DNA Methylation and Plants Response to Biotic and Abiotic Stress
DOI:
https://doi.org/10.48048/tis.2022.5102Keywords:
Genome, Methylation, Biotic stress, Abiotic stress, PlantAbstract
DNA methylation is a conserved epigenetic modification that regulates, stabilizes, and maintains genomic integrity. Loss of DNA methylation or aberrant patterns of DNA methylation causes abnormalities in the gene regulation of plants. DNA methylation in plants is regulated by the combined action of de novo methylation, maintenance of methylation, and demethylation. The enzymes that regulate DNA methylation in plants are different but have some homology to that of mammalian DNA methylation enzymes. DNA methylation helps to develop adaptation mechanisms towards various biotic and abiotic stresses. This paper provides a comprehensive review of the DNA methylation pathway and its role in biotic and abiotic stress tolerances in plants.
HIGHLIGHTS
- Plants responds to the changing climatic condition via epigenetic changes - changing the gene expression patterns, without changing the DNA sequences
- Abiotic and biotic stress leads to the differential expression of genes; furthermore, plants have memory in the form of epigenetic marks laid in the DNA sequences
- DNA methylation in plants is regulated by the combined action of de novo methylation, maintenance of methylation, and demethylation
GRAPHICAL ABSTRACT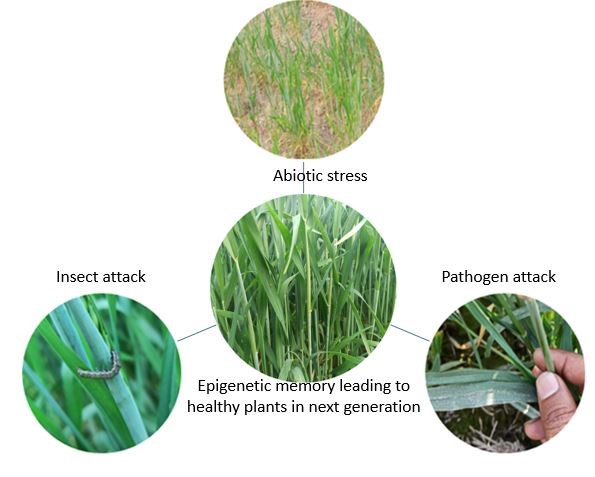
Downloads
References
DE Handy, R Castro and J Loscalzo. Epigenetic modifications: Basic mechanisms and role in cardiovascular disease. Circulation 2011; 123, 2145-56.
LD Moore, T Le and G Fan. DNA methylation and its basic function. Neuropsychopharmacology 2013; 38, 23-38.
C Zhou, C Wang, H Liu, Q Zhou, Q Liu, Y Guo, T Peng, J Song, J Zhang, L Chen, Y Zhao, Z Zeng and DX Zhou. Identification and analysis of adenine N 6-methylation sites in the rice genome. Nat. Plants 2018; 4, 554-63.
H Yang, F Chang, C You, J Cui, G Zhu, L Wang and H Ma. Whole-genome DNA methylation patterns and complex associations with gene structure and expression during flower development in Arabidopsis. Plant J. 2015; 38, 268-81.
E Zubko, M Gentry, A Kunova and P meyer. De novo DNA methylation activity of Methyltransferase 1 (MET1) partially restores body methylation in Arabidopsis thaliana. Plant J. 2012; 71, 1029-37.
W Aufsatz, MF Mette, JVD Winden, AJM Matzke and M Matzke. RNA-directed DNA methylation in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 2002; 99, 16499-506.
H Zhang, Z Lang and JK Zhu. Dynamics and function of DNA methylation in plants. Nat. Rev. Mol. Cell Biol. 2018; 19, 489-506.
J Gallego-Bartolomé, W Liu, PH Kuo, S Feng, B Ghoshal, J Gardiner, J Miao-ChiZhao, SY Park, J Chory and SE Jacobsen. Co-targeting RNA Polymerases IV and V promotes efficient De Novo DNA methylation in Arabidopsis. Cell 2019; 176, 1068-1082.e19.
SP Wongpalee, S Liu, J Gallego-Bartolomé, A Leitner, R Aebersold, W Liu, L Yen, MA Nohales, PH Kuo, AA Vashisht, JA Wohlschlegel, S Feng, SA Kay, ZH Zhou and SE Jacobsen. CryoEM structures of Arabidopsis DDR complexes involved in RNA-directed DNA methylation. Nat. Commun. 2019; 10, 3916.
S Zhong, Y Xu, C Yu, X Zhang, L Li, H, Ge and B Zheng. Anaphase-promoting complex/cyclosome regulates RdDM activity by degrading DMS3 in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 2019; 116, 3899-908.
A Zemach, MY Kim, PH Hsieh, D Coleman-Derr, L Eshed-Williams, K Thao, SL Harmer and D Zilberman. The arabidopsis nucleosome remodeler DDM1 allows DNA methyltransferases to access H1-containing heterochromatin. Cell 2013; 153, 193-205.
Y Fu, A Kawabe, M Etcheverry, T Ito, A Toyoda, A Fujiyama, V Colot, Y Tarutani and T Kakutani. Mobilization of a plant transposon by expression of the transposon-encoded anti-silencing factor. European Mol. Biol. Organ. J. 2013; 32, 2407-17.
XJ He, T Chen and JK Zhu. Regulation and function of DNA methylation in plants and animals. Cell Res. 2011; 21, 442-65.
J Gallego-Bartolomé. DNA methylation in plants: Mechanisms and tools for targeted manipulation. New Phytol. 2020; 227, 38-44.
J Du, X Zhong, YV Bernatavichute, H Stroud, S Feng, E Caro, Ajay A Vashisht, J Terragni, H Gyeong Chin, A Tu, J Hetzel, JA Wohlschlegel, S Pradhan, DJ Patel and SE Jacobsen. Dual binding of chromomethylase domains to H3K9me2-containing nucleosomes directs DNA methylation in plants. Cell 2012; 151, 167-80.
R Liu and Z Lang. The mechanism and function of active DNA demethylation in plants. J. Integr. Plant Biol. 2020; 62, 148-59.
J FitzGerald, M Luo, A Chaudhury and F Berger. DNA methylation causes predominant maternal controls of plant embryo growth. PLoS ONE. 2008; 3, e2298.
Z Gong, T Morales-Ruiz, RR Ariza, T Roldán-Arjona, L David and JK Zhu. ROS1, a repressor of transcriptional gene silencing in Arabidopsis, encodes a DNA glycosylase/lyase. Cell 2002; 111, 803-14.
AP Ortega-Galisteo, T Morales-Ruiz, RR Ariza, T Roldán-Arjona. Arabidopsis DEMETER-LIKE proteins DML2 and DML3 are required for appropriate distribution of DNA methylation marks. Plant Mol. Biol. 2008; 67, 671-81.
J Lee, H Jang, H Shin, WL Choi, YG Mok and JH Huh. AP endonucleases process 5-methylcytosine excision intermediates during active DNA demethylation in Arabidopsis. Nucleic Acids Res. 2014; 42, 11612-21.
Y Li, D Córdoba-Cañero, W Qian, X Zhu, K Tang, H Zhang, RR Ariza, T Roldán-Arjona and JK Zhu. An AP endonuclease functions in Active DNA demethylation and gene imprinting in Arabidopsis. PLoS Genet. 2015; 11, e1005198.
MI Martínez-Macías, W Qian, D Miki, O Pontes, Y Liu, K Tang, R Liu, T Morales-Ruiz, RR Ariza, T Roldán-Arjona and JK Zhu. A DNA 3’ phosphatase functions in active DNA demethylation in Arabidopsis. Mol. Cell 2012; 45, 357-70.
JT Parrilla-Doblas, T Roldán-Arjona, RR Ariza and D Córdoba-Cañero. Active DNA demethylation in plants. Int. J. Mol. Sci. 2019; 20, 4683.
C Satgé, S Moreau, E Sallet, G Lefort, MC Auriac, C Remblière, L Cottret, K Gallardo, C Noirot, MF Jardinaud and P Gamas. Reprogramming of DNA methylation is critical for nodule development in Medicago truncatula. Nat. Plants 2016; 2, 16166.
M Nagymihály, A Veluchamy, Z Györgypál, F Ariel, T Jégu, M Benhamed, A Szűcs, A Kereszt, P Mergaert, E Kondorosi. Ploidy-dependent changes in the epigenome of symbiotic cells correlate with specific patterns of gene expression. Proc. Natl. Acad. Sci. U.S.A. 2017; 114, 4543-8.
RH Dowen, M Pelizzola, RJ Schmitz, R Lister, JM Dowen, JR Nery, JE Dixon and JR Ecker. Widespread dynamic DNA methylation in response to biotic stress. Proc. Natl. Acad. Sci. U.S.A. 2012; 109, E2183-E2191.
S Lee, F Fu, S Xu, SY Lee, DJ Yun and T Mengiste. Global regulation of plant immunity by histone lysine methyl transferases. Plant. Cell 2016; 28, 1640-61.
MYK Barozai and AN Aziz, Recent plant growth and stress management related significant advancements in epigenetics. Ann. Agrar. Sci. 2018; 16, 416-21.
M Castellano, G Martinez, MC Marques, J Moreno-Romero, C Köhler, V Pallas and G Gomez, Changes in the DNA methylation pattern of the host male gametophyte of viroid-infected cucumber plants. J. Exp. Bot. 2016; 67, 5857-68.
M Castellano, G Martinez, V Pallás and G Gómez. Alterations in host DNA methylation in response to constitutive expression of Hop stunt viroid RNA in Nicotiana benthamiana plants. Plant Pathol. 2015; 64, 1247-57.
A López, V Ramírez, J García-Andrade, V Flors and P Vera. The RNA silencing enzyme RNA polymerase V is required for plant immunity. PLoS Genet. 2011; 7, e1002434.
D Secco, C Wang, H Shou, MD Schultz, S Chiarenza, L Nussaume, Joseph R EckerJames WhelanR Lister. Stress induced gene expression drives transient DNA methylation changes at adjacent repetitive elements. ELife 2015; 4, e09343.
B Zhang, DM Tieman, C Jiao, Y Xu, K Chen, Z Fei, JJ Giovannoni and HJ Klee. Chilling-induced tomato flavor loss is associated with altered volatile synthesis and transient changes in DNA methylation. Proc. Natl. Acad. Sci. U.S.A. 2016; 113, 12580-5.
J Lämke, K Brzezinka, S Altmann and I Bäurle. A hit‐and‐run heat shock factor governs sustained histone methylation and transcriptional stress memory. European Mol. Biol. Organ. J. 2016; 35, 162-75.
L Kong, Y Liu, X Wang and C Chang, Insight into the role of epigenetic processes in abiotic and biotic stress response in wheat and barley. Int. J. Mol. Sci. 2020; 21, 1480.
D Secco, J Whelan, H Rouached and R Lister. Nutrient stress-induced chromatin changes in plants. Curr. Opin. Plant Biol. 2017; 37, 1-7.
SR Eichten and NM Springer, Minimal evidence for consistent changes in maize DNA methylation patterns following environmental stress. Front. Plant Sci. 2015; 6, 308.
DH Sanchez and J Paszkowski. Heat-induced release of epigenetic silencing reveals the concealed role of an imprinted plant gene. PLoS Genet. 2014; 10, e1004806.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.