Temporal Immune Trajectory and Candidate Biomarkers in Sepsis Associated with Community-Acquired Pneumonia: A Systematic Review
DOI:
https://doi.org/10.48048/tis.2026.13450Keywords:
Community-acquired pneumonia, Sepsis, Transcriptomic, BiomarkerAbstract
Community-acquired pneumonia (CAP) with sepsis remains a leading cause of morbidity and mortality worldwide. Although numerous immune biomarkers have been proposed, integrated evidence describing how immune responses evolve in CAP-associated sepsis is limited, particularly from blood transcriptomic studies. This systematic review aimed to synthesize evidence on temporal immune trajectories and identify candidate transcriptomic biomarkers associated with outcomes in CAP-associated sepsis. A systematic search of medRxiv, PubMed, and CENTRAL was conducted for studies reporting serial blood transcriptomic or immune biomarker data in adult patients with CAP-associated sepsis. Studies with clearly defined timepoints and clinical outcomes were included. Risk of bias was assessed using an adapted Newcastle-Ottawa Scale with transcriptomic-specific criteria. Due to heterogeneity in study design and timing definitions, findings were integrated using structured narrative synthesis. Six studies met the inclusion criteria. Early sepsis was characterized by consistent upregulation of innate inflammatory pathways, including neutrophil activation, interferon-related genes, and cytokine signaling, largely independent of pathogen type. Survivors demonstrated recovery of antigen presentation and adaptive immune pathways at later time points, whereas non-survivors exhibited persistent inflammatory signatures, impaired lymphocyte-associated transcription, immune checkpoint activation, and endothelial dysfunction. Recurrent candidate biomarkers included inflammatory gene signatures, immune endotype classifications, and transcriptomic indicators of adaptive immune suppression. CAP-associated sepsis follows dynamic immune trajectories rather than a single inflammatory state. Temporal transcriptomic patterns may help identify patients at risk of immune failure and guide immunomodulatory strategies. However, heterogeneity in timing definitions, small sample sizes, and limited validation underscore the need for prospective studies before clinical implementation.
HIGHLIGHTS
- This systematic review analyzed defined temporal immune trajectories in CAP-associated sepsis using blood transcriptomic studies.
- Early hyperinflammation was pathogen-independent and not predictive of mortality.
- Failure of antigen presentation and Th1 immunity characterized poor outcomes.
- Immune checkpoint activation and endothelial dysfunction defined immunoparalysis in non-survivors.
- Candidate transcriptomic biomarkers reflect immune competence and exhaustion.
GRAPHICAL ABSTRACT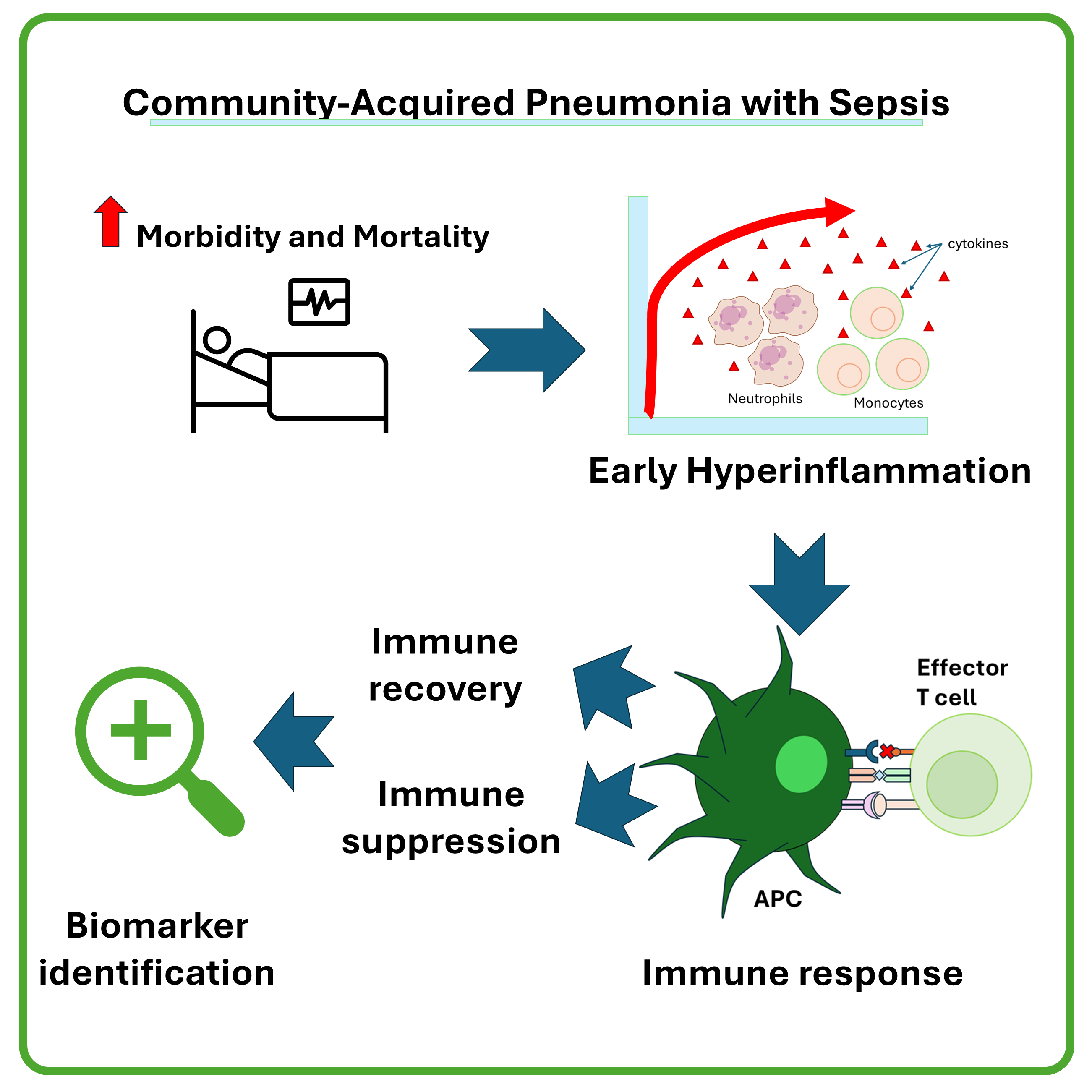
Downloads
References
JP Metlay, GW Waterer, AC Long, A Anzueto, J Brozek, K Crothers, LA Cooley, NC Dean, MJ Fine, SA Flanders, MR Griffin, ML Metersky, DM Musher, MI Restrepo and CG Whitney. Diagnosis and treatment of adults with community-acquired pneumonia. An official clinical practice guideline of the American Thoracic Society and Infectious Diseases Society of America. American Journal of Respiratory and Critical Care Medicine 2019; 200(7), e45-e67.
MN Lutfiyya, E Henley, LF Chang and SW Reyburn. Diagnosis and treatment of community-acquired pneumonia. American Family Physician 2006; 73(3), 442-450.
J Womack and J Kropa. Community-acquired pneumonia in adults: Rapid evidence review. American Family Physician 2022; 105(6), 625-630.
MN Lutfiyya, E Henley, LF Chang and SW Reyburn. Diagnosis and treatment of community-acquired pneumonia. American Family Physician 2006; 73(3), 442-450.
SM Han, PS Hon, HY Na, TTH Yong, PA Tambyah and YT Wen. Viral non-SARS-CoV-2 etiology of community-acquired pneumonia (CAP) in Southeast Asia: a review and pooled analysis. IJID Regions 2025; 15, 100672.
Z Hu, J Lin, J Chen, T Cai, L Xia, Y Liu, X Song and Z He. Overview of viral pneumonia associated with influenza virus, respiratory syncytial virus, and coronavirus, and therapeutics based on natural products of medicinal plants. Frontiers in Pharmacology 2021; 12, 630834.
W Xu, Y Zhang, Y Gao and X Li. Impact of the corona virus disease 2019 pandemic on the prevalence of common respiratory pathogens in hospitalized young patients with non-severe community-acquired pneumonia. Journal of Thoracic Disease 2025; 17(10), 107216.
V Biradar and JL Moran. SIRS, sepsis and multiorgan failure. In: R Fitridge and M Thompson (Eds.). Mechanisms of vascular disease: A reference book for vascular specialists. University of Adelaide Press, South Australia, Australia, 2011.
L La Via, G Sangiorgio, S Stefani, A Marino, G Nunnari, S Cocuzza, I La Mantia, B Cacopardo, S Stracquadanio, S Spampinato, S Lavalle and A Maniaci. The global burden of sepsis and septic shock. Epidemiologia 2024; 5(3), 456-478.
Y Li, S Ren and S Zhou. Advances in sepsis research: Insights into signaling pathways, organ failure, and emerging intervention strategies. Experimental and Molecular Pathology 2025; 142, 104963.
MA Ovali and S Percin. Sepsis-associated immunosuppression: Mechanistic insights, biomarkers, and therapeutic perspectives. Molecular Biology Reports 2025; 53(1), 148.
L Stiel, A Gaudet, S Thietart, H Vallet, P Bastard, G Voiriot, M Oualha, B Sarton, H Kallel, N Brechot, L Kreitmann, S Benghanem, J Joffre and Y Jouan. Innate immune response in acute critical illness: A narrative review. Annals of Intensive Care 2024; 14(1), 137.
X Tao, J Wang, B Liu, P Cheng, D Mu, H Du and B Niu. Plasticity and crosstalk of mesenchymal stem cells and macrophages in immunomodulation in sepsis. Frontiers in Immunology 2024; 15, 1338744.
KL Burnham, EE Davenport, J Radhakrishnan, P Humburg, AC Gordon, P Hutton, E Svoren-Jabalera, C Garrard, AVS Hill, CJ Hinds and JC Knight. Shared and distinct aspects of the sepsis transcriptomic response to fecal peritonitis and pneumonia. American Journal of Respiratory and Critical Care Medicine 2017; 196(3), 328-339.
JG Chenoweth, J Brandsma, DA Striegel, P Genzor, E Chiyka, PW Blair, S Krishnan, E Dogbe, I Boakye, GB Fogel, EL Tsalik, CW Woods, A Owusu-Ofori, C Oppong, G Oduro, T Vantha, AG Letizia, CG Beckett, KL Schully and DV Clark. Sepsis endotypes identified by host gene expression across global cohorts. Communications Medicine 2024; 4(1), 120.
RE Hancock, A An, CC dos Santos and AH Lee. Deciphering sepsis: Transforming diagnosis and treatment through systems immunology. Frontiers in Science 2025; 2, 1469417.
YY Zhang and BT Ning. Signaling pathways and intervention therapies in sepsis. Signal Transduction and Targeted Therapy 2021; 6, 407.
P Severino, E Silva, GL Baggio-Zappia, MKC Brunialti, LA Nucci, O Rigato, IDCG da Silva, FR Machado and R Salomao. Patterns of gene expression in peripheral blood mononuclear cells and outcomes from patients with sepsis secondary to community acquired pneumonia. PLoS One 2014; 9(3), e91886.
E Kyriazopoulou, Y Hasin-Brumshtein, U Midic, G Poulakou, H Milionis, S Metallidis, M Astriti, A Fragkou, A Rapti, E Taddei, I Kalomenidis, G Chrysos, A Angheben, I Kainis, Z Alexiou, F Castelli, FS Serino, P Bakakos, E Nicastri, …, EJ Giamarellos-Bourboulis. Transitions of blood immune endotypes and improved outcome by anakinra in COVID-19 pneumonia: An analysis of the SAVE-MORE randomized controlled trial. Critical Care 2024; 28, 73.
AY An, A Baghela, P Zhang, R Falsafi, AH Lee, U Trahtemberg, AJ Baker, CC dos Santos and REW Hancock. Severe COVID-19 and non-COVID-19 severe sepsis converge transcriptionally after a week in the intensive care unit, indicating common disease mechanisms. Frontiers in Immunology 2023; 14, 1167917.
T Chew, TM Pelaia, AL Phu, S Teoh, Y Wang, N Deshpande, K Kim, V Herwanto, Gunawan, T Karvunidis, Y Zerbib, KR Short, S Macdonald, I Thevarajan, D Rinchai, WS Kuan, B Knippenberg, J Iredell, PN Britton and M Shojaei. Molecular landscape of respiratory infection: A large-scale, multi-centre blood transcriptome dataset. Scientific Data 2025; 12(1), 1175.
A Muratsu, S Oda, S Onishi, J Yoshimura, H Matsumoto, Y Togami, Y Mitsuyama, H Ito, D Okuzaki, H Ogura and J Oda. Bacterial sepsis causes more dramatic pathogenetic changes in the Th1 pathway than does viral (COVID-19) sepsis: A prospective observational study of whole blood transcriptomes. Virology Journal 2024; 21(1), 190.
WJ Wiersinga, SJ Leopold, DR Cranendonk and T van der Poll. Host innate immune responses to sepsis. Virulence 2014; 5(1), 36-44.
T van der Poll, FL van de Veerdonk, BP Scicluna and MG Netea. The immunopathology of sepsis and potential therapeutic targets. Nature Reviews Immunology 2017; 17(7), 407-420.
KH Lee, A Gordon and B Foxman. The role of respiratory viruses in the etiology of bacterial pneumonia. Evolution, Medicine, and Public Health 2016; 2016(1), 95-109.
MJ Delano and PA Ward. The immune system’s role in sepsis progression, resolution, and long-term outcome. Immunological Reviews 2016; 274(1), 330-353.
C Nedeva. Inflammation and cell death of the innate and adaptive immune system during sepsis. Biomolecules 2021; 11(7), 1011.
F Franco, A Jaccard, P Romero, YR Yu and PC Ho. Metabolic and epigenetic regulation of T-cell exhaustion. Nature Metabolism 2020; 2(10), 1001-1012.
J Brady, S Horie and JG Laffey. Role of the adaptive immune response in sepsis. Intensive Care Medicine Experimental 2020; 8(1), 20.
CM Padovani and K Yin. Immunosuppression in sepsis: Biomarkers and specialized pro-resolving mediators. Biomedicines 2024; 12(1), 175.
F Wang, Y Cui, D He, G Gong and H Liang. Natural killer cells in sepsis: Friends or foes? Frontiers in Immunology 2023; 14, 1011111.
L Chiche, JM Forel, G Thomas, C Farnarier, F Vely, M Blery, L Papazian and E Vivier. The role of natural killer cells in sepsis. BioMed Research International 2011; 2011(1), 986491.
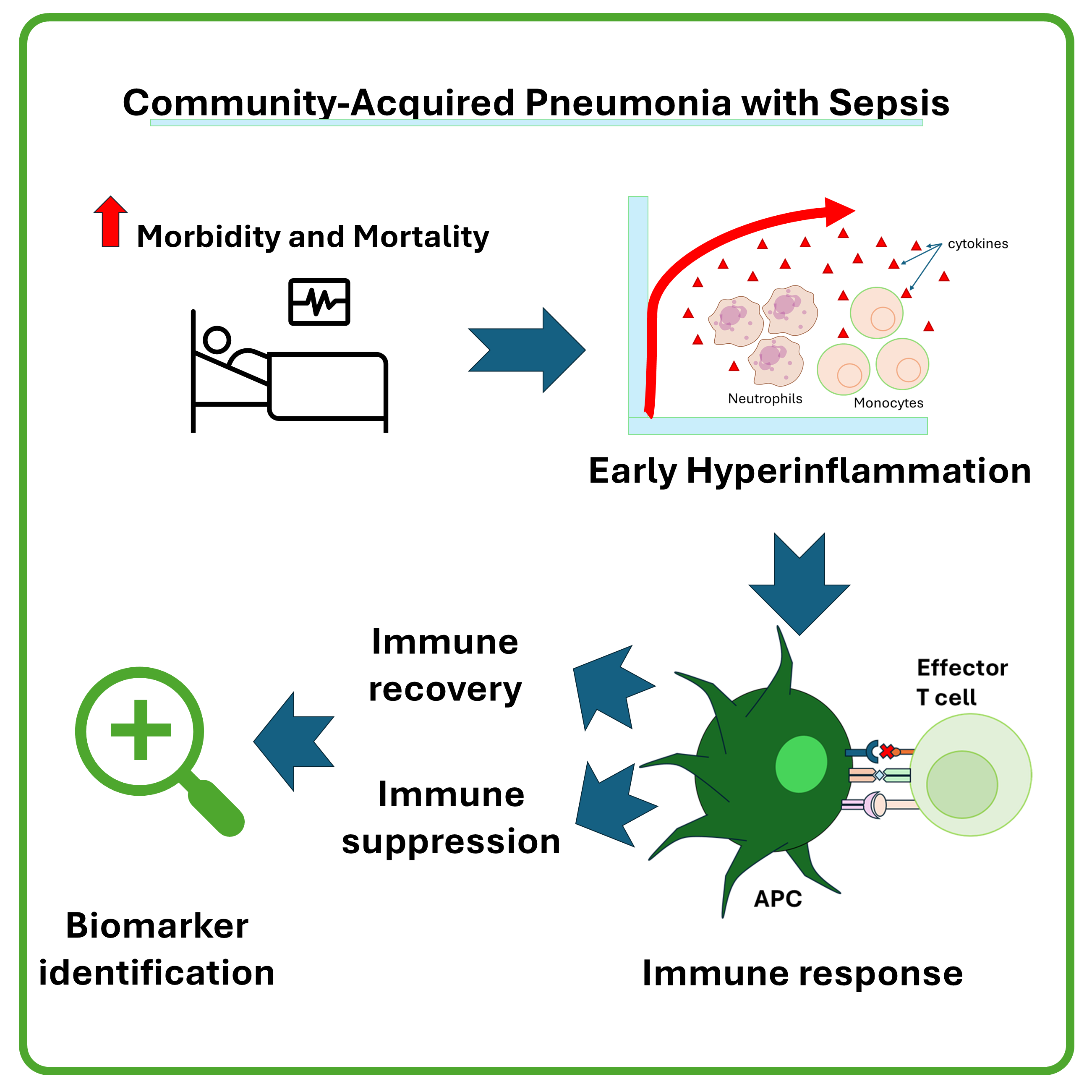
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






