Species Composition and Hydrolase Enzyme (EC.3) Activity of Fungi Isolated from Thasala Mangroves, Nakhon Si Thammarat Province, Southern Thailand
DOI:
https://doi.org/10.48048/tis.2022.6172Keywords:
Hydrolase enzyme, Manglicolous fungi, Species compositionAbstract
In the present study, we aimed to investigate diversity of manglicolous and endobiotic fungi associated with mangrove plants from Thasala mangroves, Nakhon Si Thammarat province, southern Thailand. Extracellular hydrolase enzymes (EC.3) including amylase, cellulase, lipase, protease, and pectinase from isolated fungi were evaluated. A total of 31 obligate marine fungal species was recorded from various Thasala mangroves sites. These included 24 ascomycetes (77.4 %), 1 mucoromycete (3.2 %), 3 basidiomycetes (9.7 %), 4 asexual morphs (12.9 %), and tentatively identified fungi (9.7 %). The average percentage occurrence of the fungi recorded from each site ranged from 0.4 to 16%. At all sites, the common filamentous marine fungi were Cumulospora sp., Halorosellinia oceanica, Kallichroma tethys, Leptosphaeria sp., Rimora mangrovei, Lulworthia sp., Phoma sp., Halenospora varia, Halocyphina villosa, Nia vibrissa, and Sclerococcum haliotrephum. Also, the most frequent marine fungi include Eutypella naqsii, Sammeyersia grandispora, Marinosphaera mangrovei, Nemania maritima, and Verruculina enalia. Seven mangrove plants were selected for endophytic fungal isolation including Acanthus ebracteatus, Acrostichum aureum, Avicennia alba, Bruguiera cylindrica, Rhizophora apiculata, R. mucronata, and Sonneratia alba. The isolation rate (IR) and colonization rate (CR) of fungal endobiota varied for all plant studied. The isolation rate ranged from 24 - 53.3 %, while colonization rate varied from 29.3 - 61.3 %. The fungal isolates were screened for the production of hydrolase enzymes (EC.3), of which 37.2 % exhibited enzyme activities. The 16 out of 28 obligate marine fungi (57.1 %) and 13 out of 50 fungal endophytes (26 %) had enzyme production efficiency toward specific substrates or an enzymatic index (EI) higher than 1.4. None of the fungal isolates depicted lipase activity. Additionally, the environmental stressors (temperature, pH, salinity) affecting enzyme production were evaluated in order to discover potential candidates for industrial application.
HIGHLIGHTS
- Diversity of obligate marine fungal species was estimated from Thasala mangroves
- The obligate marine fungi and fungal endophytes had hydrolase enzyme (EC.3) production efficiency toward specific substrates or an enzymatic index (EI) higher than 1.4
- The temperature, pH, and salinity affecting enzyme production were evaluated in order to discover potential candidates for industrial application
GRAPHICAL ABSTRACT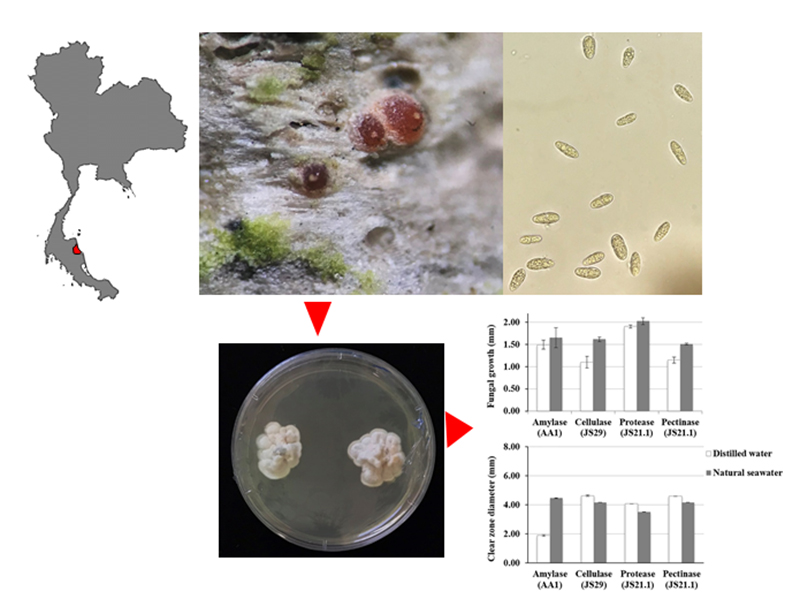
Downloads
References
WN Nakorn, W Chanchaw, P Penprapai and S Chanchaw. Diversity and evenness of mangrove trees in Thasala, Sichon and Pakpaneang District, Nakhon Si Thammarat Province, Thailand. Int. J. Agr. Tech. 2018; 14, 333-40.
J Kohlmeyer. Tropical marine fungi. Mar. Ecol. 1984; 5, 329-78.
J Koch. Some lignicolous marine fungi from Thailand, including two new species. Nord. J. Bot. 1986; 6, 497-9.
KD Hyde. Caryospora mangrovei sp. nov. and notes on marine fungi from Thailand. Trans. Mycol. Soc. Jpn. 1989; 30, 333-41.
KD Hyde. Ecology of tropical marine fungi. Hydrobiologia 1989; 178, 199-208.
KD Hyde. Aigialus striatispora sp. nov. from intertidal mangrove wood. Mycol. Res. 1992; 96, 1044-6.
KD Hyde. The genus Saccardoella from intertidal mangrove wood. Mycol. Res. 1992; 84, 803-10.
KD Hyde. Cryptosphaeria mangrovei sp. nov. (Diatrypaceae) from intertidal mangrove-wood. Trans. Mycol. Soc. Jpn. 1993; 34, 311-4.
KD Hyde. A Chalermpongse and T Boonthavikoon. Ecology of intertidal fungi at Ranong mangrove, Thailand. Trans. Mycol. Soc. Jpn.1990; 31, 17-27.
KD Hyde, A Chalermpongse and T Boonthavikoon. The distribution of intertidal fungi on Rhizophora apiculate. Hong Kong University Press, Hong Kong, 1993, p. 643-52.
EBG Jones, A Pilantanapak, I Chatmala, J Sakayaroj, S Phongpaichit and R Choeyklin. Thai marine fungal diversity. Songklanakarin J. Sci. Tech. 2006; 28, 687-708.
T Dethoup and L Manoch. Diversity of marine fungi in Eastern Thailand. Kasetsart J. Nat. Sci. 2009; 43, 100-6.
J Sakayaroj, O Supaphon, EBG Jones and S Phongpaichit. Diversity of higher marine fungi at Hat Khanom-Mu Ko Thale Tai National Park, Southern Thailand. Songklanakarin J. Sci. Tech. 2011; 33,15-22.
S Suetrong, S Preedanon, A Klaysuban, W Gundool, P Unagul, J Sakayaroj, W Promchu and T Sangtiean. Distribution and occurrence of manglicolous marine fungi from Eastern and Southern Thailand. Bot. Mar. 2017; 60, 503-14.
B Devadatha, EBG Jones, KL Pang, MA Abdel-Wahab, KD Hyde, S Sutreong, J Sakayaroj, AH Bahkali, M Calabon, VV Sarma and S Zhang. Occurrence and geographical distribution of mangrove fungi. Fungal Divers. 2021; 106, 137-227.
KD Hyde and EBG Jones. Intertidal mangrove fungi: Pedumispora gen. nov. (Diaporthales). Mycol. Res. 1992; 96, 78-80.
I Chatmala, J Sakayaroj, S Somrithipol and S Phongpaichit. Marine hyphomycetes of Thailand and Cummulospora varia sp. nov. Fungal Divers. 2004; 17, 1-9.
EBG Jones, I Chatmala and KL Pang. Two new genera isolated from marine habitats in Thailand: Pseudolignincola and Thalespora (Halosphaeriales, Ascomycota). Nova Hedwigia 2006; 83, 219-32.
T Hattori, J Sakayaroj, EBG Jones, S Suetrong, S Preedanon and A Klaysuban. Three species of Fulvifomes (Basidiomycota, Hymenochaetales) associated with rots on mangrove tree Xylocarpus granatum in Thailand. Mycoscience 2014; 55, 344-54.
S Preedanon, A Klaysuban, S Suetrong, W Promchoo, W Gundool, T Sangtiean and J Sakayaroj. Helicascus mangrovei sp. nov., a new intertidal mangrove fungus from Thailand. Mycoscience 2017; 58, 174-80.
S Chaeprasert, J Piapukiew, AJS Whalley and P Sihanonth. Endophytic fungi from mangrove plant species of Thailand: Their antimicrobial and anticancer potentials. Bot. Mar. 2010; 53, 555-64.
J Sakayaroj, S Preedanon, O Supaphon, EBG Jones and S Phongpaichit. Phylogenetic diversity of endophyte assemblages associated with the tropical seagrass Enhalus acoroides in Thailand. Fungal Divers. 2010; 42, 27-45.
J Buatong, S Phongpaichit, V Rukachaisirikul and J Sakayaroj. Antimicrobial activity of crude extracts from mangrove fungal endophytes. World J. Microbiol. Biotechnol. 2011; 27, 3005-3008.
M Doilom, IS Manawasinghe, R Jeewon, RS Jayawardena, S Tibpromma, S Hongsanan, W Meepol, S Lumyong, EBG Jones and KD Hyde. Can ITS sequence data identify fungal endobiotes from cultures? A case study from Rhizophora apiculata. Mycosphere 2017; 8, 1869-92.
P Supaphon, S Phongpaichit, J Sakayaroj, V Rukachaisirikul, N Kobmoo and JW Spatafora. Phylogenetic community structure of fungal endophytes in seagrass species. Bot. Mar. 2017; 60, 489-502.
K Mandyam and A Jumpponen. Seeking the elusive function of the root-colonising dark septate endophytic fungi. Stud. Mycol. 2005; 53, 173-89.
D Johnston-Monje and MN Raizada. Plant and endophyte relationships: Nutrient management. In: M Moo-Young (Ed.). Comprehensive biotechnology. 2nd ed. Academic Press, New Jersey, 2011, p. 713-27.
YW Choi, KD Hyde and WH Ho. Single spore isolation of fungi. Fungal Divers. 1999; 3, 29-38.
J Kohlmeyer and B Volkmann-Kohlmeyer. Illustrated key to the filamentous higher marine fungi. Bot. Mar.1991; 34, 1-61.
EBG Jones, J Sakayaroj, S Suetrong, S Somrithipol and KL Pang. Classification of marine Ascomycota, anamorphic taxa and Basidiomycota. Fungal Divers. 2009; 35, 1-203.
EBG Jones, S Suetrong, J Sakayaroj, AH Bahkali, MA Abdel-Wahab, T Boekhout and KL Pang. Classification of marine Ascomycota, Basidiomycota, Blastocladiomycota and Chytridiomycota. Fungal Divers. 2015; 3, 1-72.
TJ White, T Bruns, S Lee and J Taylor. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: MA Innis, DH Gelfand, JS Sninsky and TJ White (Eds.). PCR protocol: A guide to methods and applications. Academic Press, San Diego, California, 1990, p. 315-22.
S Landvik. Neolecta, a fruit-body-producing genus of the basal ascomycetes, as shown by SSU and LSU rDNA sequences. Mycol. Res. 1996; 100, 199-202.
T Hall. BioEdit 7.0.5.3. Department of Microbiology, North Carolina State University, Available at: http://www.mbio.ncsu.edu/BioEdit/ bioedit.html, accessed March 2021.
DL Swofford. PAUP*: Phylogenetic analysis using parsimony (*and other methods) version 4.0b10. Sinauer Associates, Sunderland, Massachusetts, 2002.
AE Magurran. Ecological diversity and its measurement. Springer, Dordrecht, Netherlands, 1998, p. 7-45.
O Petrini, J Stone and FE Carroll. Endophytic fungi in evergreen shrubs in western Oregon: A preliminary study. Can. J. Bot.1982; 60, 789-96.
EBG Jones. Marine fungi: Some factors influencing biodiversity. Fungal Divers. 2000; 4, 53-73.
A Pilantanapak, EBG Jones and RA Eaton. Marine fungi on Nypa fruticans in Thailand. Bot. Mar. 2005; 48, 365-73.
J Kohlmeyer and B Volkmann-Kohlmeyer. Biogeographic observations on Pacific marine fungi. Mycologia 1993; 85, 337-46.
EBG Jones, KL Pang, MA Abdel-Wahab, B Scholz, KD Hyde, T Boekhout, R Ebel, M Rateb, L Henderson, J Sakayaroj, S Suetrong, M Dayarathne, V Kumar, S Raghukumar, KR Sridhar, A Bahkali, F Gleason and C Norphanphoun. An online resource for marine fungi. Fungal Divers. 2019; 96, 347-433.
XK Xing, J Chen, MJ Xu, WH Lin and SX Guo. Fungal endophytes associated with Sonneratia (Sonneratiaceae) mangrove plants on the south coast of China. Forest Pathol. 2011; 41, 334-40.
AR Liu, SC Chen, WJ Jin, PY Zhao, R Jeewon and T Xu. Host specificity of endophytic Pestalotiopsis populations in mangrove plant species of South China. Afr. J. Microbiol. Res. 2012; 6, 6262-9.
Y Sun, Q Wang, XD Lu, I Okane and M Kakishima. Endophytic fungal community in stems and leaves of plants from desert areas in China. Mycol. Prog. 2012; 11, 781-90.
JL Li, X Sun, L Chen and LD Guo. Community structure of endophytic fungi of four mangrove species in Southern China. Mycology 2016; 7, 180-90.
KL Pang, LLP Vrijmoed, TK Goh, N Plaingam and EBG Jones. Fungal endophytes associated with Kandelia candel (Rhizophoraceae) in Mai Po Nature Reserve, Hong Kong. Bot. Mar. 2008; 51, 171-8.
A Zuccaro and JI Mitchell. Fungal communities of seaweeds. In: J Dighton, JF White and P Oudemans (Eds.). The fungal community. CRC Press, New York, 2005, p. 533-79.
A Zuccaro, B Schulz and JI Mitchell. Molecular detection of ascomycetes associated with Fucus serratus. Mycol. Res. 2003; 107, 1451-66.
A Zuccaro, CL Schoch, JW Spatafora, J Kohlmeyer, S Draeger and JI Mitchell. Detection and identification of fungi intimately associated with the brown seaweed Fucus serratus. Appl. Environ. Microbiol. 2008; 74, 931-41.
JE Taylor, KD Hyde and EBG Jones. Endophytic fungi associated with the temperate palm, Trachycarpus fortunei, within and outside its natural geographic range. New Phytologist 1999; 142, 335-46.
AE Arnold. Understanding the diversity of foliar endophytic fungi: Progress, challenges, and frontiers. Fungal Biol. Rev. 2007; 21, 51-66.
K Saikkonen. Forest structure and fungal endophytes. Fungal Biol. Rev. 2007; 21, 67-74.
TN Sieber. Endophytic fungi in forest trees: Are they mutualists? Fungal Biol. Rev. 2007; 21, 75-89.
VVC Bucher, KD Hyde, SB Pointing and CA Reddy. Production of wood decay enzymes, mass loss and lignin solubilization in wood by marine ascomycetes and their anamorphs. Fungal Divers. 2004; 15, 1-14.
N Velmurugan and YS Lee. 21 enzymes from marine fungi: Current research and future prospects. In: EBG Jones and KL Pang (Eds.). Marine fungi and fungal-like organisms. Walter de Gruyter, Berlin, Germany, 2012, p. 441-74.
JDP Bezerra, MGS Santos, VM Svedese, DMM Lima, MJS Fernandes, LM Paiva and CM Souza-Motta. Richness of endophytic fungi isolated from Opuntia ficus-indica Mill. (Cactaceae) and preliminary screening for enzyme production. World J. Microbiol Biotechnol. 2012; 28, 1989-95.
AB Gunjal, NN Patil and SS Shinde. Enzymes in degradation of the lignocellulosic wastes. Springer Nature, Switzerland, 2020, p. 15-113.
SB Pointing, JA Buswell, EBG Jones and LLP Vrijmoed. Extracellular cellulolytic enzyme profiles of five lignicolous mangrove fungi. Mycol. Res. 1999; 103, 696-700.
G Schimpfhauser and HP Molitoris. Enzyme activities of monokaryotic and dikaryotic strains of the marine Basidiomycete Nia vibrissa. Kieler Meeresforsch. Sonderh. 1991; 8, 361-8.
S Rohrmann and HP Molitoris. Screening for wood decay enzymes in marine fungi. Can. J. Bot. 1992; 70, 2116-23.
C Raghukumar, S Raghukumar, A Chinnaraj, D Chandramohan, TM D’Souza and CA Reddy. Laccase and other lignocellulose modifying enzymes of marine fungi isolated from the coast of India. Bot. Mar. 1994; 37, 512-23.
J Kohlmeyer and B Volkmann-Kohlmeyer. Observations on Halonectria and Kallichroma gen. nov. Mycol. Res. 1993; 97, 753-61.
S Rohrmann and HP Molitoris. Screening for wood-degrading enzymes in marine fungi. Can. J. Bot. 2011; 70, 2116-23.
GL Maria, KR Sridhar and NS Raviraja. Antimicrobial and enzyme activity of mangrove endophytic fungi of southwest coast of India. J. Agr. Tech. 2005; 1, 67-80.
V Kumaresan and TS Suryanarayanan. Endophyte assemblage in young, mature and senescent leaves of Rhizophora apiculata: Evidence for the role of endophytes in mangrove litter degradation. Fungal Divers. 2002; 9, 81-91.
S Lee, MS Park, H Lee, JJ Kim, JA Eimes and YW Lim. Fungal diversity and enzyme activity associated with the macroalgae, Agarum clathratum. Mycobiology 2019; 47, 50-8.
V Cuomo, S Pagano, MA Pecorella and P Parascandola. Evidence of the active role of ligno-cellulosic enzymes of marine fungi in degradation of Posidonia oceanica leaves. Biochem. Systemat. Ecol. 1987; 15, 635-7.
S Damare, P Singh and S Raghukumar. Biotechnology of marine fungi. In: C Raghukumar (Ed.). Biology of marine fungi. Springer Science & Business Media, Berlin, Germany, 2012, p. 277-97.
M Venkatachalam, L Gérard, C Milhau, F Vinale, L Dufossé and M Fouillaud. Salinity and temperature influence growth and pigment production in the marine-derived fungal strain Talaromyces albobiverticillius 30548. Microorganisms 2019; 7, 10.
T Zangirolami, M Carlsen, J Nielsen and SB Jørgensen. Growth and enzyme production during continuous cultures of a high amylase-producing variant of Aspergillus oryzae. Braz. J. Chem. Eng. 2002; 19, 55-68.
SP Gautam, PS Bundela, AK Pandey, J Khan, MK Awasthi and S Sarsaiya. Optimization for the production of cellulase enzyme from municipal solid waste residue by two novel cellulolytic fungi. Biotechnol. Res. Int. 2011; 2011, 810425.
S Manivannan and K Kandasamy. Alkaline protease production by Penicillium fellutanum isolated from mangrove sediment. Int. J. Biol. Chem. 2007; 1, 98-103.
M Lobo, K Sridhar and R Seetharam. Antimicrobial and enzyme activity of mangrove endophytic fungi of southwest coast of India. J. Agr. Tech. 2005; 1, 67-80.
C Muthulakshmi, G Duraisamy, DG Kumar, G Ravikumar, M Kalaiselvi and C Uma, C. Production, purification and characterization of protease by Aspergillus flavus under solid state fermentation. Jordan J. Biol. Sci. 2011; 4, 137-48.
DB Pedrolli and EC Carmona. Pectin lyase from Aspergillus giganteus: Comparative study of productivity of submerged fermentation on citrus pectin and orange waste. Appl. Biochem. Microbiol. 2009; 45, 677-83.
R Ketipally and R Ram. Optimization of pectinase production by Aspergillus oryzae RR 103. Curr. Agr. Res. 2018; 6, 37-44.
C Raghukumar, U Muraleedharan, VR Gaud and R Mishra. Xylanase of marine fungi of potential use for biobleaching of paper pulp. J. Ind. Microbiol. Biotechnol. 2004; 31, 433-41.
SB Pointing and KD Hyde. Lignocellulose‐degrading marine fungi. Biofouling 2000; 15, 221-9.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.







