In Silico Screening of Phytochemicals Targeting SmdCD of Streptococcus mutans using Molecular Docking Approach
DOI:
https://doi.org/10.48048/tis.2023.6036Keywords:
Dental caries, Streptococcus mutans, Medicinal plant, Drug-likeness rule, Virtual screeningAbstract
Introduction: Dental caries is a chronic infectious disease caused by bacteria forming a biofilm formation on the tooth surface. Drug-resistant bacteria Streptococcus mutans (S. mutans) pose a vital public health issue. To overcome this, the development of effective drugs with a novel mechanism of action is very important. Drug repurposing is considered a viable alternative approach to overcome the above issue. In the present study, we have attempted to select unique and traditional source used as in traditional medicine. To continue this effort, in this study we used phytochemical of selected to find an effective cure for dental caries. Materials and methods: Inhibition of S. mutans deoxycytidylate deaminases (SmdCDs) is a promising drug development strategy against S. mutans, responsible for biofilm formation. Since no effective drugs are available against S. mutans, we conducted a virtual screening of phytochemicals to find novel compounds against this bacterium. Hence, we created a library of 871 phytochemicals from 13 plants with reported antimicrobial activity. The phytochemical library was subjected to virtual screening against the SmdCD target. Results and discussion: Traditionally, many cultures use chewing sticks for oral hygiene maintenance. In the present work, out of 871 phytochemicals, 211 phytochemicals showed the most druggable substance with zero violation from any of the drug-likeness rules. Further, the binding energy indicates the affinity of adhesion of protein structure docked with 2 hit potential herbal compounds of which Cyclocurcumin and Androsta-1,4,6-triene-3,17-dione showed the best docking with SmdCD. Conclusions: Based on the binding energy score, we suggest that hit compounds can be used for dental caries and further could be developed as an organic mouthwash.
HIGHLIGHTS
- Streptococcus mutans is common in the human oral cavity and contributes significantly to tooth decay
- mutans deoxycytidylate deaminases (SmdCDs) are responsible for biofilm formation causing drug resistance
- SmdCDs can be used to develop effective drug against mutans, responsible for biofilm formation
- Screening phytochemicals may result potential compounds which can be used to develop new drugs against mutans
GRAPHICAL ABSTRACT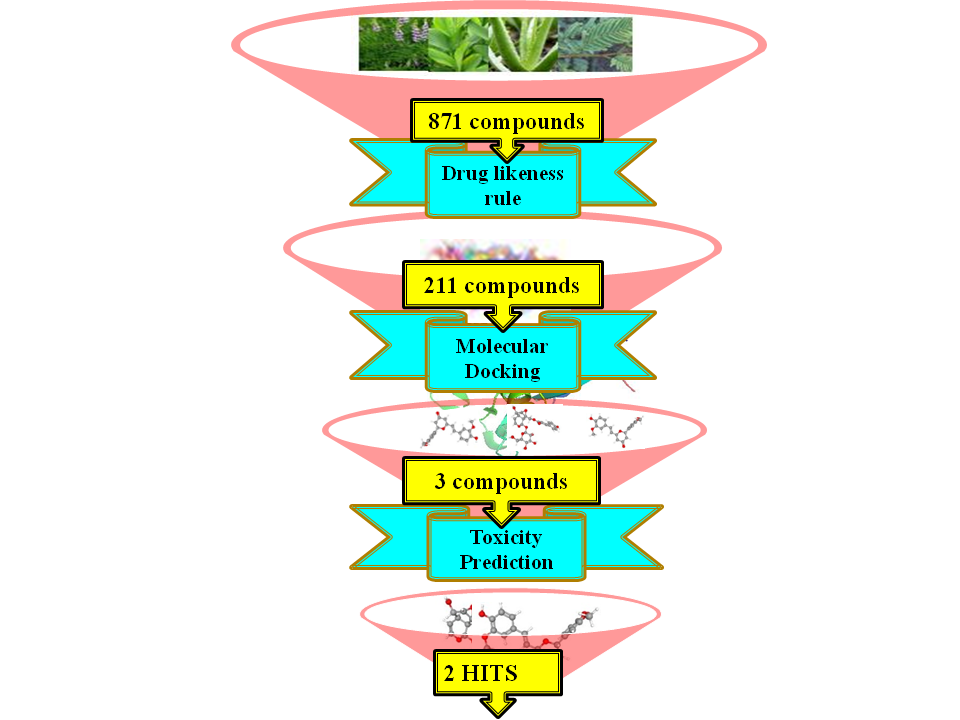
Downloads
References
L Gao, T Xu, G Huang, S Jiang, Y Gu and F Chen. Oral microbiomes: More and more importance in oral cavity and whole body. Protein Cell 2018; 9, 488-500.
J Reibel. Tobacco and oral diseases. Med. Princ. Pract. 2003; 12, 22-32.
RPJR Palmer, N Chalmers, A Rickard and P Kolenbrander. Community development in bacterial biofilms of the oral cavity. Microsc. Microanal. 2008; 14, 1554-55.
WH Bowen and H Koo. Biology of streptococcus mutans-derived glucosyltransferases: Role in extracellular matrix formation of cariogenic biofilms. Caries Res. 2011; 45, 69-86.
PD Marsh. Microbiology of dental plaque biofilms and their role in oral health and caries. Dent. Clin. 2010; 54, 441-54.
MM Abdel-Aziz, T Emam and MM Raafat. Hindering of cariogenic streptococcus mutans biofilm by fatty acid array derived from an endophytic arthrographis kalrae strain. Biomolecules 2020; 10, 811.
P Moynihan. Sugars and dental caries: Evidence for setting a recommended threshold for intake. Adv. Nutr. 2016; 7, 149-56.
A Algburi, N Comito, D Kashtanov, LMT Dicks and ML Chikindas. Control of biofilm formation: Antibiotics and beyond. Appl. Environ. Microbiol. 2017; 83, e02508-16.
SK Rath and M Singh. Comparative clinical and microbiological efficacy of mouthwashes containing 0.2 % and 0.12 % chlorhexidine. Dent. Res. J. 2013; 10, 364-9.
CG Jones. Chlorhexidine: Is it still the gold standard? Periodontol 2000; 15, 55-62.
CT Cabral and MH Fernandes. In vitro comparison of chlorhexidine and povidone-iodine on the long-term proliferation and functional activity of human alveolar bone cells. Clin. Oral Investig. 2007; 11, 155-64.
DA Ribeiro, AP Bazo, CADS Franchi, MEA Marques and DMF Salvadori. Chlorhexidine induces DNA damage in rat peripheral leukocytes and oral mucosal cells. J. Periodontal Res. 2004; 39, 358-61.
S Purkayastha and P Dahiya. Phytochemical screening and antimicrobial activity of some medicinal plants against multi-drug resistant bacteria from clinical isolates. Indian J. Pharmaceut. Sci. 2012; 74, 443.
B Josh, SK Panda, RS Jouneghani, M Liu, N Parajuli, P Leyssen, J Neyts and W Luyten. Antibacterial, antifungal, antiviral, and anthelmintic activities of medicinal plants of nepal selected based on ethnobotanical evidence. Evid. Base. Compl. Alternative Med. 2020; 2020, 1-14.
Z Ren, T Cui, J Zeng, L Chen, W Zhang, X Xu, L Cheng, M Li, J Li, X Zhou and Y Li. Molecule targeting glucosyltransferase inhibits streptococcus mutans biofilm formation and virulence. Antimicrob. Agents Chemother. 2016; 60, 126-35.
B Hernandez-Santiago, L Placidi, E Cretton-Scott, A Faraj, EG Bridges, ML Bryant, J Rodriguez-Orengo, JL Imbach, G Gosselin, C Pierra, D Dukhan and JP Sommadossi. Pharmacology of β-l-thymidine and β-l-2′-deoxycytidine in HepG2 cells and primary human hepatocytes: Relevance to chemotherapeutic efficacy against hepatitis B virus. Antimicrob. Agents Chemother. 2002; 46, 1728-33.
S Vellappally, DD Divakar, AAA Kheraif, R Ramakrishnaiah, A Alqahtani, MHN Dalati, S Anil, AA Khan and PRH Varma. Occurrence of vancomycin-resistant Staphylococcus aureus in the oral cavity of patients with dental caries. Acta Microbiol. Immunol. Hung. 2017; 64, 343-51.
Y Yang, Y Zhou, X Liu and J Tan. Health services provision of 48 public tertiary dental hospitals during the COVID-19 epidemic in China. Clin. Oral Investig. 2020; 24, 1861-4.
A Dziedzic. Special care dentistry and COVID-19 outbreak: What lesson should we learn? Dent. J. 2020; 8, 46.
C Niu and ES Gilbert. Colorimetric method for identifying plant essential oil components that affect biofilm formation and structure. Appl. Environ. Microbiol. 2004; 70, 6951-6.
L Lu, W Hu, Z Tian, D Yuan, G Yi, Y Zhou, Q Cheng, J Zhu and M Li. Developing natural products as potential anti-biofilm agents. Chin. Med. 2019; 14, 11.
S Bhandari, K Khadayat, S Poudel, S Shrestha, R Shrestha, P Devkota, S Khanal and BP Marasini. Phytochemical analysis of medicinal plants of Nepal and their antibacterial and antibiofilm activities against uropathogenic Escherichia coli. BMC Compl. Med. Ther. 2021; 21, 116.
NM O’Boyle, M Banck, CA James, C Morley, T Vandermeersch and GR Hutchison. Open babel: An open chemical toolbox. J. Cheminformatics 2011; 3, 33.
RC Lua and O Lichtarge. PyETV: A PyMOL evolutionary trace viewer to analyze functional site predictions in protein complexes. Bioinformatics 2010; 26, 2981-82.
O Trott and AJ Olson. AutoDock vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2009; 31, 455-61.
A Daina, O Michielin and V Zoete. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017; 7, 42717.
WJ Egan, KM Merz and JJ Baldwin. Prediction of drug absorption using multivariate statistics. J. Med. Chem. 2000; 43, 3867-77.
CA Lipinski. Drug-like properties and the causes of poor solubility and poor permeability. J. Pharmacol. Toxicol. Meth. 2000; 44, 235-49.
T Sander, J Freyss, MV Korff, JR Reich and C Rufener. OSIRIS, an entirely in-house developed drug discovery informatics system. J. Chem. Inform. Model. 2009; 49, 232-46.
H Yang, C Lou, L Sun, J Li, Y Cai, Z Wang, W Li, G Liu and Y Tang. admetSAR 2.0: Web-service for prediction and optimization of chemical ADMET properties. Bioinformatics 2019; 35, 1067-9.
AC Wallace, RA Laskowski and JM Thornton. LIGPLOT: A program to generate schematic diagrams of protein-ligand interactions. Protein Eng. Des. Sel. 1995; 8, 127-34.
AK Ghose, VN Viswanadhan and JJ Wendoloski. A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. a qualitative and quantitative characterization of known drug databases. ACS Combin. Sci. 1999; 1, 55-68.
F Darvas, G Keseru, A Papp, G Dorman, L Urge and P Krajcsi. In silico and ex silico ADME approaches for drug discovery. Curr. Top. Med. Chem. 2002; 2, 1287-304.
JA DiMasi, RW Hansen and HG Grabowski. The price of innovation: New estimates of drug development costs. J. Health Econ. 2003; 22, 151-85.
H Pajouhesh and GR Lenz. Medicinal chemical properties of successful central nervous system drugs. NeuroRx 2005; 2, 541-53.
A Finch and P Pillans. P-glycoprotein and its role in drug-drug interactions. Aust. Prescriber 2014; 37, 137-9.
F Cheng, Y Yu, J Shen, L Yang, W Li, G Liu, PW Lee and Y Tang. Classification of cytochrome P450 inhibitors and noninhibitors using combined classifiers. J. Chem. Inform. Model. 2011; 51, 996-1011.
EM Abdallah. Plants: An alternative source for antimicrobials. J. App. Pharmaceut. Sci. 2011; 1, 16-20.
H Luo, DF Liang, MY Bao, R Sun, YY Li, JZ Li, X Wang, KM Lu and JK Bao. In silico identification of potential inhibitors targeting Streptococcus mutans sortase A. Int. J. Oral Sci. 2017; 9, 53-62.
OM Vandeputte, M Kiendrebeogo, T Rasamiravaka, C Stévigny, P Duez, S Rajaonson, B Diallo, A Mol, M Baucher and ME Jaziri. The flavanone naringenin reduces the production of quorum sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. Microbiology 2011; 157, 2120-32.
SA Tamil, MG Dinesh, RS Satyan, B Chandrasekaran and C Rose. Leaf and seed extracts of Bixa orellana L. exerts anti-microbial activity against bacterial pathogens. J. App. Pharmaceut. Sci. 2011; 1, 116-20.
AS Chakotiya, R Chawla, P Thakur, A Tanwar, A Narula, SS Grover, R Goel, R Arora and RK Sharma. In vitro bactericidal activity of promising nutraceuticals for targeting multidrug resistant Pseudomonas aeruginosa. Nutrition 2016; 32, 890-7.
AS Chakotiya, A Tanwar, A Narula and RK Sharma. Alternative to antibiotics against Pseudomonas aeruginosa: Effects of Glycyrrhiza glabra on membrane permeability and inhibition of efflux activity and biofilm formation in Pseudomonas aeruginosa and its in vitro time-kill activity. Microb. Pathog. 2016; 98, 98-105.
G Spinetti, O Fortunato, A Caporali, S Shantikumar, M Marchetti, M Meloni, B Descamps, I Floris, E Sangalli, R Vono, E Faglia, C Specchia, G Pintus, P Madeddu and C Emanueli. MicroRNA-15a and microRNA-16 impair human circulating proangiogenic cell functions and are increased in the proangiogenic cells and serum of patients with critical limb ischemia. Circ. Res. 2013; 112, 335-46.
S Mishra and K Palanivelu. The effect of curcumin (turmeric) on Alzheimer’s disease: An overview. Ann. Indian Acad. Neurol. 2008; 11, 13.
N Goswami, MI Hussain and P Borah. Molecular dynamics approach to probe the antigenicity of PagN - an outer membrane protein of salmonella typhi. J. Biomol. Struct. Dynam. 2018; 36, 2131-46.
Y Nomin, S Charbonnier, L Miguet, N Potier, AV Dorsselaer, RA Atkinson, G Travé and B Kieffer. 1H and 15N resonance assignment, secondary structure and dynamic behaviour of the C-terminal domain of human papillomavirus oncoprotein E6. J. Biomol. NMR 2005; 31, 129-41.
CG Ullman, PI Haris, DA Galloway, VC Emery and SJ Perkins. Predicted α-helix/β-sheet secondary structures for the zinc-binding motifs of human papillomavirus E7 and E6 proteins by consensus prediction averaging and spectroscopic studies of E7. Biochem. J. 1996; 319, 229-39.

Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2022 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






