Biodegradation Potential of Glyphosate by Rhizobacteria Isolated from Tithonia diversifolia: Characterization, Glyphosate Degrading, and Molecular Identification
DOI:
https://doi.org/10.48048/tis.2026.12865Keywords:
Bioremediation, Glyphosate, Isolation, Rhizobacteria, Tithonia diversifoliaAbstract
Glyphosate is a widely used broad-spectrum systemic herbicide that effectively controls weeds by inhibiting the synthesis of specific amino acids for the formation of plant proteins. Improper and repeated use can lead to the accumulation of residue in the soil, which may remain strongly absorbed over an extended period. Investigations into the biodegradation of glyphosate in soil under various environmental conditions are crucial for bioremediation efforts. Recently, a further step has been taken considering the use of rhizobacteria for the removal of glyphosate herbicide. Our research aimed to isolate naturally occurring rhizobacteria from Tithonia diversifolia (as a source of green manure) and assess the critical ecological factors influencing their growth and glyphosate degradation. Screening began by cultivating a one gram of T. diversifolia rhizosphere soil sample in Nutrient Broth (NB) media containing 15 mg mL−1 glyphosate for a week. The six isolates initially obtained were further subcultured in Nutrient agar (NA) containing nutrient agar and 15 mg mL−1 glyphosate. The morphological and physiological properties, phosphate solubilization, and IAA production were used in isolate characterization, in addition to glyphosate biodegradation. Only two isolates, TBr1 and TBr12, have shown the capacity to survive and biodegrade glyphosate. Both isolates exhibit optimal pH levels above 6.0 and optimal activity at 30 °C, demonstrating the fastest growth rates and abilities to break down glyphosate by 67% and 76%, respectively, within 7 days. Tested in a medium minus carbon, nitrogen, and phosphorus sources, both isolates showed the ability to hydrolyze glyphosate through CN and CP bonds. However, they had different CP lyase efficacy in metabolizing glyphosate. Based on 16S rDNA gene sequence analysis, TBr1 was identified as Burkholderia cepacia strain TBr1, and TBr12 as the Bacillus velezensis strain TBr12. Therefore, the ability of both isolates to degrade glyphosate, produce IAA, and dissolve phosphates makes them promising candidates for removing these emerging contaminants from the environment.
HIGHLIGHTS
- Glyphosate inhibits EPSP synthase, potentially damaging the microbiome and soil ecosystems.
- The research focus is on the isolation and characterization of glyphosate-degrading rhizobacteria from Tithonia diversifolia.
- The strains Burkholderia cepacia TBr1 and Bacillus velezensis TBr1 hydrolyze glyphosate by 67% and 76%, respectively, within one week at a concentration of 15 mg mL-1.
- Both strains have the potential to serve as bioremediation agents for glyphosate contamination.
GRAPHICAL ABSTRACT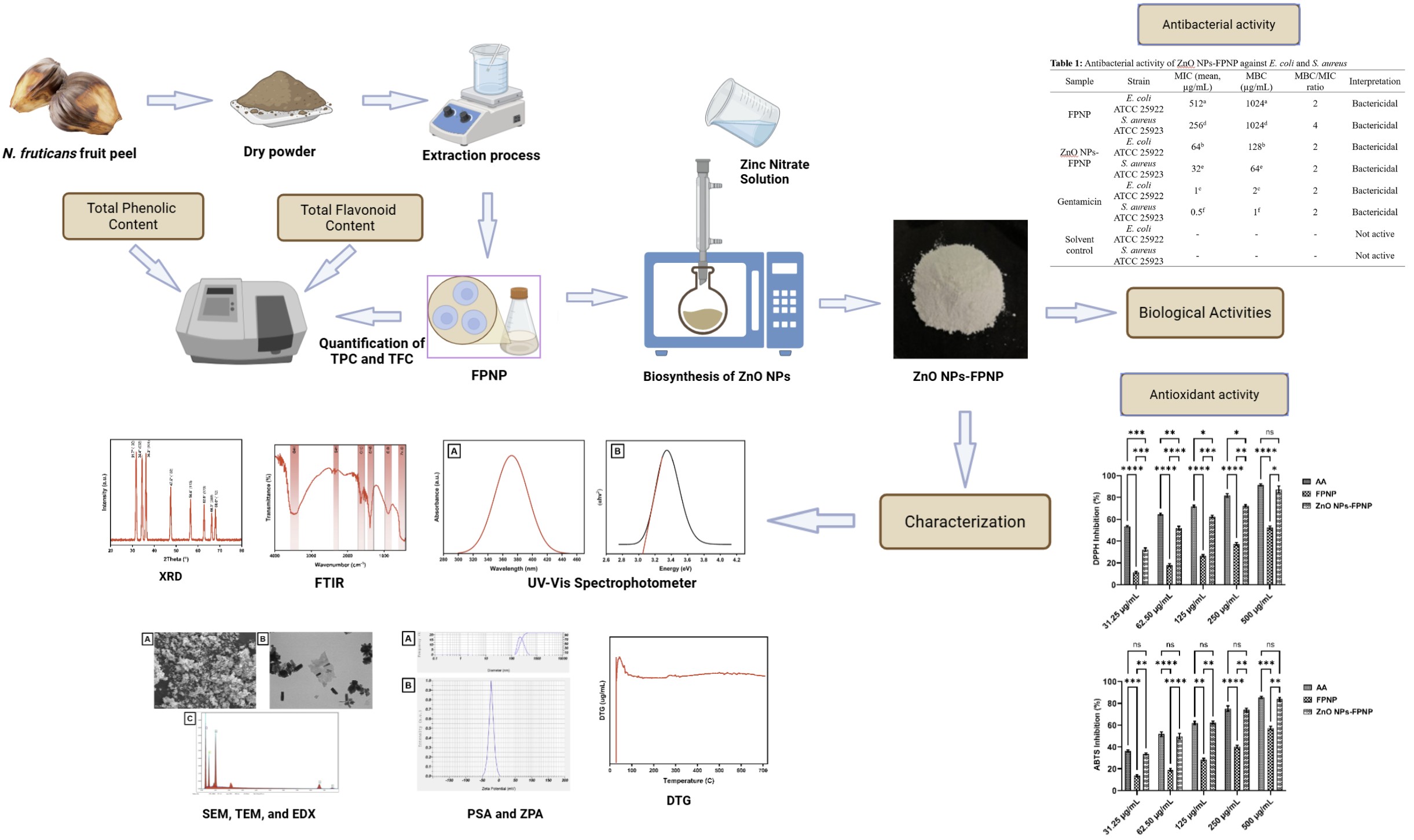
Downloads
References
F Maggi, DL Cecilia, FHM Tang and A McBratney. The global environmental hazard of glyphosate use. Science of The Total Environment 2020; 717, 137167.
JP Myers, MN Antoniou, B Blumberg, L Carroll, T Colborn, LG Everett, M Hansen, PJ Landrigan, BP Lanphear, R Mesnage, LN Vandenberg, FS vom Saal, WV Welshons and CM Benbrook. Concerns over use of glyphosate-based herbicides and risks associated with exposures: A consensus statement. Environmental Health 2016; 15, 19.
CM Benbrook. Trends in glyphosate herbicide use in the United States and globally. Environmental Sciences Europe 2016; 28, 3.
PM Dewick. The biosynthesis of shikimate metabolites. Natural Product Reports 1995; 12(6), 579-607.
AR Knaggs. The biosynthesis of shikimate metabolites. Natural Product Reports 1999; 16(4), 525-560.
JT Padilla and HM Selim. Chapter one - Environmental behavior of glyphosate in soils. Advances in Agronomy 2020; 159, 1-34.
AHC van Bruggen, MR Finckh, M He, CJ Ritsema, P Harkes, D Knuth and V Geissen. Indirect effects of the herbicide glyphosate on plant, animal and human health through its effects on microbial communities. Frontiers in Environmental Science 2021; 9, 763917.
P Chávez-Ortiz, Y Tapia-Torres, J Larsen and F Garcia-Olivia. Glyphosate-based herbicides alter soil carbon and phosphorus dynamics and microbial activity. Applied Soil Ecology 2022; 169, 104256.
VL Lozano and HN Pizarro. Glyphosate lessons: Is biodegradation of pesticides a harmless process for biodiversity? Environmental Sciences Europe 2024; 36, 55.
R Kanissery, B Gairhe, D Kadyampakeni, O Batumen and F Alferez. Glyphosate: Its environmental persistence and impact on crop health and nutrition. Plants 2019; 8(11), 499.
JA García-Pérez, E Alarcón, Y Hernández and C Hernandez. Impact of litter contaminated with glyphosate-based herbicide on the performance of Pontoscolex corethrurus, soil phosphatase activities and soil pH. Applied Soil Ecology 2016; 104, 31-41.
Y Sang, JC Mejuto, J Xiao and J Simal-Gandara. Assessment of glyphosate impact on the agrofood ecosystem. Plants 2021; 10(2), 405.
S Singh, V Kumar, JPK Gill, JPK Datta, S Singh, V Dhaka, D Kapoor, AB Wani, DS Dhanjal, M Kumar, SL Harikumar and J Singh. Herbicide glyphosate: Toxicity and microbial degradation. International Journal of Environmental Research and Public Health 2020; 17(20), 7519.
Y Chen, WJ Chen, Y Huang, J Li, J Zhong, W Zhang, Y Zou, S Mishra, P Bhatt and S Chen. Insights into the microbial degradation and resistance mechanisms of glyphosate. Environmental Research 2022; 215(1), 114153.
AV Sviridov, TV Shushkova, NF Zelenkova, NG Vinokurova, IG Morgunov, IT Ermakova and AA Leontievsky. Distribution of glyphosate and methylphosphonate catabolism systems in soil bacteria Ochrobactrum anthropi and Achromobacter sp. Applied Microbiology and Biotechnology 2012; 93, 787-796.
AN Moneke, GN Okpala and CU Anyanwu. Biodegradation of glyphosate herbicide in vitro using bacterial isolates from four rice fields. African Journal of Biotechnology 2010; 9(26), 4067-4074.
NE Ibrahim, V Sevakumaran and F Ariffin. Preliminary study on glyphosate-degrading bacteria isolated from agricultural soil. Environmental Advances 2023; 12, 100368.
ML Castrejón-Godínez, E Tovar-Sánchez, L Valencia-Cuevas, ME Rosas-Ramírez, A Rodríguez and P Mussali-Galante. Glyphosate pollution treatment and microbial degradation alternatives, a review. Microorganisms 2021; 9(11), 2322.
YV Kryuchkova, GL Burygin, NE Gogoleva, YV Gogolev, MP Chernyshova, OE Makarov, EE Fedorov and OV Turkovskaya. Isolation and characterization of a glyphosate-degrading rhizosphere strain, Enterobacter cloacae K7. Microbiological Research 2014; 169(1), 99-105.
G Saxena and RN Bharagava. Bioremediation of industrial waste for environmental safety. Springer, Singapore, 2020.
L Jia, Z Wang, L Ji, SD Neve, PC Struik, Y Yao, J Lv, T Zhou and K Jin. Keystone microbiome in the rhizosphere soil reveals the effect of long-term conservation tillage on crop growth in the Chinese Loess Plateau. Plant and Soil 2022; 473, 457-472.
T Macek, M Macková and J Káš. Exploitation of plants for the removal of organics in environmental remediation. Biotechnology Advances 2000; 18(1), 23-34.
J Montreemuk, TN Stewart and B Prapagdee. Bacterial-assisted phytoremediation of heavy metals: Concepts, current knowledge, and future directions. Environmental Technology & Innovation 2024; 33, 103488.
W Mohy-Ud-Din, MJ Akhtar, S Bashir, HN Asghar, MF Nawaz and F Chen. Isolation of glyphosate-resistant bacterial strains to improve the growth of maize and degrade glyphosate under axenic condition. Agriculture 2023; 13(4), 886.
N Hakim, R Alfina, Agustian, Hermansah and Yulnafatmawita. Bacterial inoculants to increase the biomass and nutrient uptake of Tithonia cultivated as hedgerow plants in ultisols. Malaysian Journal of Soil Science 2014; 18 115-123.
B Jama, CA Palm, RJ Buresh, A Niang, C Gachengo, G Nziguheba and B Amadalo. Tithonia diversifolia as a green manure for soil fertility improvement in western Kenya: A review. Agroforestry Systems 2000; 49, 201-221.
ST Partey. Effect of pruning frequency and pruning height on the biomass production of Tithonia diversifolia (Hemsl) A. Gray. Agroforestry Systems 2011; 83, 181-187.
NI Elarabi, AA Abdelhadi, RH Ahmed, I Saleh, IA Arif, G Osman and DS Ahmed. Bacillus aryabhattai FACU: A promising bacterial strain capable of manipulate the glyphosate herbicide residues. Saudi Journal of Biological Sciences 2020; 27(9), 2207-2214.
MR Jan, J Shah, M Muhammad and B Ara. Glyphosate herbicide residue determination in samples of environmental importance using spectrophotometric method. Journal of Hazardous Materials 2009; 169(1-3), 742-745.
J Murphy and JP Riley. A modified single solution method for the determination of phosphate in natural waters. Analytica Chimica Acta 1962; 27, 31-36.
M Sarwar, M Arshad, DA Martens and WT Frankerberger. Tryptophan-dependent biosynthesis of auxins in soil. Plant and Soil 1992; 147, 207-215.
NR Krieg, JT Staley, DR Brown, BP Hedlund, BJ Paster, NL Ward, W Ludwig and WB Whitman. Bergey’s manual of systematic bacteriology. 2nd ed. Springer, New York, 2010.
B Bearzatto, JF Durant, J Ambroise and JL Gala. Rapid, user-friendly, cost-effective DNA and library Preparation methods for whole-genome sequencing of bacteria with varying cell wall composition and GC content using minimal DNA on the illumina platform. BMC Genomics 2025; 26, 396.
AA Abdulla. Optimization of DNA extraction of Lactobacillus spp for Identification by tuf B gene –Based polymerase chain reaction. Journal of Biology, Agriculture and Healthcare 2014; 4(8), 122-127.
AA Dashti, MM Jadaon, AM Abdulsamad and HM Dashti. Heat treatment of bacteria: A simple method of DNA extraction for molecular techniques. Kuwait Medical Journal 2009; 41(2), 117-122.
N Saitou and M Nei. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Molecular Biology and Evolution 1987; 4(4), 406-425.
J Felsenstein. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985; 39(4), 783-791.
K Tamura, M Nei and S Kumar. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceedings of the National Academy of Sciences of the United States of America 2004; 101(30), 11030-11035.
S Kumar, G Stecher, M Suleski, M Sanderford, S Sharma and K Tamura. MEGA12 : Molecular evolutionary genetic analysis version 12 for adaptive and green computing. Molecular Biology and Evolution 2024; 41(12), msae263.
R Pipke and N Amrhein. Isolation and characterization of a mutant of Arthrobacter sp. strain GLP-1 which utilizes the herbicide glyphosate as its sole source of phosphorus and nitrogen. Applied and Environmental Microbiology 1988; 54(11), 2868-2870.
N Stosiek, M Talma and M Klimek-Ochab. Carbon-phosphorus Lyase—the state of the art. Applied Biochemistry and Biotechnology 2020; 190, 1525-1552.
S Firdous, S Iqbal and S Anwar. Optimization and modeling of glyphosate biodegradation by a novel Comamonas odontotermitis P2 through response surface methodology. Pedosphere 2020; 30(5), 618-627.
WJ Chen, M Liu, SF Chen, Y Zhang, H Song, MH Abdoulahi, K Bhatt, S Mishra, MA Ghorab, W Zhang and S Chen. Glyphosate bioremediation using a newly isolated Bacillus albus strain F9D: Mechanisms and kinetic studies. Microbial Cell Factories 2025; 24, 195.
J Fan, G Yang, H Zhao, G Shi, Y Geng, T Hou and K Tao. Isolation, identification and characterization of a glyphosate-degrading bacterium, Bacillus cereus CB4, from soil. The Journal of General and Applied Microbiology 2012; 58(4), 263-271.
CL Patten, AJC Blakney and TJD Coulson. Activity, distribution and function of indole-3-acetic acid biosynthetic pathways in bacteria. Critical Reviews in Microbiology 2013; 39(4), 395-415.
M Shahid, B Ahmed and MS Khan. Evaluation of microbiological management strategy of herbicide toxicity to greengram plants. Biocatalysis and Agricultural Biotechnology 2018; 14, 96-108.
R Hertel, K Schöne, C Mittelstädt, J Meißner, N Zschoche, M Collignon, C Kohler, I Friedrich, D Schneider, M Hoppert, R Kuhn, I Schwedt, P Scholz, A Poehlein, M Martienssen, T Ischebeck, R Daniel and FM Commichau. Characterization of glyphosate‐resistant Burkholderia anthina and Burkholderia cenocepacia isolates from a commercial Roundup® solution. Environmental Microbiology Reports 2022; 14(1), 70-84.
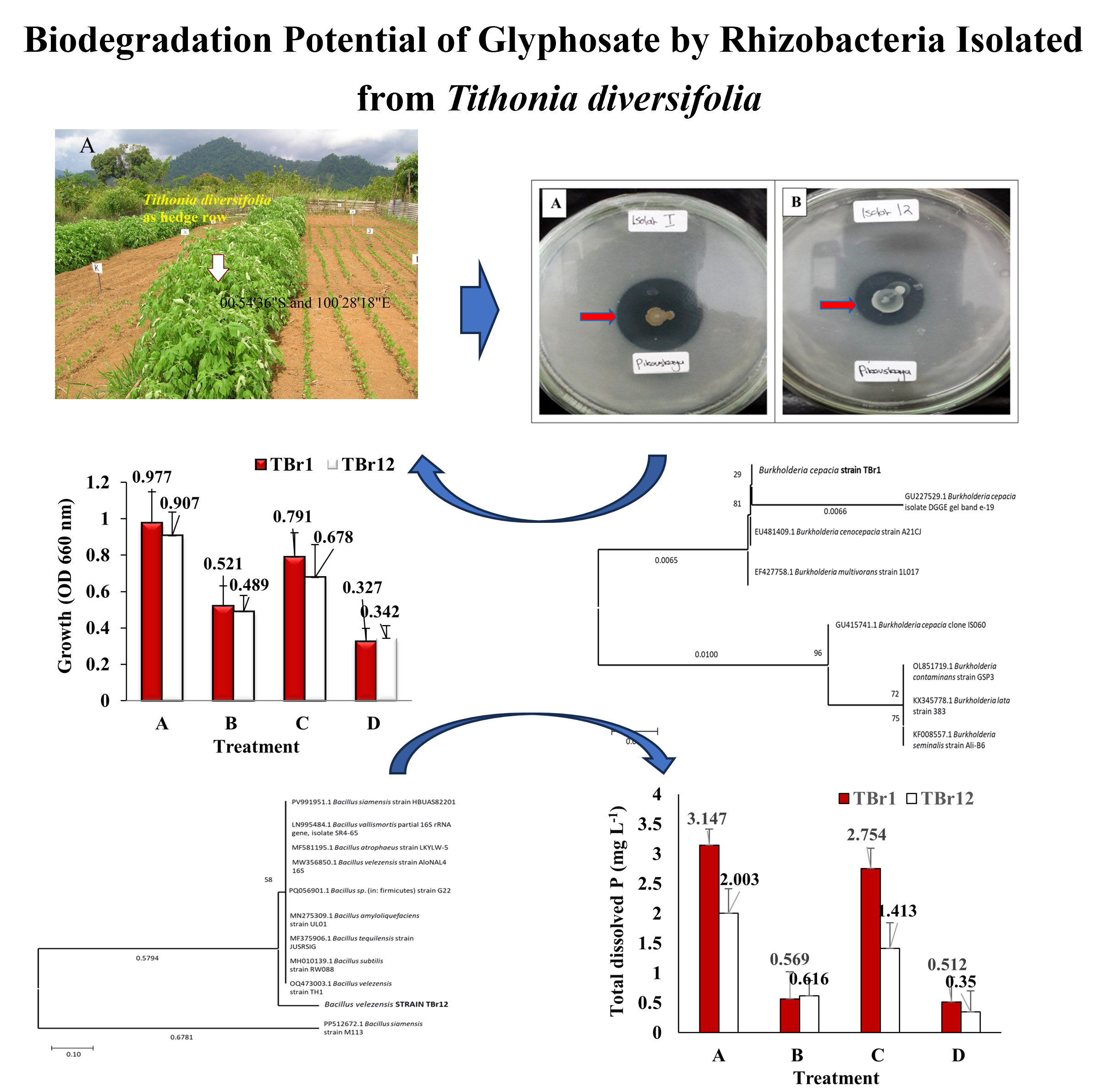
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






