Expression and in silico Analysis of CIDRα1 Recombinant Protein from Plasmodium Falciparum as a Malaria Subunit Vaccine Candidate
DOI:
https://doi.org/10.48048/tis.2022.1621Keywords:
CIDRα, In silico, Malaria vaccine, Plasmodium falciparum, Recombinant proteinAbstract
Malaria vaccination is an essential approach to combat malaria. One major protein studied for vaccine development is Plasmodium falciparum erythrocyte membrane protein-1 (PfEMP1). It contains several important domains for malaria pathogenesis. The binding of Cysteine-rich interdomain region α1 (CIDRα1) of PfEMP1 to endothelial protein C receptor (EPCR) is associated with cerebral malaria, while CIDRα1 binding to CD36 has been correlated with uncomplicated malaria. The vital function of CIDRα1 of PfEMP1 makes it a potential vaccine candidate to prevent clinical features of malaria. A long journey of vaccine development can be shortened by the advancement of bioinformatics and biotechnology techniques. This study aimed to express the recombinant CIDRα1 of PfEMP1 and investigate its potency as a malaria subunit vaccine candidate by in silico analysis. Constructed CIDRα1-PfEMP1 was expressed in E. coli BL21(DE3) after induction with Isopropyl ß-D-1-thiogalactopyranoside (IPTG) and purified using Ni-NTA column. In silico analysis on CIDRα1 of PfEMP1 sequence was conducted using ProtParam Tool for its physicochemical properties, Iterative Threading ASSEmbly Refinement (I-TASSER) server and JPred4 program to predict secondary structure, 3D modelling, and ligand-binding site, BepiPred 2.0 and Kolaskar-Tangaonkar to predict B-cell epitope, NetCTL server to determine T-cell epitope, and Vaxijen v2.0 server to predict its antigenicity. The chimeric CIDRα1 of PfEMP1 protein had a 27 kDa molecular weight and was classified as a stable protein. The secondary structure consisted of 6 helices connected with loops. It revealed similarity to CD36-binding protein, EPCR-binding domain, and protein involved in rosetting. The 3D structure modelling demonstrated conserved ligand-binding sites and accessible surface area, which are vital for receptor binding. It had B-cell and T-cell epitopes and was non-allergenic. The properties of the chimeric CIDRα of PfEMP1 indicated its potential as a malaria subunit vaccine candidate.
HIGHLIGHTS
- The binding capacity of CIDRα1 of PfEMP1 to endothelial protein C receptor (EPCR) and CD36 makes it a potential vaccine candidate to prevent clinical malaria
- The chimeric CIDRα1 of PfEMP1 protein was a stable protein and showed similarity to CD36-binding protein, EPCR-binding domain, and protein involved in rosetting, which demonstrated conserved ligand-binding sites and accessible surface area, which are vital for receptor binding
- The chimeric CIDRα1 of PfEMP1 protein had B-cell and T-cell epitopes and was non-allergenic in in silico analysis, indicating its potential as a malaria vaccine candidate
GRAPHICAL ABSTRACT 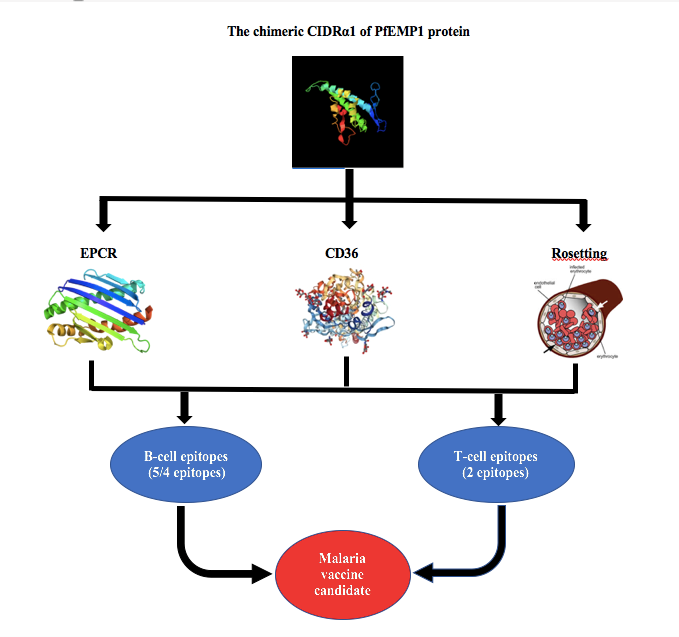
Downloads
References
World Health Organization (WHO) 2020. World Malaria Report 2020, Available at: https://www.who.int/teams/global-malaria-programme/reports/world-malaria-report-2020, accessed October 2021.
JD Smith. The role of PfEMP1 adhesion domain classification in Plasmodium falciparum pathogenesis research. Mol. Biochem. Parasitol. 2014; 195, p. 82-7.
N Obeng-Adjei, DB Larremore, L Turner, A Ongoiba, S Li, S Doumbo, TB Yazew, K Kayentao, LH Miller, B Traore, SK Pierce, CO Buckee, T Lavstsen, PD Crompton and TM Tran. Longitudinal analysis of naturally acquired PfEMP1 CIDR domain variant antibodies identifies associations with malaria protection. JCI Insight 2020; 5, PMC7406271.
JD Smith, G Subramanian, B Gamain and DI ML Baruch. Classification of adhesive domains in the Plasmodium falciparum erythrocyte membrane protein 1 family. Mol. Biochem. Parasitol. 2000; 110, 293-310.
MM Klein, AG Gittis, HP Su, MO Makobongo, JM Moore, S Singh, LH Miller and DN Garboczi. The cysteine-rich interdomain region from the highly variable Plasmodium falciparum erythrocyte membrane protein-1 exhibits a conserved structure. PLoS Pathog. 2008; 4, e1000147.
L Turner, T Lavstsen, BP Mmbando, CW Wang, PA Magistrado, LS Vestergaard, DS Ishengoma, DTR Minja, JP Lusingu and TG Theander. IgG antibodies to endothelial protein C receptor-binding cysteine-rich interdomain region domains of Plasmodium falciparum erythrocyte membrane protein 1 are acquired early in life in individuals exposed to malaria. Infect. Immun. 2015; 83, 3096-103.
CKY Lau, L Turner, JS Jespersen, ED Lowe, B Petersen, CW Wang, JEV Petersen, J Lusingu, TG Theander, T Lavstsen and MK Higgins. Structural conservation despite huge sequence diversity allows EPCR binding by the pfemp1 family implicated in severe childhood malaria. Cell Host Microbe 2015; 17, 118-29.
C Harmsen, L Turner, S Thrane, AF Sander, TG Theander and T Lavstsen. Immunization with virus-like particles conjugated to CIDRα1 domain of Plasmodium falciparum erythrocyte membrane protein 1 induces inhibitory antibodies. Malar J. Bio. Med. Central 2020; 19, 1-11.
M Bernabeu, SA Danziger, M Avril, M Vaz, PH Babar, AJ Brazier, T Herricks, JN Maki, L Pereira, A Mascarenhas, E Gomes, L Chery, JD Aitchison, PK Rathod and JD Smith. Severe adult malaria is associated with specific PfEMP1 adhesion types and high parasite biomass. Proc. Natl. Acad. Sci. U.S.A. 2016; 113, E3270-E3279.
A Kessler, S Dankwa, M Bernabeu, V Harawa, SA Danziger, F Duffy, SD Kampondeni, MJ Potchen, N Dambrauskas, V Vigdorovich, BG Oliver, SE Hochman, WB Mowrey, IJC MacCormick, WL Mandala, SJ Rogerson, DN Sather, JD Aitchison, TE Taylor, KB Seydel, JD Smith and K Kim. Linking EPCR-binding PfEMP1 to brain swelling in pediatric cerebral malaria. Cell Host Microbe 2017; 22, 601-14.
JS Jespersen, CW Wang, SI Mkumbaye, DT Minja, B Petersen, L Turner, JEV Petersen, JPA Lusingu, TG Theander and T Lavstsen. Plasmodium falciparum var genes expressed in children with severe malaria encode CIDR α1 domains. EMBO Mol. Med. 2016; 8, 839-50.
E Gasteiger, C Hoogland, A Gattiker, S Duvaud, MR Wilkins, RD Appel and A Bairoch. The proteomics protocols handbook. Humana Press, New Jersey, United States, 2005, p. 571-608.
Y Zhang. I-TASSER server for protein 3D structure prediction. BMC Bioinform. 2008; 9, 1-8.
A Roy, A Kucukural and Y Zhang. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc. 2010; 5, 725-38.
J Yang, R Yan, A Roy, D Xu, J Poisson and Y Zhang. The I-TASSER suite: Protein structure and function prediction. Nat. Methods 2014; 12, 7-8.
A Drozdetskiy, C Cole, J Procter and GJ Barton. JPred4: A protein secondary structure prediction server. Nucleic Acids Res. 2015; 43, W389-W394.
B Petersen, TN Petersen, P Andersen, M Nielsen and C Lundegaard. A generic method for assignment of reliability scores applied to solvent accessibility predictions. BMC Struct. Biol. 2009; 9, 1-10.
J Yang, Y Wang and Y Zhang. ResQ: An approach to unified estimation of B-factor and residue-specific error in protein structure prediction. J. Mol. Biol. 2016; 428, 693-701.
C Zhang, PL Freddolino and Y Zhang. COFACTOR: Improved protein function prediction by combining structure, sequence and protein-protein interaction information. Nucleic Acids Res. 2017; 45, W291-W299.
MC Jespersen, B Peters, M Nielsen and P Marcatili. BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 2017; 45, W24-W29.
AS Kolaskar and PC Tongaonkar. A semi-empirical method for prediction of antigenic determinants on protein antigens. FEBS Lett. 1990; 276, 172-4.
MV Larsen, C Lundegaard, K Lamberth, S Buus, O Lund and M Nielsen. Large-scale validation of methods for cytotoxic T-lymphocyte epitope prediction. BMC Bioinform. 2007; 8, 1-12.
B Peters, S Bulik, R Tampe, PM van Endert and HG Holzhütter. Identifying MHC class I epitopes by predicting the TAP transport efficiency of epitope precursors. J. Immunol. 2003; 171, 1741-9.
IA Doytchinova and DR Flower. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 2007; 8, 1-7.
M Mo, CL Hooi, M Kotaka, M Niang, X Gao, JK Iyer, J Lescar and P Preiser. The C-terminal segment of the cysteine-rich interdomain of Plasmodium falciparum erythrocyte membrane protein 1 determines CD36 binding and elicits antibodies that inhibit adhesion of parasite-infected erythrocytes. Infect. Immun. 2008; 76, 1837-47.
FL Hsieh, L Turner, JR Bolla, CV Robinson, T Lavstsen and MK Higgins. The structural basis for CD36 binding by the malaria parasite. Nat. Commun. 2016; 7, 1-11.
TS Rask, DA Hansen, TG Theander, AG Pedersen and T Lavstsen. Plasmodium falciparum erythrocyte membrane protein 1 diversity in seven genomes - divide and conquer. PLoS Comput. Biol. 2010; 6, e1000933.
JA Christopher, J Brown, AS Doré, JC Errey, M Koglin, FH Marshall, DG Myszka, RL Rich, CG. Tate, B Tehan, T Warne and M Congreve. Biophysical fragment screening of the β1-adrenergic receptor: Identification of high affinity arylpiperazine leads using structure-based drug design. J. Med. Chem. 2013; 56, 3446-55.
National Library of Medicine. Compound Summary, Available at: https://pubchem.ncbi.nlm. nih.gov/compound/49835947, accessed February 2022.
JL Sanchez-Trincado, M Gomez-Perosanz and PA Reche. Fundamentals and methods for T- and B-cell epitope prediction. J. Immunol. Res. 2017; 2017, 2680160.
J Heide, KC Vaughan, A Sette, T Jacobs and JS Zur Wiesch. Comprehensive review of human plasmodium falciparum-specific CD8+ T cell epitopes. Front Immunol. 2019; 10, 1-23.
K Miura. Progress and prospects for blood-stage malaria vaccines. Expert Rev. Vaccines 2016; 15, 765-81.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2022 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.