Genetic Diversity of the Endemic Species Phanera sirindhorniae in the Mekong Basin of Thailand Based on ISSR Markers
DOI:
https://doi.org/10.48048/tis.2023.7053Keywords:
Genetic diversity, ISSR, Mekong Basin of Thailand, Phanera sirindhorniaeAbstract
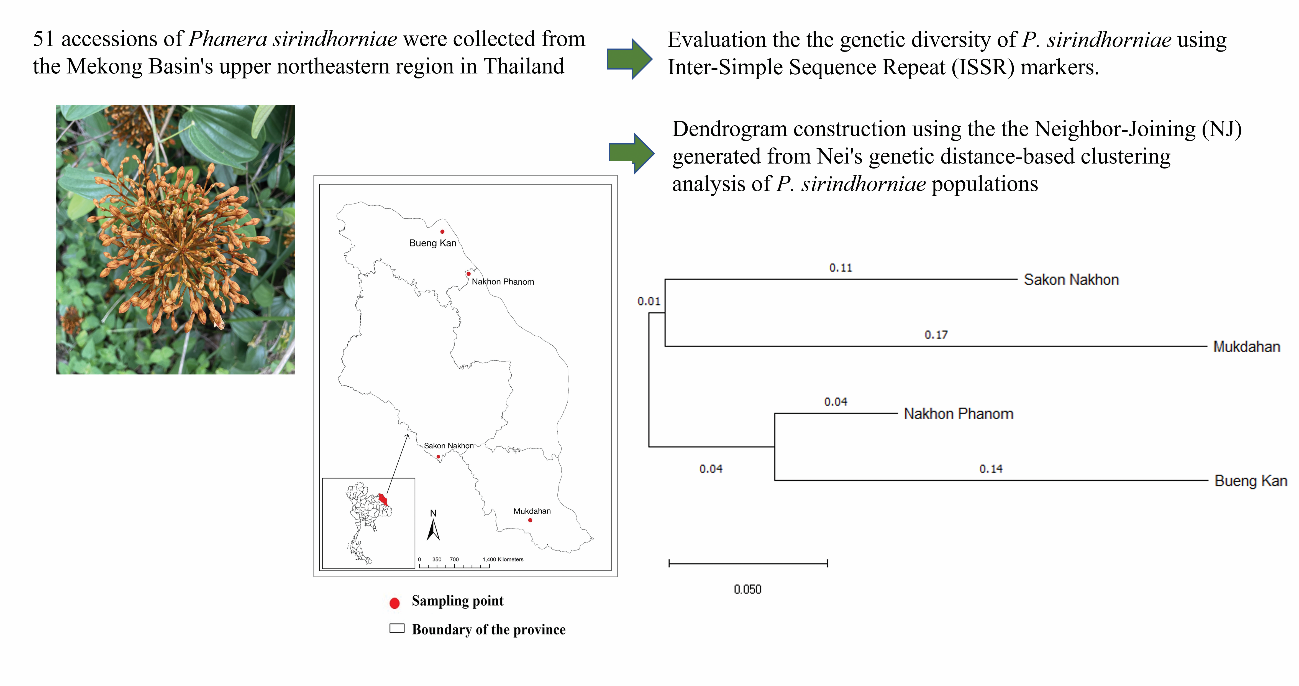
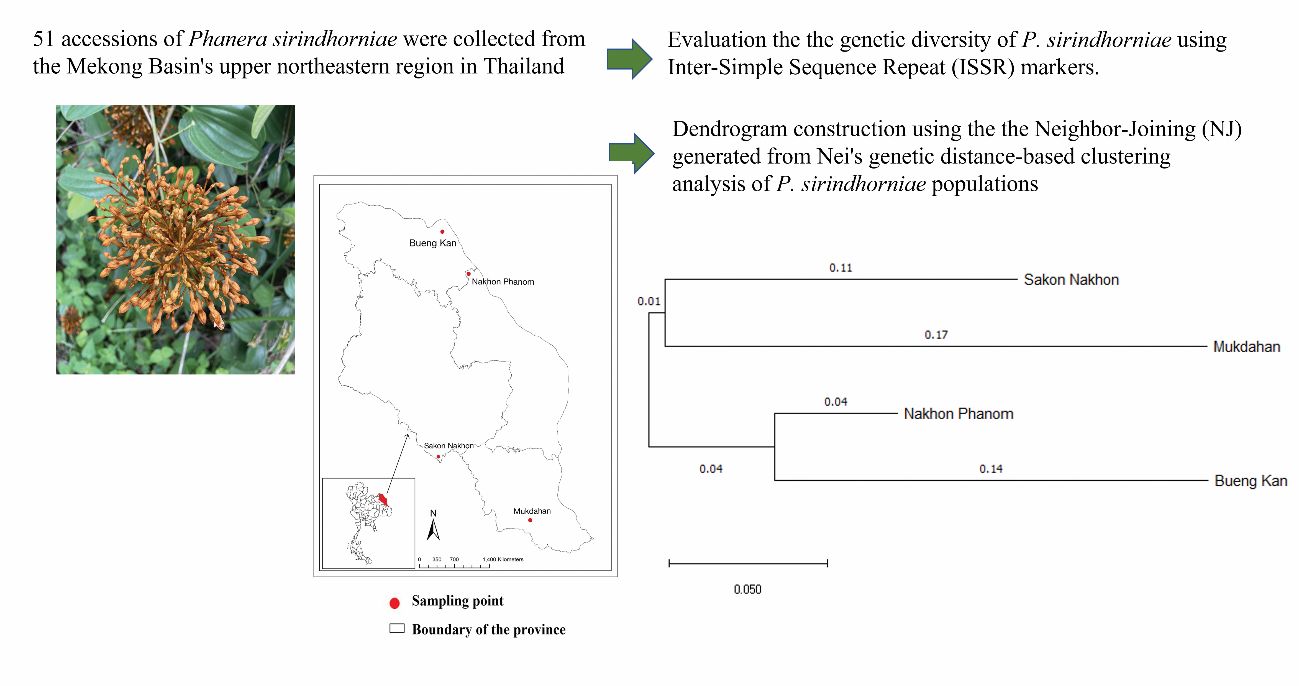
Phanera sirindhorniae, an endemic species to the Mekong Basin’s upper northeastern region in Thailand, is notable for its therapeutic properties in alleviating pain and discomfort. The objective of this study was to assess the genetic diversity of P. sirindhorniae derived from four natural sources, specifically Sakon Nakhon, Nakhon Phanom, Bueng Kan and Mukdahan provinces, utilizing Inter-Simple Sequence Repeat (ISSR) markers. A total of 51 samples were analyzed, and 10 primers exhibited distinct, robust amplified fragments and polymorphism. The amplified fragment sizes ranged from 170 to 1,300 base pairs, identifying a total of 113 amplified fragments, of which 107 (94.69 %) displayed polymorphism. Polymorphic information content (PIC) values spanned between 0.112 and 0.429, while Nei’s gene diversity ranged from 0.2088 ± 0.2184 to 0.2994 ± 0.1912. Assessment of genetic diversity within and among populations revealed the total genetic diversity (Ht = 0.3443 ± 0.0225) and the genetic diversity within populations (Hs = 0.2617 ± 0.0161). The variability among populations (Gst = 0.2401) indicated a significant degree of gene flow (Nm = 1.580). The Analysis of Molecular Variance (AMOVA) results supported this, with 74 % of genetic differentiation occurring within populations and 26 % among populations. Using the Neighbor-Joining (NJ) generated from Nei’s genetic distance-based clustering analysis, Dendrogram construction successfully classified 2 distinct groups. The analysis revealed that
P. sirindhorniae from Nakhon Phanom, and Bueng Kan constituted 1 group, whereas P. sirindhorniae from Mukdahan, and Sakon Nakhon formed another. These findings provide essential data for developing genetic conservation initiatives for this valuable genetic resource.
HIGHLIGHTS
- This study presents the inaugural analysis of genetic diversity in wild Phanera sirindhorniae populations, a species designated as endangered, utilizing Inter-Simple Sequence Repeat (ISSR) markers.
- A group of 10 Inter Simple Sequence Repeat (ISSR) primers showed values for Polymorphism Information Content (PIC), with (AG)8YC (PIC= 0.445) standing out. These results highlight their potential for use in the investigation of genetic diversity.
- The examination of P. sirindhorniae resulted in the identification of 2 distinct genetic groups, which have the potential to inform and facilitate the development of genetic conservation strategies for this crucial genetic resource.
GRAPHICAL ABSTRACT
Downloads
References
S Athikomkulchai, N Sriubolmas and N Ruangrungsi. Antibacterial activity of flavonoids from Bauhinia sirindhorniae. Thai J. Health Res. 2005; 19, 13-9.
K Larsen and SS Larsen. Bauhinia sirindhorniae sp. nov. (Leguminosa Caesalpinioideae) a remarkable new species from Thailand. Nord. J. Bot. 1997; 17, 113-8.
P Chaisri and N Laoprom. Antioxidant properties and total phenolic content of selected traditional Thai medicinal plants. Thai Pharmaceut. Health Sci. J. 2017; 12, 10-8.
A Sirivan, R Nijsiri and S Toshikazu. Chemical constituents of Bauhinia sirindhorniae. Herb. Med. J. 2003; 57, 150-3.
N Nithikulworawong. Antibacterial activity of Bauhinia sirindhorniae extract against Aeromonas hydrophila isolated from hybrid catfish. Walailak J. Sci. Tech. 2012; 9, 195-9.
N Nithikulworawong. Efficacy of Bauhinia sirindhorniae on resistance to against Streptococcus agalactiae in Nile tilapia (Oreochromis niloticus). Asia Pac. J. Sci. Tech. 2012; 17, 715-24.
B Milligan, J Leebens‐Mack and A Strand. Conservation genetics: Beyond the maintenance of marker diversity. Mol. Ecol. 1994; 3, 423-35.
MH Mills and MW Schwartz. Rare plants at the extremes of distribution: broadly and narrowly distributed rare species. Biodiversity Conservat. 2005; 14, 1401-20.
S Tomasello, I Álvarez, P Vargas and C Oberprieler. Is the extremely rare Iberian endemic plant species Castrilanthemum debeauxii (Compositae, Anthemideae) a ‘living fossil’? Evidence from a multi-locus species tree reconstruction. Mol. Phylogenet. Evol. 2015; 82, 118-30.
N Chachai, B Pensuriya, T Pinsuntiae, P Pratubkong, J Mungngam, P Nitmee, P Kaewsri, S Wongsatchanan, R Jindajia, P Triboun and J Sreesaeng. Variability of morphological and agronomical characteristics of Centella asiatica in Thailand. Trends Sci. 2021; 18, 502.
M Kamali, D Samsampour,A Bagheri, A Mehrafarin and A Homaei. Association analysis and evaluation of genetic diversity of Teucrium stocksianum Boiss. populations using ISSR markers. Genet. Resour. Crop Evol. 2023; 70, 691-709.
O Pakhrou, L Medraoui, C Yatrib, M Alami, A Filali-Maltouf and B Belkadi. Assessment of genetic diversity and population structure of an endemic Moroccan tree (Argania spinosa L.) based in IRAP and ISSR markers and implications for conservation. Physiol. Mol. Biol. Plant. 2017; 23, 651-61.
A El-Moneim, EIS ELsarag, S Aloufi, AM El-Azraq, SM ALshamrani, FAA Safhi and AA Ibrahim. Quinoa (Chenopodium quinoa Willd.): Genetic diversity according to ISSR and SCoT markers, relative gene expression, and morpho-physiological variation under salinity stress. Plants 2021; 10, 2802.
M Yaman. Determination of genetic diversity in european cranberrybush (Viburnum opulus L.) genotypes based on morphological, phytochemical and ISSR markers. Genet. Resour. Crop Evol. 2022; 69, 1889-99
F Aliabadi, A Bagheri, S Abbasi, H Saeidi and FR Blattner. High genetic diversity in an endemic and vulnerable species: Evidence from Astragalus cyclophyllon (Fabaceae) in Iran. Genet. Resour. Crop Evol. 2023; 70, 1999-2008.
NS Zvyagina and OV Dorogina. Genetic differentiation of Altai-Sayan endemic Hedysarum theinum Krasnob. (Fabaceae) evaluated by inter-simple sequence repeat analysis. Russ. J. Genet. 2013; 49, 1030-5.
SA Dev, M Shenoy and RM Borges. Genetic and clonal diversity of the endemic ant-plant Humboldtia brunonis (Fabaceae) in the Western Ghats of India. J. Biosci. 2010; 35, 267-79.
N Rahali, I Yangui, M Boussaid and C Messaoud. Assessment of genetic diversity and population structure of the endemic Hertia cheirifolia (L.) Kuntze based on ISSR and SRAP molecular markers. Biologia 2022; 77, 3429-39.
F Kazemeini, Y Asri G Mostafavi, R Kalvandi and I Mehregan. Assessment of genetic diversity, population structure and morphological analyses in an Iranian endemic species Rhabdosciadium aucheri Boiss. (Apiaceae) using ISSR markers. Biologia 2021; 76, 441-51.
A Lal, M Pant, G Palni, LMS Kumar, A Kumar and G Kumar. ISSR marker assisted genetic diversity assessment in natural populations of two endemic orchids Aerides multiflora and Rhynchostylis retusa from Uttarakhand, India. S. Afr. J. Bot. 2023; 157, 151-8.
W Jamjumroon, N Tesna and S Sangrit. Threatened plants of Phu Wua-Phu Langka forest. Omega Printing, Bangkok, Thailand, 2015, p. 52.
N Supuntee, W Jamjumroon and N Tesna. Threatened plants of Phu Wua-Phu Langka forest. In: Proceedings of the Academic Conference and Academic Presentations, Thailand Forest Ecology Research Network: Ecology Knowledge for Sustainable Management, Phitsanulok, Thailand. 2015, p. 270.
JJ Doyle and JL Doyle. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987; 19, 11-5
CP Lau, L Ramsden and RM Saunders. Hybrid origin of “Bauhinia blakeana” (Leguminosae: Caesalpinioideae), inferred using morphological, reproductive, and molecular data. Am. J. Bot. 2005; 92, 525-33.
D Botstein, RL White, M Skolnick and RW Davis. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 1980; 32, 314-31.
A Prevost and MJ Wilkinson. A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor. Appl. Genet. 1999; 98, 107-12.
FC Yen. Population genetic analysis of co-dominant and dominant markers and quantitative traits. Belgian J. Bot. 1997; 129, 157-63.
R Peakall and PE Smouse. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 2012; 28, 2537-9.
R Frankham, JD Ballou and DA Briscoe. Introduction to conservation genetics. Cambridge University Press, Cambridge, United Kingdom, 2002.
FV Rossum and OJ Hardy. Guidelines for genetic monitoring of translocated plant populations. Conservat. Biol. 2022; 36, e13670.
Y Chen, C Tang, H Li, L Qian, S Luo and X Zhou. Analysis of affinity of Bauhinia variegata germplasm resources based on ISSR markers. Southwest Chin. J. Agr. Sci. 2020; 33, 26-31.
S Sraphet, P Sukawutthiya, N Srisawad, DR Smith and K Triwitayakorn. Evaluation of the genetic diversity of Bauhinia winitii, an endemic plant of Thailand, using microsatellite markers. Philippine J. Sci. 2021; 150, 557-69.
SCH Barrett and JR Kohn. Genetic and evolutionary consequences of small population size in plant: Implications for conservation. In: DA Falk and KE Holsinger (Eds.). Genetic and conservation of rare plants. Oxford University Press, Oxford, United Kingdom, 1991, p. 3-30.
TM Faske, AC Agneray, JP Jahner, LM Sheta, EA Leger and TL Parchman. Genomic and common garden approaches yield complementary results for quantifying environmental drivers of local adaptation in rubber rabbitbrush, a foundational Great Basin shrub. Evol. Appl. 2021; 14, 2881-900.
M Cai, Y Wen, K Uchiyama, Y Onuma and Y Tsumura. Population genetic diversity and structure of ancient tree populations of Cryptomeria japonica var. sinensis based on RAD-seq data. Forests 2020; 11, 1192.
CML Serrote, LRS Reiniger, KB Silva, SMDS Rabaiolli and CM Stefanel. Determining the polymorphism information content of a molecular marker. Gene 2020; 726, 144175.
W Powell, GC Machray and J Provan. Polymorphism revealed by simple sequence repeats. Trends Plant Sci. 1996; 1, 215-22.
RRA Giachino. Investigation of the genetic variation of anise (Pimpinella anisum L.) using RAPD and ISSR markers. Genet. Resour. Crop Evol. 2020; 67, 763-80.
J Hadian, A Karami, A Azizi and A Khadivi-Khub. Ubiquitous genetic diversity among and within wild populations of Satureja rechingeri assessed with ISSR markers. Plant Syst. Evol. 2015; 301, 923-30.
ALD Silva Júnior, RLR Cabral, L Sartori, FDD Miranda, MVW Caldeira, SO Moreira, TD Oliveira Godinho and FSD Oliveira. Molecular markers applied to the genetic characterization of Dalbergia nigra: Implications for conservation and management. Trees 2022; 36, 1539-57
L Liao, QS Guo, ZY Wang, L Liu and ZB Zhu. Genetic diversity analysis of Prunella vulgaris in China using ISSR and SRAP markers. Biochem. Syst. Ecol. 2012; 45, 209-17.
S Saboori, Z Noormohammadi, M Sheidai and S Marashi. SCoT molecular markers and genetic fingerprinting of date palm (Phoenix dactylifera L.) cultivars. Genet. Resour. Crop Evol. 2020; 67, 73-82.
SG Jian, T Tang, Y Zhong and SH Shi. Conservation genetics of Heritiera littoralis (Sterculiaceae), a threatened mangrove in China, based on AFLP and ISSR markers. Biochem. Syst. Ecol. 2010; 38, 924-30.
BR Subedee, GR Tripathi, NN Munankarmi and RP Chaudhary. Genetic diversity in populations of Girardinia diversifolia from Nepal Himalaya using ISSR markers. Ecol. Genet. Genom. 2022; 23, e100120.
E Pons, A Navarro, P Ollitrault and L Pena. Pollen competition as a reproductive isolation barrier represses transgene flow between compatible and co-flowering citrus genotypes. PLoS One 2022; 6, e25810
FM Sereshkeh, A Azizi and A Noroozisharaf. Structure of genetic diversity among and within populations of the endemic Iranian plant Dracocephalum kotschyi. Horticult. Environ. Biotechnol. 2019; 60, 767-77.
S Li-jian and S Nuchit. The characteristics, formation and exploration progress of the potash deposits on the Khorat Plateau, Thailand and Laos, Southeast Asia. Chin. Geol. 2020; 3, 67-82
AK Bishoyi, VV Pillai, KA Geetha and S Maiti. Assessment of genetic diversity in Clitoria ternatea populations from different parts of India by RAPD and ISSR markers. Genet. Resour. Crop Evol. 2014; 61, 1597-609.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2023 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






