PCR-based SNP Markers for Sex Identification in Date Palm (Phoenix dactylifera L.) cv. KL1
DOI:
https://doi.org/10.48048/tis.2021.26Keywords:
Date palm, Markers, PCR, Sex determination, Single Nucleotide Polymorphisms (SNP)Abstract
The date palm (Phoenix dactylifera L.) is a valuable commercial plant in the Arecaceae family. It is a sweet, edible fruit containing a lot of nutrients including carbs, sugar, protein, and potassium. Maejo 36 or KL1 is a Thai date palm with excellent fresh fruit quality that is mostly grown in Thailand's northern and northeastern areas. The main issue with date palm farming is that there is no reliable way to discriminate between male and female seedlings prior to flowering, which takes 3 - 5 years after planting. An increase in the number of female date palm plants in commercial orchards may increase in date production and profitable investment. The present research aims to develop Single Nucleotide Polymorphisms (SNP) markers for the sex identification of date palm cv. KL1. The SNP markers were designed by analysis of nucleotide sequence difference from GenBank (accession number GL746268.1). The results revealed that SNP primers; PH02F - PdaNu01 (PdaNU02 or PdaNU03) - PdaNU05 used in multiplex PCR technique can precisely distinguish the sex of date palm by generating 2 different sizes of PCR-products in male plants, and only 1 size of PCR-product in female plants. This research indicates that SNP markers with multiplex-PCR technique are a high potential method for identifying the sex of date palm cv. KL1 may be used to increase female plants for cultivation at the seedling stage, which is highly beneficial for commercial date palm production.
HIGHLIGHTS
- Date palm is currently an economic fruits crops in Thailand
- The sex of date palm can be identified by phenotype in flowering stage until five years after planting
- The amplicon patterns in male and female date palm generated by SNP marker are applicable to predict their genders at an early seedling stage
- The use of SNP marker is highly beneficial for commercial date palm production
GRAPHICAL ABSTRACT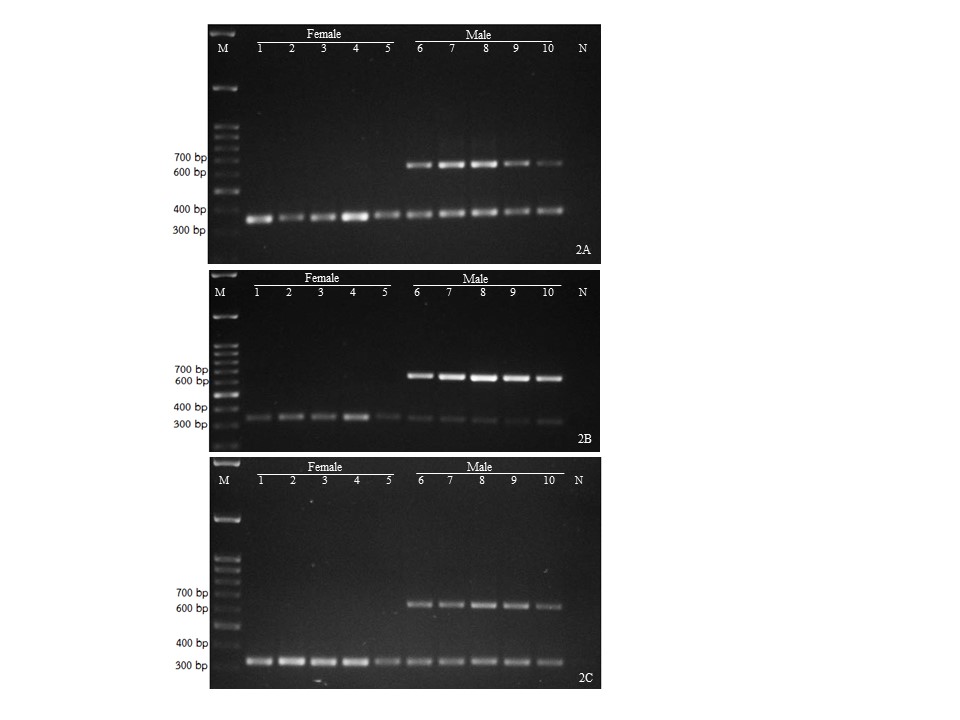
Downloads
References
CT Chao and RR Krueger. The date palm (Phoenix dactylifera L.): Overview of biology, uses, and cultivation. HortScience 2007; 42, 1077-82.
W Al-shahib and RJ Marshall. The fruit of the date palm: Its possible use as the best food for the future? Int. J. Food Sci. Nutr. 2003; 54, 247-59.
SM Al-Orf, MH Ahmed, N Al-Atwai, H Al-Zaidi, A Dehwah and S Dehwah. Nutritional properties and benefits of the date fruits (Phoenix dactylifera L.). Bull. Natl. Nutr. Inst. Arab Republic Egypt 2012; 39, 97-129.
AME Sulieman, IAA Elhafise and AM Abdelrahim. Comparative study on five Sudanese date (Phoenix dactylifera L.) fruit cultivars. Food Nutr. Sci. 2012; 3, 1245-51.
PK Vayalil. Antioxidant and antimutagenic properties of aqueous extract of date fruit (Phoenix dactylifera L. Arecaceae). J. Agr. Food Chem. 2002; 50, 610-7.
F Biglari, AFM AlKarkhi and AM Easa. Antioxidant activity and phenolic content of various date palm (Phoenix dactylifera) fruits from Iran. Food Chem. 2008; 107, 1636-41.
AR Tapas, DM Sakarkar and RB Kakde. Flavonoids as nutraceuticals: A review. Trop. J. Pharmaceut. Res. 2008; 7, 1089-99.
N Intha and P Chaiprasart. Sex determination in date palm (Phoenix dactylifera L.) by PCR based marker analysis. Sci. Hortic. 2018; 236, 251-5.
SA Bekheet and M Hanafy. Towards sex determination of date palm. In: SM Jain, JM Al-Khayri and DV Johnson (Eds.). Date palm biotechnology. Springer, Dordrecht, 2011, p. 551-66.
PJ Ainsworth and V Buchanan-Wollaston. Sex determination in plants. Curr. Top. Dev. Biol. 1998; 38, 167-223.
SM Jain. Date palm biotechnology: Current status and prospective-an overview. Emir. J. Food Agr. 2012; 24, 386-99.
D Johnson. Introduction: date palm biotechnology from theory to practice. In: SM Jain, JM Al-Khayri and DV Johnson (Eds.). Date Palm Biotechnology. Springer, Dordrecht, 2011, p. 1-11.
S Siljak-Yakovlev, M Cerbah, A Sarr, S Benmalek, N Bounaga, TCDL Pena and S Brown. Chromosomal sex determination and heterochromatin structure in date palm. Sex. Plant Reprod. 1996; 9, 127-32.
RAA Younis, OM Ismail and SS Soliman. Identification of sex-specific DNA markers for date Palm (Phoenix dactylifera L.) using RAPD and ISSR techniques. Res. J. Agr. Biol. Sci. 2008; 4, 278-84.
EK Al-Dous, B George, ME Al-Mahmoud, MY Al-Jaber, H Wang, YM Salameh, EK Al-Azwani, S Chaluvadi, AC Pontaroli, J DeBarry, V Arondel, J Ohlrogge, IJ Saie, KM Suliman-Elmeer, JL Bennetzen, RR Kruegger and JA Malek. De novo genome sequencing and comparative genomics of date palm (Phoenix dactylifera). Nat. Biotechnol. 2011; 29, 521-8.
E Cherif, S Zehdi-Azouzi, A Crabos, K Castillo, N Chabrillange, JC Pintaud, A Salhi-Hannachi, S Glemin and F Aberlenc-Bertossi. Evolution of sex chromosomes prior to speciation in the dioecious Phoenix species. J. Evol. Biol. 2016; 29, 1513-22.
S Kafkas, M Khodaeiaminjan, M Güney and E Kafkas. Identification of sex-linked SNP markers using RAD sequencing suggests ZW/ZZ sex determination in Pistacia vera L. BMC genom. 2015; 16, 98.
K Sujipuli, S Urtgam, N Kunpratum and T Jongjitvimo. Development of Specific-Molecular Marker for sex determination among papaya cultivars grown in Phitsanulok province, Thailand. NU. Int. J. Sci. 2016; 13, 1-10.
ME Al-Mahmoud, EK Al-Dous, EK Al-Azwani, and JA Malek. DNA-based assays to distinguish date palm (Arecaceae) gender. Am. J. Bot. 2012; 99, e7-e10.
JG Williams, AR Kubelik, KJ Livak, JA Rafalski and SV Tingey. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 1990; 18, 6531-5.
A Modi, K Suthar, P Thakkar, MC Mankad, S Kumari, S Narayanan, AS Singh and N Kumar. Evaluation of sex specific RAPD and SCAR markers linked to papaya (Carica papaya L.). Biocatal. Agr. Biotechnol. 2018; 16, 271-6.
J Kumar, M Heikrujam, K Sharma and V Agrawal. SRAP and SSR marker-assisted genetic diversity, population structure analysis and sex identification in Jojoba (Simmondsia chinensis). Ind. Crop. Prod. 2019; 133, 118-32.
Y Zhou, W Wu, Z Ning and R Zhou. Identification and characterization of sex-specific markers in the milky mangrove Excoecaria agallocha using double digest restriction site-associated DNA sequencing. Aquat. Bot. 2018; 144, 54-60.
MED Solliman, HAA Mohasseb, AA Al-Khateeb, SA Al-Khateeb, K Chowdhury, HA El-Shemy and MI Aldaej. Identification and sequencing of Date-SRY Gene: A novel tool for sex determination of date palm (Phoenix dactylifera L.). Saudi J. Biol. Sci. 2019; 26, 514-23.
PY Kwok. Methods for genotyping single nucleotide polymorphisms. Annu. Rev. Genomics Hum. Genet. 2001; 2, 235-258.
N Urasaki, K Tarora, A Shudo, H Ueno, M Tamaki, N Miyagi, S Adaniya and H Matsumura. Digital transcriptome analysis of putative sex-determination genes in papaya (Carica papaya). PLoS One. 2012; 7, e40904.
Y Hong, MK Pandey, Y Liu, X Chen, H Liu, RK Varshney, X Liang and S Huang. Identification and evaluation of single-nucleotide polymorphisms in allotetraploid peanut (Arachis hypogaea L.) based on amplicon sequencing combined with high resolution melting (HRM) analysis. Front. Plant Sci. 2015; 6, 1068.
A Ribeiro, A Golicz, CA Hackett, I Milne, G Stephen, D Marshall, AJ Flavell and M Bayer. An investigation of causes of false positive single nucleotide polymorphisms using simulated reads from a small eukaryote genome. BMC Bioinformatics 2015; 16, 382.
R Horn, A Radanovic, L Fuhrmann, Y Sprycha, S Hamrit, M Jockovic, D Miladinovic and C Jansen. Development and validation of markers for the fertility restorer gene Rf1 in Sunflower. Int. J. Mol. Sci. 2019; 20, 1260.
T Koyama, A Ozaki, K Yoshida, J Suzuki, K Fuji, JY Aoki, W Kai, Y Kawabata, T Tsuzaki, K Araki and T Sakamoto. Identification of sex-linked SNPs and sex-determining regions in the yellowtail genome. Mar. Biotechnol. 2015; 17, 502-10.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.







