Analysis of Mutational Profiles of SARS-CoV-2 Structural and Non-Structural Proteins with Emphasis on Spike Protein Variants
DOI:
https://doi.org/10.48048/tis.2022.5768Keywords:
D614G spike mutation, MEGA X (10.0), Phylogenetic tree, Parsimony informative sites, Synonymous and non-synonymous mutationsAbstract
SARS-CoV-2 has very recently posed a potential threat to humanity due to its very rapid rate of mutations and repairing mechanism. The spread of this virus is considered to have occurred in Wuhan, China in December 2019. Characterized by high rates of transmission, the virus is constantly evolving towards attaining higher rates of stability and transmissibility through acquiring mutations in its genome. Therefore, this study aims to analyse the mutational profiles of SARS-CoV-2 isolates. Analysis of the mutational profiles in individual SARS-CoV-2 proteins will allow us to look into the rates of mutations associated with each protein. Frequently mutated residues have been identified in this research by aligning 688 SARS-CoV-2 nucleotide sequences, which were downloaded from NCBI (National Center For Biotechnology Information) repository. Further, mutational frequencies of these mutated residues have been studied, which is instrumental in identifying the proteins that are resistant to changes, as well as the ones that have a greater proclivity towards incorporating mutations.
HIGHLIGHTS
- The rates, frequencies and proclivities of mutation (w.r.t each protein) in SARS-CoV-2 were analyzed to facilitate vaccine development , supplemented with a phylogenetic tree (w.r.t spike protein)
- D614G mutation (frequency of more than 40 %) is crucial in imparting greater viral stability and transmissibility
- Both, at gene and protein level, number of mutations per 100 bases are high in Nucleocapsid , ORF3a and ORF8
- Envelope and Membrane proteins have housekeeping functions
GRAPHICAL ABSTRACT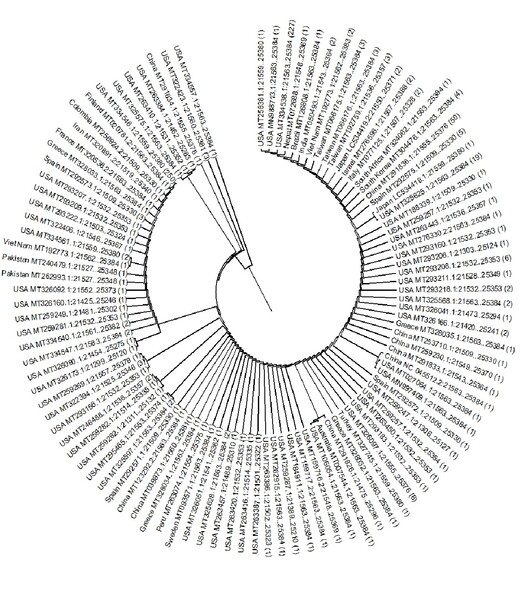
Downloads
References
S Alai, N Gujar, M Joshi, M Gautam and S Gairola. Pan-India novel coronavirus SARS-CoV-2 genomics and global diversity analysis in spike protein. Heliyon 2020; 7, e06564.
F Begum, D Mukherjee, D Thagriki, S Das, PP Tripathi, AK Banerjee and U Ray. Analyses of spike protein from first deposited sequences of SARS-CoV2 from West Bengal, India. F1000Research 2020; 9, 371.
JR Byrnes, XX Zhou, I Lui, SK Elledge, JE Glasgow, SA Lim, RP Loudermilk, CY Chiu, TT Wang, MR Wilson, KK Leung and JA Wells. Competitive SARS-CoV-2 serology reveals most antibodies targeting the spike receptor-binding domain compete for ACE2 binding. mSphere 2020; 5, e00802-20.
Y Chen, Q Liu and D Guo. Emerging coronaviruses: Genome structure, replication, and pathogenesis. J. Med. Virol. 2020; 92, 418-23.
AM Davidson, J Wysocki and D Batlle. Interaction of SARS-CoV-2 and other coronavirus with ACE (Angiotensin-Converting Enzyme)-2 as their main receptor: Therapeutic implications. Hypertension 2020; 76, 1339-49.
V Gupta, S Haider, M Verma, N Singhvi, K Ponnusamy, MZ Malik, H Verma, R Kumar, U Sood, P Hira, S Satija, Y Singh and R Lal. Comparative genomics and integrated network approach unveiled undirected phylogeny patterns, Co-mutational hot spots, functional cross talk, and regulatory interactions in SARS-CoV-2. mSystems 2021; 6, e00030-21.
SW Huang, SO Miller, Y Chia-Hung and W Sheng-Fan. Impact of genetic variability in ACE2 expression on the evolutionary dynamics of SARS-CoV-2 spike D614G mutation. Genes 2020; 12, 16.
A Hussain, A Hasan, MMN Babadaei, SH Bloukh, MEH Chowdhury, M Sharifi, S Haghighat and M Falahati. Targeting SARS-CoV2 Spike protein receptor binding domain by therapeutic antibodies. biomed pharmacother. Biomed. Pharmacother. 2020; 130, 110559.
MR Islam, MN Hoque, MS Rahman, ASMRU Alam, M Akther, JA Puspo, S Akter, M Sultana, KA Crandall and MA Hossain. Genome-wide analysis of SARS-CoV-2 virus strains circulating worldwide implicates heterogeneity. Sci. Rep. 2020; 10, 14004.
S Laha, J Chakraborty, S Das, SK Manna, S Biswas and R Chatterjee. Characterizations of SARS-CoV-2 mutational profile, spike protein stability and viral transmission. Infect. Genet. Evol. 2020; 85, 104445.
S Lee, L Mi-Kyeong, H Na, J Ahn, G Hong, Y Lee, J Park, Y Kim, K Yun-Tae, K Chang-Ki, L Hwan-Sub and L Kyoung-Ryul. Comparative analysis of mutational hotspots in the spike protein of SARS-CoV-2 isolates from different geographic origins. Gene Rep. 2021; 23, 101100.
AV Letarov, VV Babenko and EE Kulikov. Free SARS-CoV-2 spike protein S1 particles may play a role in the pathogenesis of Covid-19 infection. Biochemistry 2021; 86, 257-61.
I Mercurio, V Tragni, F Busto, AD Grassi and CL Pierri. Protein structure analysis of the interactions between SARS-CoV-2 spike protein and the human ACE2 receptor: From conformational changes to novel neutralizing antibodies. Cell. Mol. Life. Sci. 2021; 78, 1501-22.
A Shah, F Rashid, A Aziz, AU Jan and M Suleman. Genetic characterization of structural and open reading Fram-8 proteins of SARS-CoV-2 isolates from different countries. Gene Rep. 2020. 21: 100886.
Y Wan, J Shang, R Graham, RS Baric and F Li. Receptor recognition by the novel coronavirus from wuhan: An analysis based on decade-long structural studies of SARS coronavirus. J. Virol. 2020; 94, e0012720.
Y Weisblum, F Schmidt, F Zhang, J DaSilva, D Poston, JC Lorenzi, F Muecksch, M Rutkowska, H Hans-Heinrich, E Michailidis, C Gaebler, M Agudelo, A Cho, Z Wang, A Gazumyan, M Cipolla, L Luchsinger, CD Hillyer, M Caskey, DF Robbiani, ..., PD Bieniasz. Escape from neutralizing antibodies by SARS-CoV-2 spike protein variants. Elife 2020; 9, e61312.
L Zhang, CB Jackson, H Mou, A Ojha, H Peng, BD Quinlan, ES Rangarajan, A Pan, A Vanderheiden, MS Suthar, W Li, T Izard, C Rader, M Farzan and H Choe. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat. Comm. 2020; 11, 6013.
R Wang, J Chen, K Gao, Y Hozumi, C Yin and W Guo-Wei. Analysis of SARS-CoV-2 mutations in The United States suggest presence of four sub-strains and novel variants. Comm. Biol. 2021; 4, 228.
DC Groves, SL Rowland-Jones and A Angyal. The D614G mutations in the SARS-CoV-2 spike protein: Implications for viral infectivity, disease severity and vaccine design. Biochem. Biophys. Res. Comm. 2021; 538, 104-7.
AJ McAuley, MJ Kuiper, PA Durr, MP Bruce, J Barr, S Todd, GG Au, K Blasdell, M Tachedjian, S Lowther, GA Marsh, S Edwards, T Poole, R Layton, R Sarah-Jane, TW Drew, JD Druce, TRF Smith, KE Broderick and SS Vasan. Experimental and in silico evidence suggests vaccines are unlikely to be affected by D614G mutation in SARS-CoV-2 spike protein. npj Vaccines 2020; 5, 96.
TN Starr, AJ Greaney, A Addetia, WW Hannon, MC Choudhary, AS Dingens, JZ Li and JD Bloom. Prospective mapping of viral mutations that escape antibodies used to treat Covid-19. Science 2021; 371, 850-4.
S Jakhmola, O Indari, D Kashyap, N Varshney, A Das, E Manivannan and HC Jha. Mutational analysis of structural proteins of SARS-CoV-2. Heliyon 2021; 7, e06572.
D Wrapp, N Wang, KS Corbett, JA Goldsmith, H Ching-Lin, O Abiona, BS Graham and JS McLellan. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020; 367, 1260-63.
Y Cai, J Zhang, T Xiao, H Peng, SM Sterling, RMW Jr, S Rawson, S Rits-Volloch and B Chen. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020; 369, 1586-92.
S Kumar, G Stecher, M Li, C Knyaz and K Tamura. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018; 35, 1547-9
J Hadfield, C Megill, SM Bell, J Huddleston, B Potter, C Callender, P Sagulenko, T Bedford and RA Neher. Nextstrain: Real-time tracking of pathogen evolution. Bioinformatics 2018; 34, 4121-3.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.