Molecular Characterization of Antibiotic Resistant Genes among Gram Negative Clinical Isolates
DOI:
https://doi.org/10.48048/tis.2022.2687Keywords:
Antibiotic resistance, ESBL, MBL, AmpC, IntegronsAbstract
Increasing antibiotic resistance is a threat worldwide. Deep studies have to be done for correct and appropriate usage of antibiotics for treating infections efficiently and to curtail the spread of resistance. Hence, this study is aimed at understanding the prevalence of antibiotic resistance genes among 35 gram-negative bacterial isolates. By performing biochemical tests, the isolates were identified to be Escherichia coli (n = 15), Klebsiella oxytoca (n = 3), Klebsiella pneumoniae (n = 5), Proteus mirabilis (n = 4), Proteus vulgaris (n = 1) and Pseudomonas aeruginosa (n = 7). Antibiotic susceptibility tests showed that most E. coli isolates were resistant to ampicillin, levofloxacin and cefazolin. Ampicillin resistance was found in all of Klebsiella oxytoca, Klebsiella pneumoniae and Proteus mirabilis isolates. All Proteus mirabilis isolates were resistant to ciprofloxacin and co-trimoxazole. Pseudomonas aeruginosa showed maximum resistance to gentamycin and tobramycin. All 35 isolates were sensitive to amikacin, imipenem and meropenem. Polymerase Chain Reaction showed that AmpC and SHV genes were predominant in E. coli and Klebsiella species respectively. Among Proteus mirabilis and Pseudomonas aeruginosa, IntI gene was more prevailing. NDM1 gene was not detected in any of the isolates. This study also showed the prevalence of isolates resistant to multiple drugs due to the co-existence of resistant genes. Therefore, control of spread of infections, routine study in this field and proper guidelines for prescribing antibiotics is very necessary.
HIGHLIGHTS
- This study is aimed at understanding the prevalence of antibiotic resistance genes among 35 gram-negative bacterial isolates
- A considerable number of isolates had antibiotic resistance genes and some isolates showed resistance to multiple drugs due to the co-existence of resistant genes
- Infections caused by bacteria that are resistant to antibiotics are difficult to treat and may sometimes lead to death
- Regular study and proper use of antibiotics is essential to curtail the spread of antibiotic resistant bacteria. Regular
GRAPHICAL ABSTRACT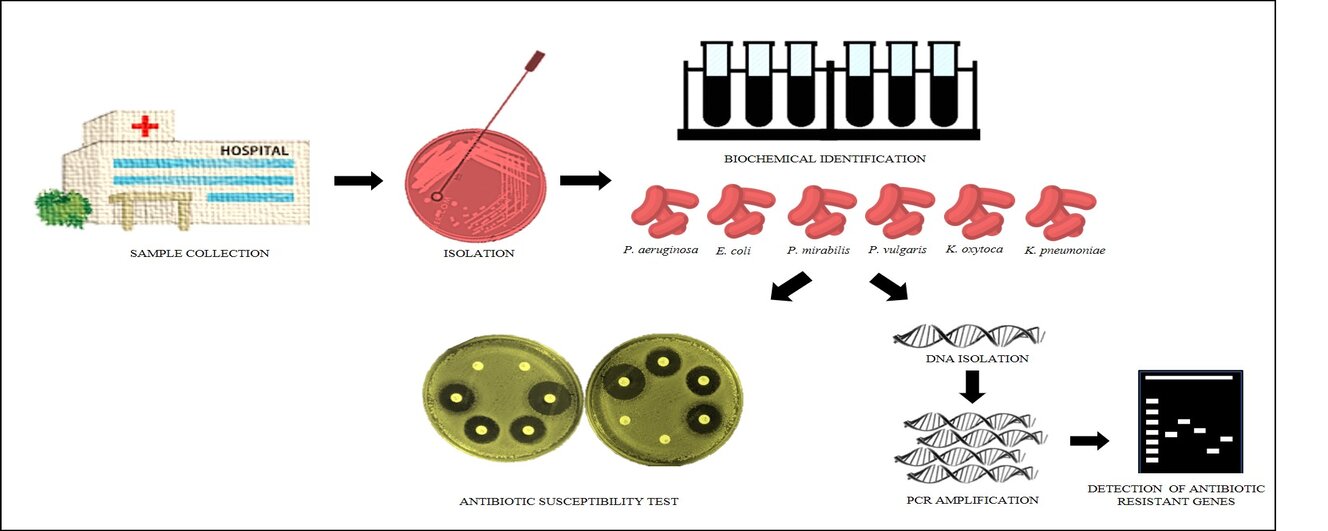
Downloads
References
MI Hutchings, AW Truman and B Wilkinson. Antibiotics: Past, present and future. Curr. Opin. Microbiol. 2019; 51,72-80.
H Landecker. Antibiotic resistance and the biology of history. Body Soc. 2016; 22,19-52.
M Sharma, B Eriksson, G Marrone, S Dhaneria and CS Lundborg. Antibiotic prescribing in two private sector hospitals; one teaching and one non-teaching: A cross-sectional study in Ujjain, India. BMC Infect. Dis. 2012; 12, 155.
SL Gorbach. The need to improve antimicrobial use in agriculture. Clin. Infect. Dis. 2002; 34, S73-S144.
IF Jaja, J Oguttu, E Green, V Muchenje. Prevalence and distribution of antimicrobial resistance determinants of Escherichia coli isolates obtained from meat in South Africa. PloS One 2019; 15, e0216914.
C Dahms, NO Hübner, A Kossow, A Mellmann, K Dittmann and A Kramer. Occurrence of ESBL-producing Escherichia coli in livestock and farm workers in Mecklenburg-Western Pomerania, Germany. PloS One 2015; 10, e0143326.
HC Neu. The crisis in antibiotic resistance. Science 1992; 257, 1064-73.
Z Pang, R Raudonis, BR Glick, TJ Lin and Z Cheng. Antibiotic resistance in Pseudomonas aeruginosa: Mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 2019; 37, 177-92.
G Kaufman. Antibiotics: Mode of action and mechanisms of resistance. Nurs. Stand. 2011; 25, 49-55.
GD Wright. Molecular mechanisms of antibiotic resistance. Chem. Commun. 2011; 14, 4055-61.
JMA Blair, MA Webber, AJ Baylay, DO Ogbolu and LJV Piddock. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015; 1, 42-51.
MD Iseman. Treatment of multidrug-resistant tuberculosis. N. Engl. J. Med. 1993; 11, 784-91.
SB Levy and B Marshall. Antibacterial resistance worldwide: Causes, challenges and responses. Nat. Med. 2004; 10, S122-S129.
M Bassetti, E Righi, A Vena, E Graziano E, A Russo and M Peghin. Risk stratification and treatment of ICU-acquired pneumonia caused by multidrug-resistant/extensively drug-resistant/pandrug-resistant bacteria. Curr. Opin. Crit. Care 2018; 24, 385-93.
L Peters, L Olson, DT Khu, S Linnros, NK Le, H Hanberger, NT Hoang, DM Tran and M Larsson. Multiple antibiotic resistance as a risk factor for mortality and prolonged hospital stay: A cohort study among neonatal intensive care patients with hospital-acquired infections caused by gram-negative bacteria in Vietnam. PloS One 2019; 14, e0215666.
S Gandra, KK Tseng, A Arora, B Bhowmik, ML Robinson, B Panigrahi, R Laxminarayan and EY Klein. The mortality burden of multidrug-resistant pathogens in india: A retrospective, observational study. Clin. Infect. Dis. 2019; 69, 563-70.
AS Jarrell, RM Kruer, LD Berescu, PJ Pronovost and JB Trivedi. Factors associated with in-hospital mortality among critically ill surgical patients with multidrug-resistant Gram-negative infections. J. Crit. Care 2018; 43, 321-6.
World Health Organization. Antimicrobial resistance: Global report on surveillance. World Health Organization, Geneva, Switzerland, 2014.
AD Khosravi, H Hoveizavi, A Mohammadian, A Farahani and A Jenabi. Genotyping of multidrug-resistant strains of Pseudomonas aeruginosa isolated from burn and wound infections by ERIC-PCR. Acta Cir. Bras. 2016; 31, 206-11.
R Sheridan, J Weber, P Chang, J Schulz, J Goverman, J Friedstat, C Duran, J Benjamin and M Pasternack. Multi-drug resistant gram negative bacteria colonization and infection in burned children: Lessons learned from a 20-year experience. Burns Open 2018; 2, 43-6.
HM Zowawi, PN Harris, MJ Roberts, PA Tambyah, MA Schembri, MD Pezzani, DA Williamson and DL Paterson. The emerging threat of multidrug-resistant Gram-negative bacteria in urology. Nat. Rev. Urol. 2015; 12, 570-84.
M Ali, KA Garba and MS Abdallah. Antibiotic susceptibility profile of bacteria responsible for urinary tract infection (UTI). S. Asian J. Biol. Res. 2018; 1, 12-27.
BJ Gardiner, AJ Stewardson, IJ Abbott and AY Peleg. Nitrofurantoin and fosfomycin for resistant urinary tract infections: Old drugs for emerging problems. Aust. Prescr. 2019; 42, 14-9.
FJ Pérez-Pérez and ND Hanson. Detection of plasmid-mediated AmpC β-lactamase genes in clinical isolates by using multiplex PCR. J. Clin. Microbiol. 2002; 6, 2153-62.
G Cuzon, T Naas, P Bogaerts, Y Glupczynski and P Nordmann. Evaluation of a DNA microarray for the rapid detection of extended-spectrum β-lactamases (TEM, SHV and CTX-M), plasmid-mediated cephalosporinases (CMY-2-like, DHA, FOX, ACC-1, ACT/MIR and CMY-1-like/MOX) and carbapenemases (KPC, OXA-48, VIM, IMP and NDM). J. Antimicrob. Chemoth. 2012; 8, 1865-9.
T Tada, JI Sekiguchi, S Watanabe, K Kuwahara-Arai, N Mizutani, I Yanagisawa, T Hishinuma, KN Zan, S Mya, HH Tin and T Kirikae. Assessment of a newly developed immunochromatographic assay for NDM-type metallo-β-lactamase producing Gram-negative pathogens in Myanmar. BMC Infect. Dis. 2019; 19, 565.
PM Bennett. Integrons and gene cassettes: A genetic construction kit for bacteria. J. Antimicrob. Chemoth. 1999; 43, 1-4.
MC Ploy, T Lambert, JP Couty and F Denis. Integrons: An antibiotic resistance gene capture and expression system. Clin. Chem. Lab. Med. 2000; 6, 483-7.
HR Goli, MR Nahaei, MA Rezaee, A Hasani, HS Kafil, M Aghazadeh and V Sheikhalizadeh. Prevalence and molecular characterization of class 1 integrons among clinical isolates of Pseudomonas aeruginosa in Northwest of Iran. Mol. Genet. Microbiol. Virol. 2017; 32, 109-15.
D Skurnik, AL Menac’h, D Zurakowski, D Mazel, P Courvalin, E Denamur, A Andremont and R Ruimy. Integron-associated antibiotic resistance and phylogenetic grouping of Escherichia coli isolates from healthy subjects free of recent antibiotic exposure. Antimicrob. Agents Chemother. 2005; 49, 3062-5.
R Coico. Gram staining. Curr. Protoc. Microbiol. 2006; 1, A.3O.1-A.3O.2.
MP Mishra, NK Debata and RN Padhy. Surveillance of multidrug resistant uropathogenic bacteria in hospitalized patients in Indian. Asian Pac. J. Trop. Biomed. 2013; 3, 315-24.
DH Bergey, FC Harrison, RS Breed, BW Hammer and FM Huntoon. Bergey’s manual of determinative bacteriology. The Williams & Wilkins Co, Maryland. 1923.
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. Clinical and Laboratory Standards Institute, Pennsylvania, 2017.
D Gislin, D Sudarsanam, GA Raj and K Baskar. Antibacterial activity of soil bacteria isolated from Kochi, India and their molecular identification. J. Genet. Eng. Biotechnol. 2018; 16, 287-94.
M Sunde. Prevalence and characterization of class 1 and class 2 integrons in Escherichia coli isolated from meat and meat products of Norwegian origin. J. Antimicrob. Chemoth. 2005; 56, 1019-24.
P Nordmann, T Naas and L Poirel. Global spread of carbapenemase-producing enterobacteriaceae. Emerg. Infect. Dis. 2011; 17, 1791-8.
PS Maneesh, M Sowmiya, T Bharani, HN Madhavan and J Malathi. Characterization of antibiotic resistance profiles of ocular enterobacteriaceae isolates. Eur. J. Microbiol. Immunol. 2016; 6, 40-8.
AH Mondal, MT Siddiqui, I Sultan and QM Haq. Prevalence and diversity of bla TEM, bla SHV and bla CTX-M variants among multidrug resistant Klebsiella spp. from an urban riverine environment in India. Int. J. Environ. Health Res. 2019; 29, 117-29.
N Sharma, AK Gupta, G Walia and R Bakhshi. A retrospective study of the changing trends of antimicrobial resistance of Klebsiella pneumoniae isolated from urine samples over last 3 years (2012-2014). J. Nat. Sci. Biol. Med. 2016; 7, 39-42.
M Baziboroun, M Bayani, Z Poormontaseri, M Shokri, T Biazar. Prevalence and antibiotic susceptibility pattern of extended spectrum beta lactamases producing Escherichia coli isolated from outpatients with Urinary Tract Infections in Babol, Northern of Iran. Curr. Issues Pharm. Med. Sci. 2018; 31, 61-4.
M Exner, S Bhattacharya, B Christiansen, J Gebel, P Goroncy-Bermes, P Hartemann, P Heeg, C Ilschner, A Kramer, E Larson and W Merkens. Antibiotic resistance: What is so special about multidrug-resistant Gram-negative bacteria? GMS Hyg. Infect. Control. 2017; 12, Doc05.
L Johnson, A Sabel, WJ Burman, RM Everhart, M Rome, TD MacKenzie, J Rozwadowski, PS Mehler and CS Price. Emergence of fluoroquinolone resistance in outpatient urinary Escherichia coli isolates. Am. J. Med. 2008; 121, 876-84.
S Gandra, J Joshi, A Trett and L Sankhil. Scoping report on antimicrobial resistance in India. Center for Disease Dynamics Economics & Policy, Washington, DC, 2017.
P Bhatt, KR Rathi, S Hazra, A Sharma and V Shete. Prevalence of multidrug resistant Pseudomonas aeruginosa infection in burn patients at a tertiary care centre. Indian J. Burns 2015; 23, 56-9.
R Dawra, R Sharma, R Bachhiwal and A Vyas. High incidence of multidrug resistant Pseudomonas aeruginosa isolated from infected burn wounds in a tertiary hospital. Int. J. Curr. Microbiol. Appl. Sci. 2017; 6, 1134-9.
DM Drekonja, SE Beekmann, S Elliott, D Mukundan, H Polenakovik, ME Rosenthal, PD Tamma, PM Polgreen and SJ Weissman. Challenges in the management of infections due to carbapenem-resistant enterobacteriaceae. Infect. Control Hosp. Epidemiol. 2014; 35, 437-9.
VN Kulkarni and MV Kulkarni. Trends in antibiotic resistance among major bacterial pathogens isolated from different specimens at a tertiary care hospital in India. J. Adv. Med. Pharm. Sci. 2018; 18, 1-7.
S Verma, SS Thorat, K Bisure. Prevalence of carbapenem resistant enterobacteriaceae: A study in a tertiary care hospital in Mumbai. J. Evolution Med. Dent. Sci. 2018; 7, 4909-12.
XA Marialouis and A Santhanam. Antibiotic resistance, RAPD-PCR typing of multiple drug resistant strains of Escherichia coli from urinary tract infection (UTI). J. Clin. Diagn. Res. 2016; 10, DC05-DC09.
JY Madec, M Haenni, P Nordmann and L Poirel. Extended-spectrum β-lactamase/AmpC-and carbapenemase-producing Enterobacteriaceae in animals: A threat for humans? Clin. Microbiol. Infect. 2017; 23, 826-33.
G Rajivgandhi, M Maruthupandy, G Ramachandran, M Priyanga and N Manoharan. Detection of ESBL genes from ciprofloxacin resistant Gram negative bacteria isolated from urinary tract infections (UTIs). Front. Lab. Med. 2018; 2, 5-13.
MM Khatun, MM Sharif, AE Haque, S Arifin and KM Ali. Bacteriological profile of ESBL producing bacteria with their antibiotic resistance pattern. KYAMC J. 2019; 10, 7-12.
B Fouzia and AS Damle. Prevalence and characterization of extended spectrum beta-lactamase production in clinical isolates of Klebsiella pneumonia. J. Med. Microb. Diagn. 2015; 4, 182.
R Gupta, A Malik, M Rizvi, M Ahmed and A Singh. Epidemiology of multidrug-resistant Gram-negative pathogens isolated from ventilator-associated pneumonia in ICU patients. J. Glob. Antimicrob. Resist. 2017; 9, 47-50.
VR Gajamer, A Bhattacharjee, D Paul, B Ingti, A Sarkar, J Kapil, AK Singh, N Pradhan and HK Tiwari. High prevalence of carbapenamase genes, AmpC genes and aminoglycoside resistance genes in extended spectrum β-lactamase positive uropathogens from Northern India. J. Glob. Antimicrob. Resist. 2020; 20, 197-203.
Z Cizmeci, E Aktas, B Otlu, O Acikgoz and S Ordekci. Molecular characterization of carbapenem-resistant Enterobacteriaceae yields increasing rates of NDM-1 carbapenemases and colistin resistance in an OXA-48-endemic area. J. Chemother. 2017; 29, 344-50.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.







