Characterization and Expression Analysis of HbC3H66: Implications for Transcriptional Regulation in Rubber Biosynthesis and Abiotic Stress Responses in Hevea brasiliensis
DOI:
https://doi.org/10.48048/tis.2023.6835Keywords:
CCCH-type zinc finger protein, Hevea brasiliensis, Water deficit, Transcription factor, Ethylene, Drought stress, Salt stressAbstract
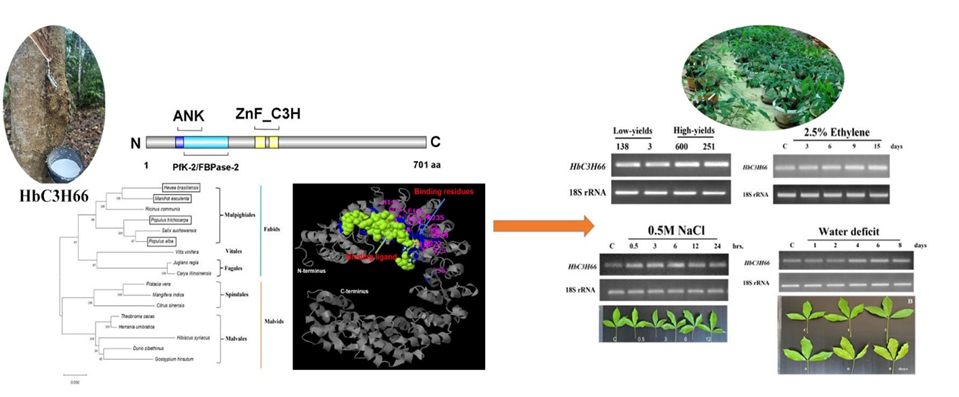
CCCH-type zinc-finger protein (ZnF_CCCH) family is one of the most important transcription factors (TFs) linked to various biotic and abiotic stressors and physiological and developmental processes in plants. This study aims to clone and characterise the latex gene expression patterns, evolution and characteristics of the Hevea brasiliensis ZnF_CCCH domain-containing protein 66 (HbC3H66) gene. These results showed that the open reading frame (ORF) of the HbC3H66 gene was 2,106 bp, encoding 701 aa and the calculated molecular weight of the encoded protein was 76.52 kDa. The N-terminal region of HbC3H66 contains 2 ANK repeats in, PfK_2/FBPase-2 and 2 type zinc finger motifs. Phylogenetic analysis showed that the C3H66 amino acid from Hevea and other plant C3H66 were clustered into 1 group and could be used for evolutionary analysis. Semi-quantitative RT-PCR (sqRT-PCR) revealed that the HbC3H66 mRNA was abundance in high-yielding Hevea clones (RRIT251 and RRIM600) and treatment with 2.5 % of ethephon (Eth) induced HbC3H66 mRNA expression in the latex of 15 years old Hevea trees. The HbC3H66 gene was induced by water deficit and 0.5 M NaCl in the latex of 3-month-old Hevea. In conclusion, the HbC3H66 protein may play a role in DNA-binding transcriptional regulation in NR pathways. Which would provide a theoretical basis for understanding the evolution and functions of the Hevea C3H66 gene in rubber biosynthesis.
HIGHLIGHTS
- CCCH-type zinc-finger proteins (ZnF_CCCH) are essential transcription factors (TFs) for protein binding activity in plant growth and stress tolerance.
- Hevea brasiliensis zinc finger CCCH domain-containing protein 66 (HbC3H66) gene was clone and characterized.
- HbC3H66 contains 2 ANK repeat in, Pfk_2/FBPase-2 and 2 type zinc finger motifs.
- Semi-quantitative RT-PCR revealed that HbC3H66 gene was abundance in high-yielding Hevea clones (RRIT251 and RRIM600) and ethylene was induced HbC3H66 mRNA expression in the latex of Hevea The HbC3H66 gene was induced by water deficit and 0.5 M NaCl in the latex of seedling of Hevea trees.
- The HbC3H66 protein play a role in DNA-binding transcriptional regulation in NR pathway.
GRAPHICAL ABSTRACT
Downloads
References
S Cherian, SB Ryu and K Cornish. Natural rubber biosynthesis in plants, the rubber transferase complex, and metabolic engineering progress and prospects. Plant Biotechnol. J. 2019; 17, 2041-61.
X Men, F Wang, GQ Chen, HB Zhang and M Xian. Biosynthesis of natural rubber: Current state and perspectives. Int. J. Mol. Sci. 2019; 20, 50.
T Yamaguchi, Y Kurihara, Y Makita, EO Kurihara, A Kageyama, E Osada, S Shimad, H Tsuchida, H Shimada and M Matsui. Regulatory potential of bHLH-type transcription factors on the road to rubber biosynthesis in Hevea brasiliensis. Plants 2020; 674, 2-13.
D Guo, HY Yi, HL Li, C Liu, ZP Yang and QS Peng. Molecular characterization of HbCZF1, a Hevea brasiliensis CCCH-type zinc finger protein that regulates hmg1. Plant Cell Rep. 2015; 34, 1569-78.
D Guo, HL Li, JH Zhu, Y Wang and SQ Peng. HbTGA1, a TGA transcription factor from Hevea brasiliensis, regulates the expression of multiple natural rubber biosynthesis genes. Front. Plant Sci. 2022; 13, 909098.
G Han, Z Qiao, Y Li, C Wang and B Wang. The roles of CCCH zinc-finger proteins in plant abiotic stress tolerance. Int. J. Mol. Sci. 2021; 22, 8327.
PJ Blackshear. Tristetraprolin and other CCCH tandem zinc finger proteins in the regulation of mRNA turnover. Biochem. Soc. Trans. 2002; 30, 945-52.
HY Seok, H Bae, T Kim, MMM Syed, LV Nguyen, SY Lee and YH Moo. Non-TZF protein AtC3H59/ZFWD3 is involved in seed germination, seedling development, and seed development, interacting with PPPDE family protein Desi1 in Arabidopsis. Int. J. Mol. Sci. 2021; 22, 4738.
H Seok, LV Nguyen, H Park, VN Tarte, J Ha, S Lee and Y Moon. Arabidopsis non-TZF gene AtC3H17 functions as a positive regulator in salt stress response. Biochem. Biophys. Res. Comm. 2018; 498, 954-9.
S Bogamuwa and JC Jang. Plant tandem CCCH zinc finger proteins interact with ABA, drought, and stress response regulators in processing-bodies and stress granules. PLos One 2016; 11, e0151574.
KK Vinod, JR Meenattoor, PM Priyadarshan, J Pothen, D Chaudhuri, AK Krishnakumar, MR Sethuraj and SN Potty. Early performance of some clones of Hevea brasiliensis in Tripura. Indian J. Nat. Rubber Res. 1996; 9, 123-9.
Y Zhang, J Leclercq and P Montoro. Reactive oxygen species in Hevea brasiliensis latex and relevance to tapping panel dryness. Tree Physiol. 2016; 37, 261-9.
P Suwanmanee, N Sirinupong and W Suvachittanont. Regulation of 3-hydroxy-3-methylglutaryl-CoA synthase and 3-hydroxy-3-methylglutaryl-CoA reductase and rubber biosynthesis of Hevea brasiliensis (B.H.K.) Mull. Arg. In: TJ Bach and M Rohmer (Eds.). Isoprenoid synthesis in plants and microorganism. Springer, New York, 2013, p. 315-27.
P Suwanmanee, S chocknukul, N Sirinupong, T Phattamanont, C Chantrapradist and P Jewtragoon. Selection of gene marker for identification of high latex yielding Hevea clone. In: Proceedings of the 19th Thaksin University annual conference, Songkhla, Thailand. 2009, p. 304-11.
E Gasteiger, C Hoogland, A Gattiker, S Duvaud, MR Wilkins, RD Appel and A Bairoch. Protein identification and analysis tools on the ExPASy server. In: JM Walker (Ed.). The proteomics protocols handbook. Humana Press, New Jersey, 2005, p. 571-607.
TA Hall. Bioedit: A user-friendly biological sequence alignment editor analysis program for windows 95/98/NT. Nucleic Acids Symp. 1999; 41, 95-8.
K Tamura, G Stecher and S Kumar. MEGS11: Molecular evolutionary genetic analysis version 11. Mol. Biol. Evol. 2021; 38, 3022-7.
I Lectunic, S Khedka and P Bork. SMART: Recent updates, new developments, and status in 2020. Nucleic Acids Res. 2021; 49, 458-60.
GE Crooke, G Hon, JM Chandonia and SE Brenner. WebLogo: A sequence logo generator. Genome Res. 2004; 14, 1188-90.
W Liu, Y Xie, J Ma, X Luo, P Nie, Z Zuo, U Lahrmann, Q Zhao, Y Zheng, Y Zhao, Y Xue and R Ren. IBS: An illustrator for the presentation and visualization of biological sequences. Bioinformatics 2015; 31, 3359-61.
S Kosuki, M Hasebe, M Tomita and H Yanagawa. Systematic identification of yeast cell cycle-dependent nucleocytoplasmic shutting protein by prediction of composite motifs. Proc. Natl. Acad. Sci. Unit. States Am. 2009; 106, 10171-6.
W Zheng, W Zhang, Y Li, R Pearce and EW Bell. Folding non-homology proteins by coupling deep-learning contact map with I-TASSER assembly simulations. Cell Rep. Meth. 2021; 1, 100014.
WL Delano, The PyMOL molecular graphics system, Available at: http://www.Pymol.org, accessed January 2022.
VB Chen, WB Arendall, JJ Headd, DA Keedy, RM Immormino, GJ Kapral, LW Murray, JS Richardson and DC Richardson. MolProbity: All-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 2010; 66, 12-21.
Z Xie, W Lin, G Yu, Q Cheng, B Xu and B Huang. Improved cold tolerance in switchgrass by a novel CCCH-type zinc finger transcription factor gene, PvC3H72, associated with ICEI-CBF-COR regulation and ABA-responsive genes. Biotechnol. Biofuels 2019; 12, 224.
J Sun, H Jiang, Y Xu, H Li, X Wu and C Li. The CCCH-type zinc finger proteins AtSZF1 and AtSZF2 regulate salt stress response in Arabidopsis. Plant Cell Physiol. 2007; 48, 1148-58.
MC Pomeranz, C Hah, P Lin, SG Kang, JJ Finer, PJ Blackshear and JC Jang. The Arabidopsis tandem zinc finger protein AtTZF1 traffics between the nucleus and cytoplasmic foci and binds both DNA and RNA. Plant Physiol. 2010; 152, 151-65.
ZJ Lorkovic and A Barta. Genome analysis: RNA recognition motif (RRM) and K homology (KH) domain RNA binding proteins from the flowering plant Arabidopsis thaliana. Nucleic Acids Res. 2002; 30, 623-35.
MH Rider, L Bertrand, D Vertommen, PA Michels, GG Rousseau and L Hue. 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase: head-to-head with a bifunctional enzyme that controls glycolysis. Biochem. J. 2004; 381, 561-78.
EB Tikhonova, AS Ethayathulla, Y Su, P Harharan, S Xie and L Guan. A transcription blocker isolated from a designed repeat protein combinatorial library by in vivo functional screen. Sci. Rep. 2014; 5, 8070.
C Mi, L Zhang, G Huang, G Shao, F Yang, X You, MQ Dong, S Sun and SF Sui. Structural basis for assembly of TRAPPII complex and specific activation of GTPase Ypt31/32. Sci. Adv. 2022; 8, eabi5603.
P Barraud, S Banerjee, WI Mohamed, MF Jantsh and FHT Allain. A bimodular nuclear localization signal assembled via an extended double-strand RNA-binding domain acts as an RNA sensing signal for transpotin1. Proc. Natl. Acad. Sci. Unit. States Am. 2014; 111, E1853-E1861.
B Pi, J Pan, M Xio, X Hu, L Zhang, M Chen, B Liu, Y Ruan and Y Huang. Systematic analysis of CCCH zinc finger family in Brasica napus showed that BnRR-TZFs are involved in stress resistance. BMC Plant Biol. 2021; 21, 555.
S Xin, Y Hua, J Li, X Dai, X Yang, J Udayabhanu, H Huang and T Huang. Comparative analysis of latex transcriptomes reveals the potential mechanisms underlying rubber molecular weight variations between the Hevea brasiliensis clones RRIM600 and Reyan 7-33-97. BMC Plant Biol. 2021; 21, 244.
D Li, L Hao, H Liu, M Zhao, Z Deng, Y Li, R Zeng and W Tian. Next-generation sequencing, assembly, and comparative analyses of the latex transcriptomes from two elite Hevea brasiliensis varieties. Tree Genet. Genomes 2015; 11, 98.
J Santos, LE Oliveira, VT Coelho, G Lopes, T Souza, AC Porto, J Lira, R Massote, C Rocha and MP Gomes. Performance of Hevea brasiliensis under drought conditions on osmoregulation and antioxidant activity through evaluation of vacuolar invertase and reducing sugars. Plant Sci. Today 2021; 8, 312-23.
TP Liu, YF Zhuang, XL Guo and YJ Li. Molecular mechanism of ethylene stimulation of latex yield in rubber tree (Hevea brasiliensis) revealed by de novo sequencing and transcriptome analysis. BMC Genom. 2016; 17, 257.
B Pi, X He, HY Ruan, JC Jang and Y Huang. Genome-wide analysis and stress-responsive expression of CCCH zinc finger family genes in Brassica rapa. BMC Plant Biol. 2018; 18, 373.
N Chevalier, L Bertrand, MH Rider, FR Opperdoes, DJ Ringden and PAM Michels. 6-phosphofructo-2-kinase and fructose-2,6-bisphosphatase in Trypanosomatidae. FEBS J. 2005; 272, 3542-60.
T Banzai, N Hanagata, Z Dubinsky and I Karube. Fructose-2,6-bisphosphate contents were increased in response to salt, water, and osmotic stress in leaves of Bruguiera gymnorrhiza by differential changes in the activity of the bifunctional enzyme 6-phosphofructo-2-kinase/fructose-2,6-bisphosphate 2 phosphatase. Plant Mol. Biol. 2003; 53, 51-9.
TH Nielson, JH Ruang and D Villadson. Fructose-2,6-bisphosphate: A traffic signal in plant metabolism. Trends Plant Sci. 2004; 9, 557-63.
X Guo and K Chong. Cold signalling in plants: Insights into mechanisms and regulation. J. Integr. Plant Biol. 2018; 60, 745-56.
R Bagur and G Hajnóczky. Intracellular Ca2+ sensing: Role in calcium homeostasis and signaling. Mol. Cell 2017; 66, 780-8.
Y Aoki, S Takahashi, D Takayama, Y Ogata, N Sakurai, H Suzuki, K Assawatreeratanakul, D Wititsuwannakul, R Wititsuwannakul, D Shibata, T Koyama and T Nakayama. Identification of laticifer-specific genes and their promoter regions from a natural rubber producing plant Hevea brasiliensis. Plant Sci. 2014; 225, 1-8.

Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2023 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






