for in silico discovery of beta-lactamases. Briefings in Bioinformatics 2021; 22(4), bbaa301.
Y Bengio, P Simard and P Frasconi. Learning long-term dependencies with gradient descent is difficult. IEEE Transactions on Neural Networks 1994; 5(2), 157-166.
K Cho, BV Merrienboer, C Gulcehre, F Bougares, H Schwenk and Y Bengio. Learning phrase representations using RNN encoder-decoder for statistical machine translation. In: Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing, Doha, Qatar. 2014, p. 1724-1734.
A Graves, A Mohamed and G Hinton. Speech recognition with deep recurrent neural networks. In: Proceedings of the 2013 IEEE International Conference on Acoustics, Speech and Signal Processing, Vancouver, Canada. 2013, p. 6645-6649.
W Kopp, R Monti, A Tamburrini, U Ohler and A Akalin. Deep learning for genomics using Janggu. Nature Communications 2020, 11, 3488.
KM Chen, EM Cofer, J Zhou and OG Troyanskaya. Selene: A PyTorch-based deep learning library for sequence data. Nature Methods 2019; 16(4), 315-318.
S Budach and A Marsico. Pysster: Classification of biological sequences by learning sequence motifs with convolutional neural networks. Bioinformatics 2018; 34(17), 3035-3037.
ŽAvsec, R Kreuzhuber, J Israeli, N Xu, J Cheng, A Shrikumar, A Banerjee, DS Kim, T Beier, L Urban, A Kundaje, O Stegle and J Gagneur. The Kipoi repository accelerates community exchange and reuse of predictive models for genomics. Nature Biotechnology 2019; 37(6), 592-600.
TM Poolman, A Townsend-Nicholson and A Cain. Teaching genomics to life science undergraduates using cloud computing platforms with open datasets. Biochemistry and Molecular Biology Education 2022; 50(5), 446-449.
A Rodriguez, Y Kim, TN Nandi, K Keat, R Kumar, R Bhukar, M Conery, M Liu, J Hessington, K Maheshwari, D Schmidt, VA Million Veteran Program, E Begoli, G Tourassi, S Muralidhar, P Natarajan, BF Voight, K Cho, JM Gaziano, SM Damrauer, KP Liao, W Zhou, JE Huffman, A Verma and RK Madduri (2024). Accelerating genome- and phenome-wide association studies using GPUs - A case study using data from the Million Veteran Program. bioRxiv 2024, https://doi.org/10.1101/2024.05 .17.594583
T Ohta, T Tanjo and O Ogasawara. Accumulating computational resource usage of genomic data analysis workflow to optimize cloud computing instance selection. GigaScience 2019; 8(4), giz052.
A Taylor-Weiner, F Aguet, NJ Haradhvala, S Gosai, S Anand, J Kim and G Getz. Scaling computational genomics to millions of individuals with GPUs. Genome Biology 2019; 20, 228.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2024 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






