LC–HRMS Metabolomics Fingerprints and Pathway Signatures Authenticate Geographic Origin of Pempek (Channa striata)
DOI:
https://doi.org/10.48048/tis.2026.12122Keywords:
LC-HRMS metabolomics, Chemometric modeling, Pempek authentication, Channa striata, Food authenticity, geographic origin, Biomarker discovery, LC–HRMS metabolomics, Chemometric modeling, Pempek authentication, Channa striata, Food authenticity, Geographic origin, Biomarker discoveryAbstract
This study demonstrates that non‑targeted liquid chromatography–high‑resolution mass spectrometry (LC–HRMS) metabolomics provides a robust framework for authenticating the geographic origin of pempek, a traditional fish‑based food prepared from Channa striata. The work addresses the limited application of untargeted LC–HRMS to processed freshwater fish products across multiple regions by explicitly testing whether molecular fingerprints capture ecological and artisanal differences between Jambi and South Sumatra. Samples were processed under standardized protocols and analyzed with LC–HRMS, followed by variance stabilization and chemometric modeling in MetaboAnalyst (a web-based platform for metabolomics data analysis). Principal component analysis (PCA) revealed origin‑based clustering, and orthogonal projections to latent structures–discriminant analysis (OPLS-DA), supported by extensive permutation validation, confirmed significant separation. Discriminatory metabolites included carnitines, choline, and creatinine enriched in Jambi products, contrasted with ether‑linked phosphatidylcholines and sphingomyelins enriched in South Sumatra products. Pathway enrichment analyses linked these differences to membrane lipid biosynthesis and fatty‑acid beta‑oxidation, while receiver operating characteristic (ROC) curves based on a multi‑marker panel demonstrated near‑perfect discrimination. These findings indicate that both endogenous metabolic traits and exogenous processing signatures jointly shape pempek metabolomes. The study advances food authenticity science by moving beyond descriptive profiling toward mechanistic interpretation and translational application. By establishing evidence‑based fingerprints for pempek, it provides a scientific foundation for protecting the cultural and economic value of this traditional Indonesian food. Future directions should include cross‑season and interlaboratory validation, targeted assays with authentic standards, and expanded regional comparisons to strengthen regulatory translation and consumer trust.
HIGHLIGHTS
- LC–HRMS fingerprints distinguish pempek origins (Jambi vs. South Sumatra)
- Chemometric modeling (PCA, OPLS-DA, ROC) ensures robust authentication.
- Key markers: carnitine, choline (Jambi) vs. phosphatidylcholines (South Sumatra)
- Pathway enrichment links origin to lipid and energy metabolism differences
- Validated marker panel enables accurate and culturally significant authentication
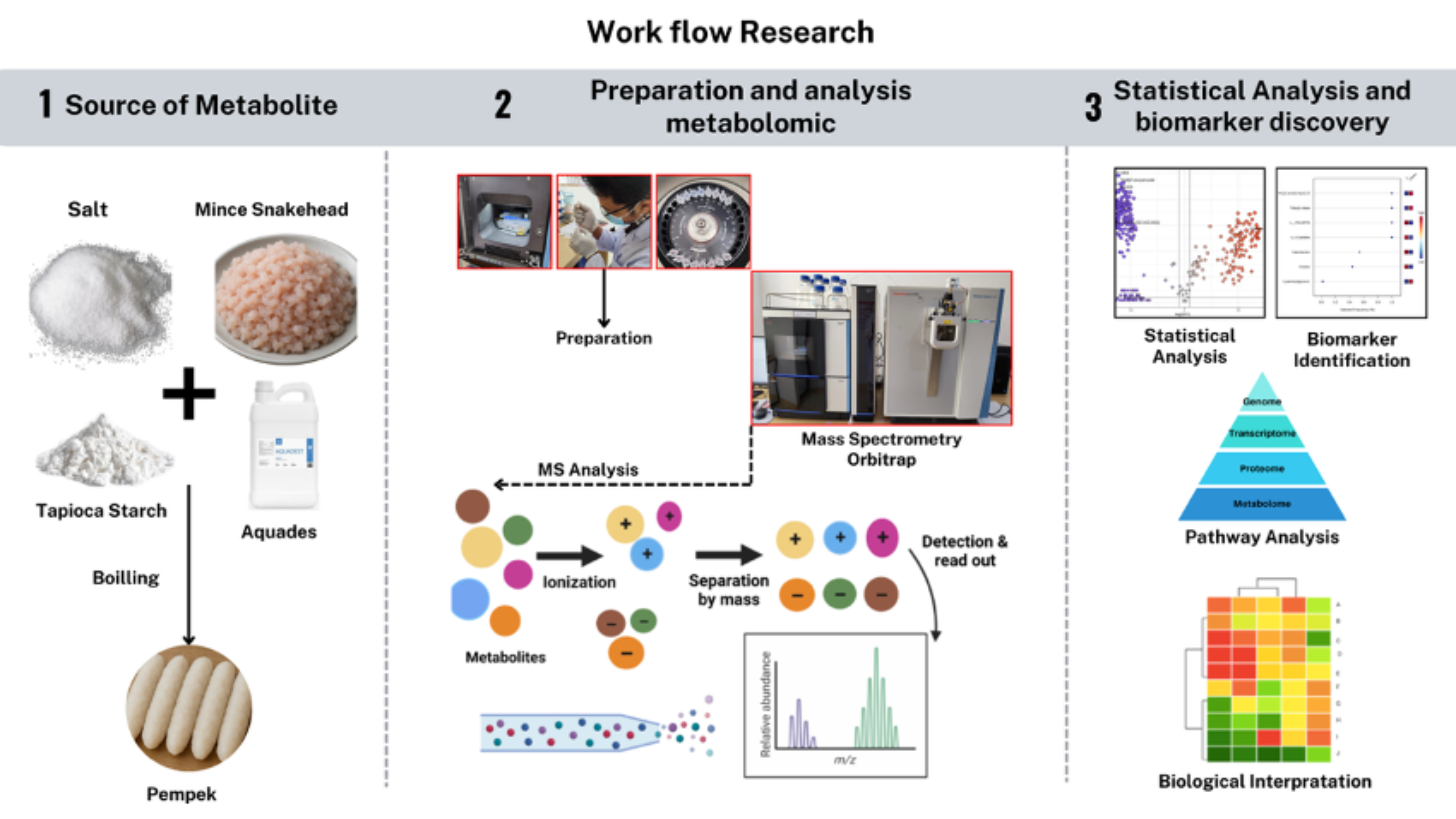
GRAPHICAL ABSTRACT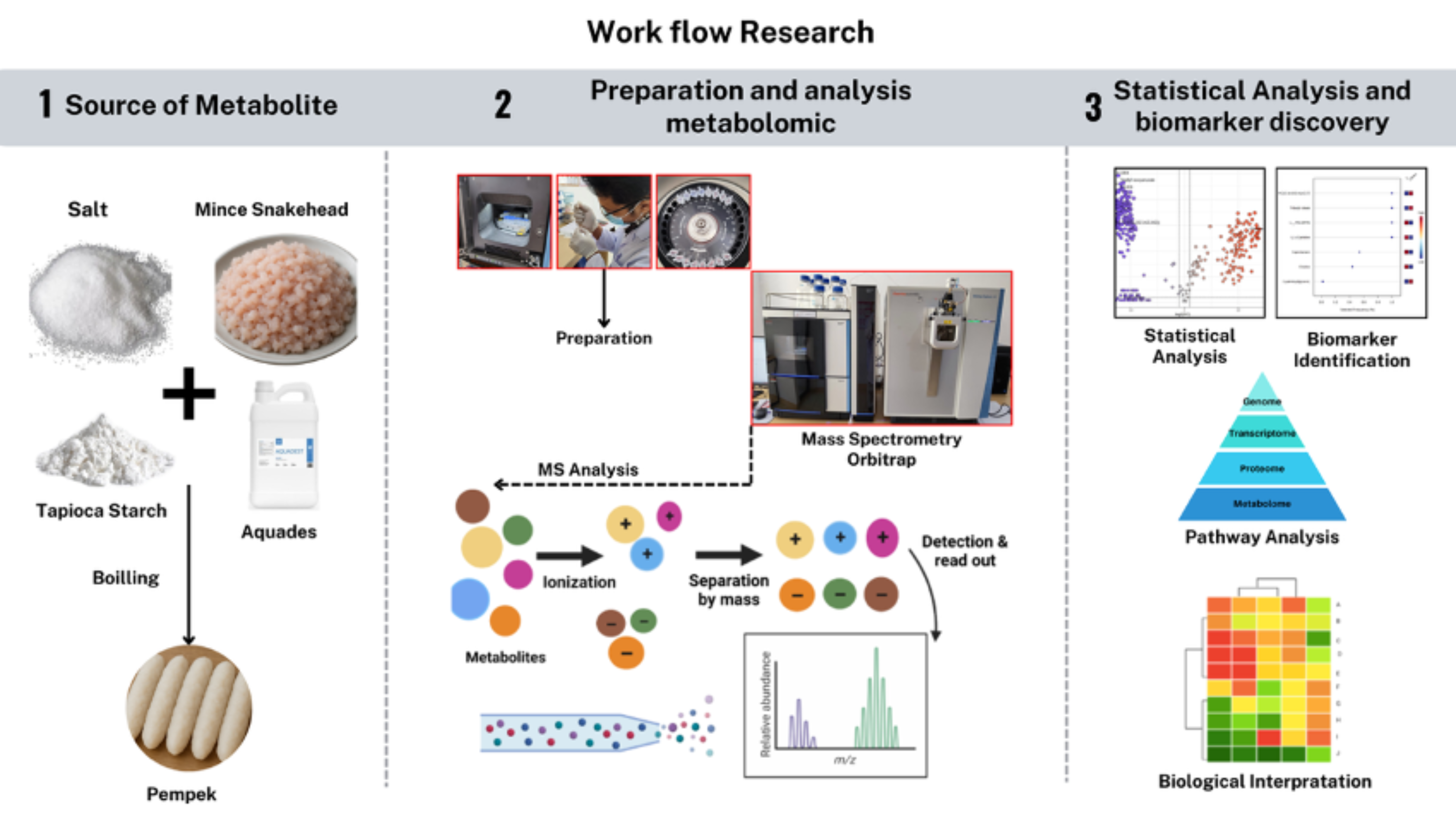
Downloads
References
X Ma, D Su, J Zhu, J Miao, X Liu, K Leng, X Wang and W Xie. Comparative metabolomics study on the quality of Antarctic krill (Euphausia superba) stored at different temperatures. International Journal of Food Science & Technology 2024; 59(7), 4489-4499.
X Wang, Y Li, L Chen and J Zhou. Analytical strategies for LC-MS-Based untargeted and targeted metabolomics approaches reveal the entomological origins of honey. Journal of Agricultural and Food Chemistry 2022; 70(4), 1358-1366.
S Hemmer, SK Manier, S Fischmann, F Westphal, L Wagmann and MR Meyer. Comparison of three untargeted data processing workflows for evaluating LC-HRMS metabolomics data. Metabolites 2020; 10(9), 378.
CK Asef, MA Rainey, BM. Garcia, GJ Gouveia, AO Shaver, FE Leach, AM Morse, AS Edison, LM McIntyre and FM Fernández. Unknown metabolite identification using machine learning collision cross-section prediction and tandem mass spectrometry. Analytical Chemistry 2023; 95(2), 1047-1056.
JM Malinowska, T Palosaari, J Sund, D Carpi, GR Lloyd, RJM Weber, M Whelan amd MR Viant. Automated sample preparation and data collection workflow for high-throughput in vitro metabolomics. Metabolites 2022; 12(1), 52.
H Liang and K Song. Comprehensive metabolomics and transcriptomics analysis reveals protein and amino acid metabolic characteristics in liver tissue under chronic hypoxia. PLoS One 2023; 18(9), e0291798.
DG Wu, CR Harris, KM Kalis, M Bowen, JF Biddle and IF Farag. Comparative metagenomics of tropical reef fishes show conserved core gut functions across hosts and diets with diet-related functional gene enrichments. Applied and Environmental Microbiology 2025; 91(2), e0222924.
N Zhou, T Gu, M Duan, Y Tian, L Chen, T Zeng, X Hou, X Wang, Q Xu, Y Zhang and L Lu. Gut microbiota dysbiosis exacerbates polystyrene microplastics-induced liver inflammation via activating LPS/TLR4 signaling pathway in ducks. Poultry Science 2025; 104(11), 105757.
L Malm, E Palm, A Souihi, M Plassmann, J Liigand and A. Kruve. Guide to semi-quantitative non-targeted screening using LC/ESI/HRMS. Molecules 2021; 26(12), 3524.
S Li, A Siddiqa, M Thapa, Y Chi and S Zheng. Trackable and scalable LC-MS metabolomics data processing using asari. Nature Communications 2023; 14(1), 4113.
J Zhang, M Sun, AH Elmaidomy, KA Youssif, AMM Zaki, HH Kamal, AM Sayed and UR Abdelmohsen. Emerging trends and applications of metabolomics in food science and nutrition. Food & Function 2023; 14(20), 9050-9082.
D Dörfel, S Rohn and E Jantzen. Electrostatic repulsion hydrophilic interaction liquid chromatography (ERLIC) for the quantitative analysis of polyamines. Journal of Chromatography 2024; 1720, 464820.
RE Danczak, RK Chu, SJ Fansler, AE Goldman, EB Graham, MM Tfaily, J Toyoda and JC Stegen. Using metacommunity ecology to understand en¬vironmental metabolomes. Nature Communica¬tions 2020; 11(1), 6369.
Z Pang, J Chong, S Li and J Xia. Metaboanalystr 3.0: Toward an optimized workflow for global metabolomics. Metabolites 2020; 10(5), 186.
A Supriadi, S Ridhowati, D Saputra and SD Lestari. Untargeted metabolomics profiling for the geographical authentication of traditional pempek using high-resolution orbitrap mass spectrometry. Food Chemistry Advances 2025; 6, 100914.
A Windarsih, Suratno, HD Warmiko, AW Indrianingsih, A Rohman and YI Ulumuddin. Untargeted metabolomics and proteomics approach using liquid chromatography-Orbitrap high resolution mass spectrometry to detect pork adulteration in Pangasius hypopthalmus meat. Food Chem 2022; 386, 132856.
A Supriadi, S Ridhowati, D Saputra, Wulandari and SD Lestari. Untargeted metabolomics profiling for the geographical authentication of traditional pempek using high-resolution orbitrap mass spectrometry. Food Chemistry Advances 2025; 6, 100914.
J Kusuma, Analianasari, A Wahyudi, MK Abdullah, AZ Hasan, I Asrowardi, Fitriani and M Tahir. Diversity of the non-targeted metabolomic data across various varieties of Cloves (Syzygium spp.). Data in Brief 2024; 58, 111237.
MT Fernández-del-Campo-García, AM Casas-Ferreira, E Rodríguez-Gonzalo and JLP Pavón. Combining Orbitrap-HRMS acquisition modes and direct injection by a guard column for targeted analysis of underivatized amino acids in urine. Microchemical Journal 2024; 196, 109663,
A Windarsih, Suratno, HD Warmiko, AW Indrianingsih, A Rohman and YI Ulumuddin. Untargeted metabolomics and proteomics approach using liquid chromatography-Orbitrap high resolution mass spectrometry to detect pork adulteration in Pangasius hypopthalmus meat. Food Chemistry 2022; 386, 132856.
CH Yong, SA Muhammad, FA Aziz, FI Nasir, MZ Mustafa, B Ibrahim, SD Kelly, A Cannavan and EK Seow. Detecting adulteration of stingless bee honey using untargeted 1H NMR metabolomics with chemometrics. Food Chemistry 2022; 368, 130808.
Y Pan, Hui-Wen Gu, Y Lv, XL Yin, Y Chen, W Long, H Fu and Y She. Untargeted metabolomic analysis of Chinese red wines for geographical origin traceability by UPLC-QTOF-MS coupled with chemometrics. Food Chemistry 2022; 394, 133473.
A Akbar AJr Siddiqui, A Raza, A Zia, K Nakyinsige, K Hakiimu and SG Musharraf. Untargeted-metabolomics differentiation of unripened cow milk cheese produced from different sources of rennet. Food Control 2025; 171, 111113.
G Riquelme, N Zabalegui, P Marchi, CM Jones and ME Monge. A python-based pipeline for preprocessing LC–MS data for untargeted metabolomics workflows. Metabolites 2020; 10(10), 416.
JD Mosley, TB Schock, CW Beecher, WB Dunn, J Kuligowski, MR Lewis, G Theodoridis, CZU Holland, D Vuckovic, ID Wilson and KA Zanetti. Establishing a framework for best practices for quality assurance and quality control in untargeted metabolomics. Metabolomics 2024; 20(2), 20.
X Shen, R Wang, X Xiong, Y Yin, Y Cai, Z Ma, N Liu and ZJ Zhu. Metabolic reaction network-based recursive metabolite annotation for untargeted metabolomics. Nature Communications 2019; 10(1), 1516.
P Bonini, T Kind, H Tsugawa, DK Barupal and O Fiehn. Retip: Retention time prediction for compound annotation in untargeted metabolomics. Analytical chemistry 2020; 92(11), 7515-7522.
CM Vitale, A Lommen, C Huber, K Wagner, BG Molina, R Nijssen, EJ Price, M Blokland, F Tricht, HGJ Mol, M Krauss, L Debrauwer, O Pardo, N Leon, J Klanova and JP Antignac. Harmonized quality assurance/quality control provisions for nontargeted measurement of urinary pesticide biomarkers in the HBM4EU multisite SPECIMEn study. Analytical Chemistry 2022, 94(22), 7833-7843.
H Villalba, M Llambrich, J Gumá, J Brezmes and R Cumeras. A metabolites merging strategy (MMS): Harmonization to enable studies’ intercomparison. Metabolites 2023; 13(12), 1167.
S Hemmer, SK Manier, S Fischmann, F Westphal, L Wagmann and MR Meyer. Comparison of three untargeted data processing workflows for evaluating LC-HRMS metabolomics data. Metabolites 2020; 10(9), 378.
R Gil‐Solsona, J Nácher-Mestre, L Lacalle-Bergeron, JV Sancho, JA Calduch-Giner, F Hernández and J Pérez-Sánchez. Untargeted metabolomics approach for unraveling robust biomarkers of nutritional status in fasted gilthead sea bream (Sparus Aurata). PeerJ 2017; 5, e2920.
M Schwaiger-Haber, H Schoeny, YE Abiead, G Hermann, E Rampler and G Koellensperger. Merging metabolomics and lipidomics into one analytical run. Analyst 2019; 144(1), 220-229
KN Tran, HG Sutherland, A Mallett, LR Griffiths and RA Lea. New composite phenotypes enhance chronic kidney disease classification and genetic associations. PLoS Genetics 2025; 21(5), e1011718
Y Xu, L Cao, Y Chen, Z Zhang, W Liu, H Li, C Ding, J Pu, K Qian and W Xu. integrating machine learning in metabolomics: A path to enhanced diagnostics and data interpretation. Small Methods 2024; 8(12), e2400305.
N Sousa, MJ Moreira, C Saraiva and JMMMD Almeida. Applying fourier transform mid infrared spectroscopy to detect the adulteration of salmo salar with oncorhynchus mykiss. Foods 2018; 7(4), 55.
M Barranco-Altirriba, P Solà-Santos, S Picart-Armada, S Kanaan-Izquierdo, J Fonollosa and A Perera-Lluna. MWISE: An Algorithm for Context-Based Annotation of Liquid Chromatography-Mass Spectrometry Features through Diffusion in Graphs. Analytical Chemistry 2021; 93(31), 10772-10778
I Blaženović, T Kind, J Ji and O Fiehn. Software tools and approaches for compound identification of LC-MS/MS data in metabolomics. Metabolites 2018; 8(2), 31.
JA Etzel and TS Braver. MVPA permutation schemes: Permutation testing in the land of cross-validation. In Proceedings - 2013 3rd International Workshop on Pattern Recognition in Neuroimaging, Philadelphia, United States. 2013, pp. 140-143.
D Schütz, E Achten, M Creydt, J Riedl and M Fischer. Non-targeted LC-MS metabolomics approach towards an authentication of the geographical origin of grain maize (Zea mays L.) samples. Foods 2021; 10(9), 2160.
A Maléchaux, YL Dréau, J Artaud and N Dupuy. Control chart and data fusion for varietal origin discrimination: Application to olive oil. Talanta 2020; 217, 121115.
X Wang, C Li, Z Li, Y Qi, X Zhang, X Zhao, C Zhao, X Lin, X Lu and G Xu. A structure-guided molecular network strategy for global untargeted metabolomics data annotation. Analytical Chemistry2023; 95(31), 11603-11612.
TN Bayissa, M Geerardyn, D Vanhauteghem, M Wakjira and GPJ Janssens. Nutrient-related metabolite profiles explain differences in body composition and size in Nile tilapia (Oreochromis niloticus) from different lakes. Scientific Reports 2021; 11(1), 16824
M Schwaiger, H Schoeny, YE Abiead, G Hermann, E Rampler and G Koellensperger. Merging metabolomics and lipidomics into one analytical run. Analyst 2018; 144(1), 220-229.
S Nishiumi Y Izumi, A Hirayama, M Takahashi, M Nakao, K Hata, D Saigusa, E Hishinuma, N Matsukawa, SM Tokuoka, Y Kita, F Hamano, N Okahashi, K Ikeda, H Nakanishi, K Saito, MY Hirai, M Yoshida, Y Oda, F Matsuda and T Bamba. Comparative evaluation of plasma metabolomic data from multiple laboratories. Metabolites 2022; 12(2), 135.
YC Sun, SC Han, MZ Yao, YM Wang, LW Geng, P Wang, WH Lu and HB Liu. High-throughput metabolomics method based on liquid chromatography-mass spectrometry: Insights into the underlying mechanisms of salinity–alkalinity exposure-induced metabolites changes in Barbus capito. Journal of Separation Science 2021; 44(2), 497-512.
X Liang, Q He, G Qin, G Li, Q Li, H Tan, Z Wang, M Fan and D Xu. Effectively removing the homodimer in bispecific antibodies by weak partitioning mode of anion exchange chromatography. Journal of Chromatography B 2023; 1225, 123767,
GJ Gouveia, AO Shaver, BM Garcia, AM Morse, EC Andersen, AS Edison and LM McIntyre. Long-term metabolomics reference material. Analytical Chemistry 2021; 93(26), 9193-9199.
C Claassen, J Kuballa and S Rohn. Polar lipids in starch-rich commodities to be analyzed with LC-MS-Based metabolomics—optimization of ionization parameters and high-throughput extraction protocols. Metabolites 2019; 9(8), 167.
J Ivanišević and EJ Want. From samples to insights into metabolism: Uncovering biologically relevant information in LC-HRMS metabolomics data. Metabolites 2019; 9(12), 308.
R González-Domínguez, A Sayago and Á Fernández-Recamales. An overview on the application of chemometrics tools in food authenticity and traceability. Foods 2022; 11(23), 3940.
M Bongaerts R Bonte, S Demirdas, EH Jacobs, E Oussoren, AT Ploeg, MAEM Wagenmakers, RMW Hofstra, HJ Blom, MJT Reinders and GJG Ruijter. Using out-of-batch reference populations to improve untargeted metabolomics for screening inborn errors of metabolism. Metabolites 2020; 11(1), 8.
S Zheng. Safeguarding food heritage through social media? between heritagization and commercialization. International Journal of Gastronomy and Food Science 2023; 31, 100678.
A Scalbert, L Brennan, O Fiehn, T Hankemeier, BS Kristal, B Ommen, E Pujos-Guillot, E Verheij, D Wishart and S Wopereis. Mass-spectrometry-based metabolomics: Limitations and recommendations for future progress with particular focus on nutrition research. Metabolomics 2009; 5(4), 435-458.
C Lavarello, S Barco, M Bartolucci, I Panfoli, E Magi, G Tripodi, A Petretto and G Cangemi. Development of an accurate mass retention time database for untargeted metabolomic analysis and its application to plasma and urine pediatric samples. Molecules 2021; 26(14), 4256.
YP Tsentalovich, EA Zelentsova, LV Yanshole, VV Yanshole and IM Odud. Most abundant metabolites in tissues of freshwater fish pike-perch (Sander lucioperca). Scientific Reports 2020; 10, 17128.
G Pimentel, D Burnand, LH Münger, FP Pralong, N Vionnet, R Portmann and G Vergères. Identification of milk and cheese intake biomarkers in healthy adults reveals high interindividual variability of lewis system–Related oligosaccharides. Journal of Nutrition 2020; 150(5), 1058-1067,
M Kortesniemi, S Noerman, A Kårlund, J Raita, T Meuronen, V Koistinen, R Landberg and K Hanhineva. Nutritional metabolomics: Recent developments and future needs. Current Opinion in Chemical Biology 2023; 77, 102400.
M Vinaixa, EL Schymanski, S Neumann, M Navarro, RM Salek and O Yanes. Mass spectral databases for LC/MS- and GC/MS-based metabolomics: State of the field and future prospects. TrAC - Trends in Analytical Chemistry 2016; 78, 23-35.
RM Boiteau, DW Hoyt, CD Nicora, HA Kinmonth-Schultz, JK Ward and K Bingol. Structure elucidation of unknown metabolites in metabolomics by combined NMR and MS/MS prediction. Metabolites 2018; 8(1), 8.
YP Tsentalovich, EA Zelentsova, ED Savina, VV Yanshole and RZ Sagdeev. Influence of ecological factors on the metabolomic composition of fish lenses. Biology 2022; 11(12), 1709.
Y Peng, C Zheng, S Guo, F Gao, X Wang, Z Du, F Gao, F Su, W Zhang, X Yu, G Liu, B Liu, C Wu, Y Sun, Z Yang, Z Hao and X Yu. Metabolomics integrated with machine learning to discriminate the geographic origin of Rougui Wuyi rock tea. NPJ Science of Food 2023; 7, 7.
B Zhang, S Hu, E Baskin, A Patt, JK Siddiqui and EA Mathé. RaMP: A comprehensive relational database of metabolomics pathways for pathway enrichment analysis of genes and metabolites. Metabolites 2018; 8(1), 16.
P Bonini, T Kind, H Tsugawa, DK Barupal and O Fiehn. Retip: Retention time prediction for compound annotation in untargeted metabolomics. Analytical Chemistry 2020; 92(11), 7515-7522.
L Cottret, D Wildridge, F Vinson, MP Barrett, H Charles, MF Sagot and F Jourdan. MetExplore: A web server to link metabolomic experiments and genome-scale metabolic networks. Nucleic Acids Research 2010; 38(S2), W132-W137.
M Bayerlová, K Jung, F Kramer, F Klemm, A Bleckmann and T Beißbarth. Comparative study on gene set and pathway topology-based enrichment methods. BMC Bioinformatics 2015; 16(1), 344.
S Klockmann, E Reiner, N Cain and M Fischer. Food targeting: Geographical origin determination of hazelnuts (corylus avellana) by LC-QqQ-MS/MS-Based targeted metabolomics application. Journal of Agricultural and Food Chemistry 2017; 65(7), 1456-1466.
B Musio, R Ragone, S Todisco, A Rizzuti, E Iorio, M Chirico, ME Pisanu, N Meloni, P Mastrorilli and V Gallo. Non-targeted nuclear magnetic resonance analysis for food authenticity: A comparative study on tomato samples. Molecules 2024; 29(18), 4441.
T Martinović, M Šrajer Gajdošik and D Josić. Sample preparation in foodomic analyses. Electrophoresis 2018; 39(13), 1527-1542.
AM Teixeira, JMGD Queiroz, BC Garrido, AJR Silva, A Bauermeister and RM Borges. Quality as¬surance in untargeted metabolomics. Quim Nova 2025; 48(3), e-20250048.
G Riquelme, EE Bortolotto, M Dombald and M Eugenia. Model-driven data curation pipeline for LC–MS-based untargeted metabolomics. Metabolomics 2023; 19(3), 15.
M Lenski, S Maallem, G Zarcone, G Garçon, JM Lo-Guidice, S Anthérieu and D Allorge. Prediction of a Large-Scale database of collision cross-section and retention time using machine learning to reduce false positive annotations in untargeted metabolomics. Metabolites 2023; 13(2), 282.
E Sinclair, KA Hollywood, C Yan, R Blankley, R Breitling and P Barran. Mobilising ion mobility mass spectrometry for metabolomics. Analyst 2018; 143(19), 4783-4788.
VM Koistinen, AB Silva, L Abrankó, D Low, RG Villalba, FT Barberán, R Landberg, O Savolainen, I Alvarez-Acero, SD Pascual-Teresa, CV Poucke, C Almeida, L Petrásková, K Valentová, S Durand, W Wiczkowski, D Szawara-Nowak, R González-Domínguez, R Llorach, C Andrés-Lacueva, … MR Bronze. Interlaboratory coverage test on plant food bioactive compounds and their metabolites by mass spectrometry-based untargeted metabolomics. Metabolites 2018; 8(3), 46.
ZA Quinlan, I Koester, AT Aron, D Petras, LI Aluwihare, PC Dorrestein, CE Nelson and LW Kelly. ConCISE: Consensus annotation propagation of ion features in untargeted tandem mass spectrometry combining molecular networking and In Silico metabolite structure prediction. Metabolites 2022; 12(12), 1275.
Y Cui, D Pan, Ji Feng, D Zhao, M Liu, Z Dong, S Liu and S Wang. Untargeted metabolomics and soil community metagenomics analyses combined with machine learning evaluation uncover geographic differences in ginseng from different locations. Journal of Agricultural and Food Chemistry 2024; 72(39), 21922-21934.
Y Wang, RA Hodge, VL Stevens, TJ Hartman and ML McCullough. Identification and reproducibility of urinary metabolomic biomarkers of habitual food intake in a cross-sectional analysis of the cancer prevention study-3 diet assessment sub-study. Metabolites 2021; 11(4), 248.
M Bongaerts, R Bonte, S Demirdas, EH Jacobs, E Oussoren, AT Ploeg, MAEM Wagenmakers, RMW Hofstra, HJ Blom, MJT Reinders and GJG Ruijter. Using out-of-batch reference populations to improve untargeted metabolomics for screening inborn errors of metabolism. Metabolites 2020; 11(1), 8.
X Wu, W Huang, K Wang, Y Han and Z Liu. Visual bibliometric analysis of the research status and hotspots of myopia in children and adolescents in the past five years. Chinese Journal of Optometry Ophthalmology and Visual Science 2024; 26(9), 677-684.
G Theodoridis, A Pechlivanis, NS Thomaidis, A Spyros, CA Georgiou, T Albanis, I Skoufos, S Kalogiannis, GT Tsangaris, AS Stasinakis, I Konstantinou, A Triantafyllidis, K Gkagkavouzis, AS Kritikou, ME Dasenaki, H Gika, C Virgiliou, D Kodra, N Nenadis, I Sampsonidis, … OBO The FoodOmicsGR Ri Consortium. Foodomicsgr_ri. a consortium for comprehensive molecular characterisation of food products. Metabolites 2021; 11(2), 74.
J Wang, Y Xue, P Wang, J Chen and L Yao. Participation mode and production efficiency enhancement mechanism of Geographical Indication products in rural areas: A meta-frontier analysis. Physics and Chemistry of the Earth 2021; 121, 102982.
A Kritikos, I Aska, S Ekonomou, A Mallouchos, FF Parlapani, SA Haroutounian and IS Boziaris. Volatilome of chill-stored european seabass (Dicentrarchus labrax) fillets and atlantic salmon (salmo salar) slices under modified atmosphere packaging. Molecules 2020; 25(8), 1981.
T Cajka. Liquid chromatography–mass spectrometry–based metabolomics approaches for foodomics research. Current Opinion in Food Science 2024; 58, 101201.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






