A Computational Alanine Scanning of Heptapeptide RRRSAGM Targeting Dengue NS2B/NS3 Protease
DOI:
https://doi.org/10.48048/tis.2022.4621Keywords:
Computational alanine scanning, Molecular docking, Dengue virus, NS2B/NS3 protease, Peptide-based inhibitorAbstract
The pathogenic dengue virus (DV) transmitted mainly by the Aedes aegypti mosquito has been the most rapidly spreading mosquito-borne viral disease and became a serious threat to global health issue in tropical and sub-tropical countries. The clinical symptoms ranging from mild to a fatal dengue hemorrhagic fever and dengue shock syndrome, characterized by failure of the circulation system that may lead to death. There are still no drugs or vaccines available in the market to prevent or treat dengue infections. Hence, the development of successful drugs that are safe and offer a long-lasting protection against dengue viruses is needed. Recently, the discovery of peptides with high specificity, selectivity, and efficacy; and relatively safer than small-molecule drugs have become a promising agent for a new drug. Previously, a molecular dynamic simulation reported that peptide NRRRRSAGMI from the capsid’s cleavage region had the most hydrogen bonds with NS2B/NS3 protease, thus it leads into a starting linear heptapeptide R(P3)-R(P2)-R(P1)-S(P1’)-A(P2’)-G(P3’)-M(P4’). This study aimed to investigate the importance and specificity of individual amino acid residue of the heptapeptide substrate on the binding affinity and interaction to NS2B/NS3 dengue protease through computational alanine scanning. It showed that the changing of P1 and P2 arginine to alanine resulted to higher values of binding affinity. Thus, the arginine is an important residue at the P1 and P2 positions for the binding of the substrate peptide to DENV NS2B/NS3 protease, hence the amino acid residues which are essential for the enzyme-substrate interactions can be predicted through computational alanine scanning.
HIGHLIGHTS
- Dengue is a mosquito-transmitted virus cause of arthropod-borne viral disease
- The discovery of peptides became a potential agent for dengue virus and previously peptide NRRRRSAGMI has been reported as starting linear heptapeptide RRRSAGM
- A computational alanine scanning based on the peptide RRRSAGM has been performed
- S1 and S2 pockets show strong preferences for basic amino acids, whereas the S3 pocket is less selective for basic residue
- The arginine at P1 and P2 positions of the heptapeptide substrate contribute significantly to the substrate binding affinity
GRAPHICAL ABSTRACT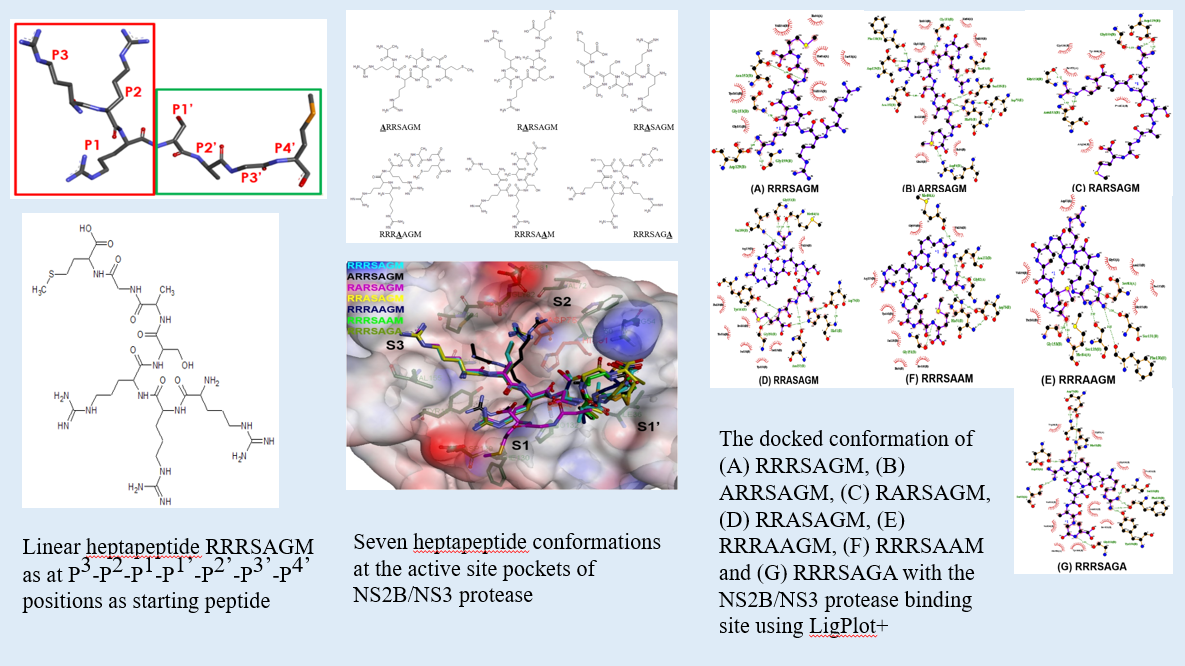
Downloads
References
World Health Organization. Update on the dengue situation in the western pacific region. World Health Organization, 2020.
BL Ligon. Dengue fever and dengue hemorrhagic fever: a review of the history, transmission, treatment, and prevention. Semin Pediatr Infect Dis. 2005; 16, 60-5.
MF Chew, KS Poh and CL Poh. Peptides as therapeutic agents for dengue virus. Int. J. Med. Sci. 2017; 14, 1342.
K Morita, M Tanaka and A Igarashi. Rapid identification of dengue virus serotypes by using polymerase chain reaction. J. Clin. Microbiol. 1991; 29, 2107-10.
AK Ageep, AA Malik and MS Elkarsani. Clinical presentations and laboratory findings in suspected cases of dengue virus. Saudi Med. J. 2006; 27, 1711-3.
RJ Kuhn, W Zhang, MG Rossmann, SV Pletnev, J Corver, E Lenches, CT Jones, S Mukhopadhyay, PR Chipman, EG Strauss, TS Baker and JH Strauss. Structure of dengue virus: implications for flavivirus organization, maturation, and fusion. Cell 2002; 108, 717-25.
BD Lindenbach, HJ Thiel and CM Rice. Flaviviridae: The viruses and their replication. In: DM Knipe and OM Howley (Eds.). Fields virology, 5th Eds. Lippincot William & Wilkins, Philadelphia, 2011, p. 1101-51.
B Falgout, R Miller and C Lai. Deletion analysis of dengue virus type 4 nonstructural protein NS2B: identification of a domain required for NS2B-NS3 protease activity. J. Virol. 1993; 67, 2034-42.
IA Rodenhuis-Zybert, J Wilschut and JM Smit. Dengue virus life cycle: viral and host factors modulating infectivity. Cell. Mol. Life Sci. 2010; 67, 2773-86.
R Assenberg, E Mastrangelo, TS Walter, A Verma, M Milani, RJ Owens, DI Stuart, JM Grimes and EJ Mancini. Crystal structure of a novel conformational state of the flavivirus NS3 protein: Implications for processing and viral replication. J. Virol. 2009; 83, 12895-906.
J Chen, H Jiang, F Li, B Hu, Y Wang, MX Wang, J Wang and M Cheng. Computational insight into dengue virus NS2B-NS3 protease inhibition: A combined ligand-and structure-based approach. Comput. Biol. Chem. 2018; 77, 261-71.
S Xu, H Li, X Shao, C Fan, B Ericksen, J Liu, C Chi and C Wang. Critical effect of peptide cyclization on the potency of peptide inhibitors against Dengue virus NS2B-NS3 protease. J. Med. Chem. 2012; 55, 6881-7.
C Mazeaud, W Freppel and L Chatel-Chaix. The multiples fates of the flavivirus RNA genome during pathogenesis. Front. Genet. 2018; 9, 595.
A Huther and U Dietrich. The emergence of peptides as therapeutic drugs for the inhibition of HIV-1. AIDS Rev. 2007; 9, 208-17.
USF Tambunan and S Alamudi. Designing cyclic peptide inhibitor of dengue virus NS3-NS2B protease by using molecular docking approach. Bioinformation 2010; 5, 250.
J Brunetti, C Falciani, G Roscia, S Pollini, S Bindi, S Scali, UC Arrieta, V Gómez-Vallejo, L Quercini, E Ibba, M Prato, GM Rossolini, J Llop, L Bracci and A Pini. In vitro and in vivo efficacy, toxicity, bio-distribution and resistance selection of a novel antibacterial drug candidate. Sci. Rep. 2016; 6, 26077.
M Wang, KP Rakesh, J Leng, WY Fang, L Ravindar, DC Gowda and HL Qin. Amino acids/peptides conjugated heterocycles: A tool for the recent development of novel therapeutic agents. Bioorg. Med. Chem. 2018; 76, 113-29.
HA Rothan, AY Abdulrahman, PG Sasikumer, S Othman, NA Rahman and R Yusof. Protegrin-1 inhibits dengue NS2B-NS3 serine protease and viral replication in MK2 cells. J. Biomed. Biotechnol. 2012; 2012, 251482.
P Yotmanee, T Rungrotmongkol, K Wichapong, SB Choi, HA Wahab, N Kungwan and S Hannongbua. Binding specificity of polypeptide substrates in NS2B/NS3pro serine protease of dengue virus type 2: A molecular dynamics study. J. Mol. Graph. Model. 2015; 60, 24-33.
J Li, SP Lim, D Beer, V Patel, D Wen, C Tumanut, DC Tully, JA Williams, J Jiricek, JP Priestle, JL Harris and SG Vasudevan. Functional profiling of recombinant NS3 proteases from all four serotypes of dengue virus using tetrapeptide and octapeptide substrate libraries. J. Biol. Chem. 2005; 280, 28766-74.
K Wichapong, S Pianwanit, W Sippl and S Kokpol. Homology modeling and molecular dynamics simulations of Dengue virus NS2B/NS3 protease: insight into molecular interaction. J. Mol. Recognit. 2010; 23, 283-300.
MF Sanner. A component-based software environment for visualizing large macromolecular assemblies. Structure 2005; 13, 447-62.
R Huey and GM Morris. Using AutoDock 4 with AutoDocktools: A tutorial. The Scripps Research Institute, California, 2008, p. 54-6.
Z Yin, SJ Patel, WL Wang, WL Chan, KRR Rao, G Wang, X Ngew, V Patel, D Beer, J E Knox, NL Ma, C Ehrhardt, SP Lim, SG Vasudevan and TH Keller. Peptide inhibitors of dengue virus NS3 protease. Part 2: SAR study of tetrapeptide aldehyde inhibitors. Bioorg. Med. Chem. Lett. 2006; 16, 40-3.
J Gasteiger and M Marsili. Iterative partial equalization of orbital electronegativity: A rapid access to atomic charges. Tetrahedron 1980; 36, 3219-28.
NA Abdalrazaq and EE Kamarulzaman. Rational design of cyclic tetra and pentapeptides as therapeutic agents for dengue NS2B/NS3 protease using structure-based molecular docking (MOE and AutoDock 4.2). Int. J. Pharm. Sci. Res. 2020; 11, 5501-10.
AC Wallace, RA Laskowski and JM Thornton. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. Des. Sel. 1995; 8, 127-34.
DS Biovia. BIOVIA workbook, release 2017; BIOVIA pipeline pilot, release 2017. San Diego: Dassault Systèmes, 2020.
B Podlogar, I Muegge and L Brice. Computational methods to estimate drug development parameters. Curr. Opin. Drug Discov. 2001; 4, 102-9.
S Cosconati, S Forli, AL Perryman, R Harris, DS Goodsell and AJ Olson. Virtual screening with AutoDock: theory and practice. Expert Opin. Drug Discov. 2010; 5, 597-607.
MAF Nasution, RN Aini and USF Tambunan. Virtual screening of commercial cyclic peptides as NS2B-NS3 protease inhibitor of dengue virus serotype 2 through molecular docking simulation. IOP Conf. Series: Mat. Sci. Eng. 2017; 188, 012017.
I Massova and PA Kollman. Computational alanine scanning to probe protein-protein interactions: A novel approach to evaluate binding free energies. J. Am. Chem. Soc. 1999; 121, 8133-43.
GM Morris, DS Goodsell, RS Halliday, R Huey,WE Hart, RK Belew and AJ Olson. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 1998; 19, 1639-62.
EE Kamarulzaman, R Vanderesse, AM Gazzali, M Barberi-Heyob, C Boura, C Frochot, O Shawkataly, A Aubry and HA Wahab. Molecular modelling, synthesis and biological evaluation of peptide inhibitors as anti-angiogenic agent targeting neuropilin-1 for anticancer application. J. Biomol. Struct. Dyn. 2017; 35, 26-45.
R Yusof, S Clum, M Wetzel, HM Murthy and R Padmanabhan. Purified NS2B/NS3 serine protease of dengue virus type 2 exhibits cofactor NS2B dependence for cleavage of substrates with dibasic amino acids in vitro. J. Biol. Chem. 2000; 275, 9963-9.
E Arunan, GR Desiraju, RA Klein, J Sadlej, S Scheiner, I Alkorta, DC Clary, RH Crabtree, JJ Dannenberg, P Hobza, HG Kjaergaard, AC Legon, B Mennucci and DJ Nesbitt. Definition of the hydrogen bond (IUPAC Recommendations 2011). Pure Appl. Chem. 2011; 83, 1637-41.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.







