Assessment of Bacterial Community Profile in the Rearing Pond Environment and the Intestinal Tract of Pacific White Shrimp Litopenaeus vannamei in Lampung Province, Indonesia using 16S rRNA Gene Amplicon Sequencing: A Short Research Investigation
DOI:
https://doi.org/10.48048/tis.2023.3418Keywords:
Aquaculture environment, Bacterial diversity, Intestinal tract, Metagenomic, Whiteleg shrimpAbstract
Along with the high demand for shrimp, the production challenges faced by practitioners are increasing in Indonesia, one of which is in Lampung region. Various basic monitoring techniques regarding the environment and farmed shrimp are needed to control production sustainability. This present study aimed to identify the bacterial community profile in the rearing pond water and intestinal tract of Pacific white shrimp Litopenaeus vannamei. Illumina-based sequencing was chosen to determine the bacterial community using the V3-V4 region of the bacterial 16S rRNA gene. Sequence data revealed the differences in bacterial community structure between the rearing water and shrimp intestines. Proteobacteria was the most prevalent phylum in the rearing water (W.B), accounting for 45.29 %, followed by Cyanobacteria, Firmicutes, Bacteroidetes, Actinobacteria, and Fusobacteria. In the shrimp intestinal tract (S.B), Cyanobacteria (35.15 %) dominated the microbiota, followed by Proteobacteria, Saccharibacteria, Actinobacteria, Bacteroidetes, Verrucomicrobia, TM6 (Dependentiae), and Firmicutes. Cyanobacteria were higher in the shrimp intestines (35.15 %) than in the rearing water (26.63 %). In addition, Escherichia-Shigella was the most common genera in the rearing water and shrimp intestines with different relative abundance. Cyanobacteria and Escherichia-Shigella highly detected in the rearing water and shrimp’s intestines might indicate that pond water had been polluted. Further investigation is necessary on the correlation of Cyanobacteria in the shrimp intestines with water pollution, proven by the dominance of Cyanobacteria and Escherichia-Shigella. These findings provide basic information to enhance our understanding of the microbial community and their roles in the shrimp culture environment to improve the quantity and quality of the yield and support its sustainability.
HIGHLIGHTS
- Bacterial community profile in the rearing water and the intestinal tract of Pacific whiteleg shrimp play important roles in shrimp production
- Bacterial community structure differed between the rearing water and shrimp intestines
- Proteobacteria was the most abundant phylum in the rearing water (W.B), while Cyanobacteria dominated the microbiota in the intestinal tract of Pacific whiteleg shrimp
- Cyanobacteria and Escherichia-Shigella highly detected in the rearing water and shrimp’s intestines might indicate that pond water had been polluted
GRAPHICAL ABSTRACT 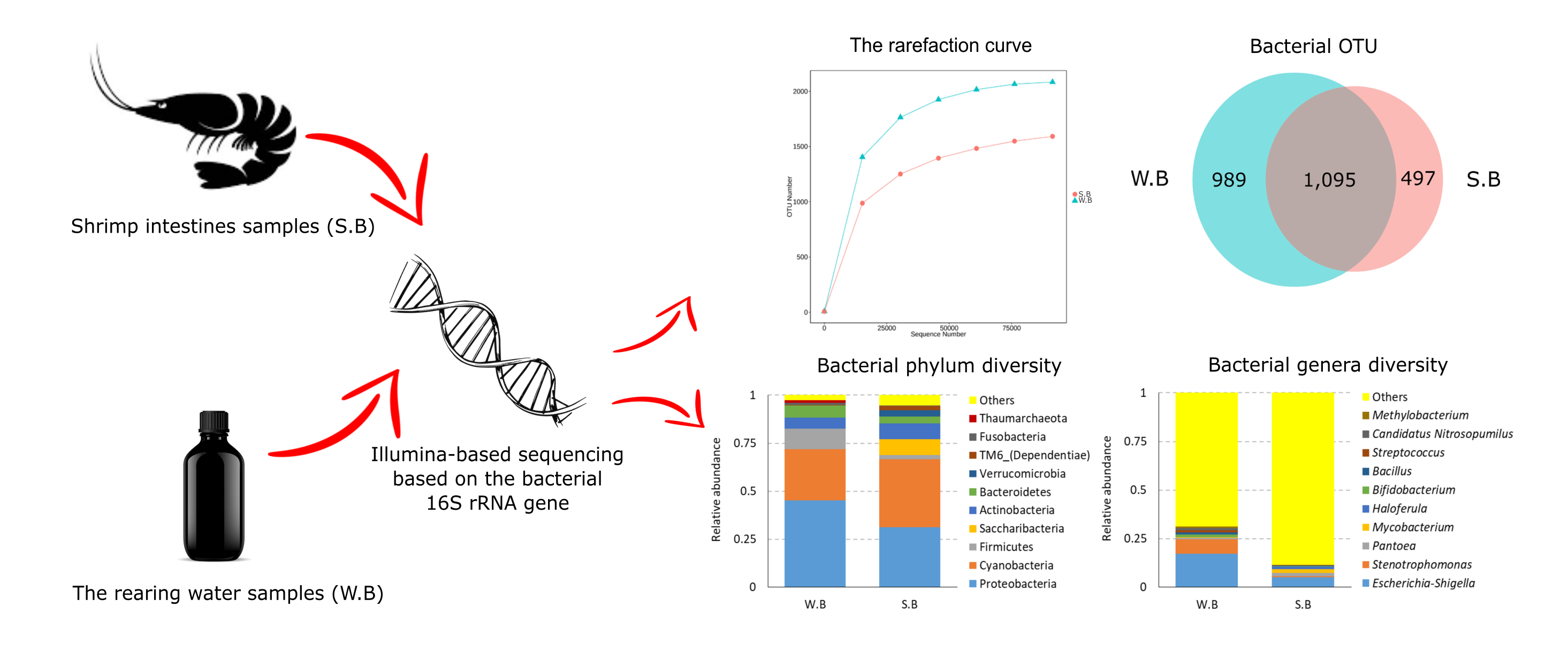
Downloads
References
FAO. The state of world fisheries and aquaculture: Meeting the sustainable development goals. Food and Agriculture Organization of the United Nations, Rome, Italy, 2018.
KKP. Peta lalu lints benuh ikan dan benur udang nasional 2018 (in Indonesia). Available at: https://kkp.go.id/bkipm/artikel/5880-peta-lalulintas-benih-ikan-dan-benur-udang-nasional-2018, accessed April 2020.
YP Hastuti. Nitrification and denitrification in pond. Jurnal Akuakultur Indonesia 2011; 10, 89-98.
SF Snieszko. The effects of environmental stress on outbreaks of infectious diseases of fishes. J. Fish. Biol. 1974; 6, 197-208.
MWI Schmidt, MS Torn, S Abiven, T Dittmar, G Guggenberger, IA Janssens, M Kleber, I Kögel-Knabner, J Lehmann, DAC Manning, P Nannipieri, DP Rasse, S Weiner and SE Trumbore. Persistence of soil organic matter as an ecosystem property. Nature 2011; 478, 49-56.
D Griggs, M Stafford-Smith, O Gaffney, J Rockstrom, MC Ohman, P Shyamsundar, W Steffen, G Glaser, N Kanie and I Noble. Sustainable development goals for people and planet. Nature 2013; 495, 305-7.
PA Venail and JV Martha. Positive effects of bacterial diversity on ecosystem functioning driven by complementarity effects in a bioremediation context. PLoS One 2013; 8, e72561.
YP Hastuti, I Rusmana, K Nirmala, R Affandi and S Tridesianti. Identification and characterization of nitrifying bacteria in mud crab (Scylla serrata) recirculation aquaculture system by 16S rRNA sequencing. Biodiversitas 2019; 20, 1339-43.
YP Hastuti, YS Fatma, H Pitoyo, Y Wardiatno and S Tridesianti. Bacterial and plankton communities in mariculture water source: A case study in Nampu and Sembukan seawaters, Wonogiri, Indonesia. Nat. Croat. 2021; 30, 341-56.
YP Hastuti, YS Fatma, H Pitoyo, W Nurussalam and J Ruhyana. Assesment of total bacterial diversity in whiteleg shrimp and its aquaculture environment in Pangkajene and Banyuwangi, Indonesia. Asia Pac. J. Mol. Biol. Biotechnol. 2021; 29, 26-37.
F Huang, L Pan, M Song and CS Tian. Microbiota assemblages of water, sediment, and intestine and their associations with environmental factors and shrimp physiological health. Appl. Microbiol. Biotechnol. 2018; 102, 8585-98.
JF Rawls, NS Samuel and JI Gordon. Gnotobiotic zebrafish reveal evolutionaily conserved response to the gut microbiota. Proc. Natl. Acad. Sci. U.S.A. 2004; 101, 4596-601.
W Rungrassamee, A Klanchui, S Maibunkaew, S Chaiyapechara, P Jiravanichpaisal and N Karoonuthaisiri. Characterization of intestinal bacteria in wild and domesticated adult black tiger shrimp (Penaeus monodon). PLoS One 2014; 9, e91853.
J Xiong, J Zhu, W Dai, C Dong, Q Qiu and C Li. Integrating gut microbiota immaturity and disease-discriminatory taxa to diagnose the initiation and severity of shrimp disease. Environ. Microbiol. 2017; 19, 1490-501.
Y Yu, C Lee, J Kim and S Hwang. Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol. Bioeng. 2005; 89, 670-9.
T Magoc and SL Salzberg. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011; 27, 2957-63.
NA Bokulich, S Subramanian, JJ Faith, D Gevers, JI Gordon, R Knight, DA Mills and JG Caporaso. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Meth. 2013; 10, 57-9.
JG Caporaso, J Kuczynski, J Stombaugh, K Bittinger, FD Bushman, EK Costello, N Fierer, AG Peña, JK Goodrich, JI Gordon, GA Huttley, ST Kelley, D Knights, JE Koenig, RE Ley, CA Lozupone, D McDonald, BD Muegge, M Pirrung, J Reeder, JR Sevinsky, PJ Turnbaugh, WA Walters, J Widmann, T Yatsunenko, J Zaneveld and R Knight. QIIME allows analysis of high-throughput community sequencing data. Nat. Meth. 2010; 7, 335-6.
RC Edgar, BJ Haas, JC Clemente, C Quince and R Knight. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011; 27, 2194-200.
BJ Haas, D Gevers, AM Earl, M Feldgarden, DV Ward, G Giannoukos, D Ciulla, D Tabbaa, SK Highlander, E Sodergren, B Methé, TZ DeSantis, Human Microbiome Consortium, JF Petrosino, R Knight and BW Birren. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res. 2011; 21, 494-504.
RC Edgar. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Meth. 2013; 10, 996-8.
CZ Wang, GR Lin, T Yan, ZP Zheng, B Chen and FL Sun. The cellular community in the intestine of the shrimp Penaeus penicillatus and its culture environments. Fish. Sci. 2014; 80, 1001-7.
C Quast, E Pruesse, P Yilmaz, J Gerken, T Schweer, P Yarza, J Peplies and FO Glöckner. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013; 41, D590-D596.
E Cardona, Y Gueguen, K Magré, B Lorgeoux, D Piquemal, F Pierrat, F Noguier and D Saulnier. Bacterial community characterization of water and intestine of the shrimp Litopenaeus stylirostris in a biofloc system. BMC Microbiol. 2016; 16, 157.
A Landsman, BS Pierre, MR Leija, M Brown and W Gibbons. Impact of aquaculture practices on intestinal bacterial profiles of pacific whiteleg shrimp Litopenaeus vannamei. Microorganisms 2019; 7, 93.
S Zeng, Z Huang, D Hou, J Liu, S Weng and J He. Composition, diversity and function of intestinal microbiota in pacific white shrimp (Litopenaeus vannamei) at different culture stages. PeerJ 2017; 5, e3986.
S Abubucker, N Segata, J Goll, AM Schubert, J Izard, BL Cantarel, B Rodriguez-Mueller, J Zucker, M Thiagarajan, B Henrissat, O White, ST Kelley, B Methé, PD Schloss, D Gevers, M Mitreva and C Huttenhower. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 2012; 8, e1002358.
O Gainza, C Ramírez, AS Ramos and J Romero. Intestinal microbiota of white shrimp Penaeus vannamei under intensive cultivation conditions in Ecuador. Microb. Ecol. 2018; 75, 562-68.
ZB Huang, XY Li, LP Wang and ZZ Shao. Changes in the intestinal bacterial community during the growth of white shrimp, Litopenaeus vannamei. Aquac. Res. 2016; 47, 1737-46.
W Rungrassamee, A Klanchui, S Chaiyapechara, S Maibunkaew, S Tangphatsornruang, P Jiravanichpaisal and N Karoonuthaisiri. Bacterial population in intestines of the black tiger shrimp (Penaeus monodon) under different growth stages. PLoS One 2013; 8, e60802.
H Bothe, O Schmitz, MG Yates and WE Newton. Nitrogen fixation and hydrogen metabolism in Cyanobacteria. Microbiol. Mol. Biol. Rev. 2010; 74, 529-51.
R Alonso-Rodríguez and F Paez-Osuna. Nutrients, phytoplankton and harmful algal blooms in shrimp ponds: A review with special reference to the situation in the Gulf of California. Aquaculture 2003; 219, 317-36.
L Massaut and J Ortiz. Cyanobacteria toxicity tested in shrimp larvae - Oscillatoria brevis reduces P. vannamei survival. Available at: https://www.globalseafood.org/advocate/cyanobacteria-toxicity-tested-in-shrimp-larvae/, accessed August 2020.
Y Sha, M Liu, B Wang, K Jiang, C Qi and L Wang. Bacterial population in intestines of Litopenaeus vannamei fed different probiotics or probiotic supernatant. J. Microbiol. Biotechnol. 2016; 26, 1736-45.
F Cornejo-Granados, L Gallardo-Becerra, M Leonardo-Reza, JP Ochoa-Romo and A Ochoa-Leyva. A meta-analysis reveals the environmental and host factors shaping the structure and function of the shrimp microbiota. PeerJ 2018; 6, e5382
JD Dubreuil, RE Isaacson and DM Schifferli. Animal enterotoxigenic Escherichia coli. EcoSal Plus 2016; 7, 1-80.
SCRD Rienzi, I Sharon, KC Wrighton, O Koren, LA Hug, BC Thomas, JK Goodrich, JT Bell, TD Spector, JF Banfield and RE Ley. The human gut and groundwater harbor non photosynthetic bacteria belonging to a new candidate phylum sibling to Cyanobacteria. eLife 2013; 2, e01102.
RM Soo, BJ Woodcroft, DH Parks, GW Tyson and P Hugenholtz. Back from the dead; the curious tale of the predatory cyanobacterium Vampirovibrio chlorellavorus. PeerJ 2015; 3, e968.
J Gao, H Zuo, L Yang, JH He, S Niu, S Weng, J He and X Xu. Long-term influences of Cyanobacterial bloom on the immune system of Litopenaeus vannamei. Fish Shellfish Immunol. 2017; 61, 79-85.
Q Wang, GM Garrity, JM Tiedje and JR Cole. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007; 73, 5261-7.
A Pangastuti, A Suwanto, Y Lestari and MT Suhartono. Bacterial communities associated with white shrimp (Litopenaeus vannamei) larvae at early developmental stages. Biodiversitas 2010; 11, 65-8.
W Soonthornchai, W Rungrassamee, N Karoonuthaisiri, P Jarayabhand, S Klinbunga, K Soderhall, et al. Expression of immune-related genes in the digestive organ of shrimp, Penaeus monodon, after an oral infection by Vibrio harveyi. Dev. Com. Immunol. 2010; 34, 19-28.
J Fan, L Chen, G Mai, H Zhang, J Yang, D Deng and Y MA. Dynamics of the gut microbiota in developmental stages of Litopenaeus vannamei reveal its association with body weight. Nature 2019; 9, 734.

Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2022 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






