Histone Methyltransferase and Demethylase Gene in Pediatric Acute Lymphoblastic Leukemia: A Molecular Insights
DOI:
https://doi.org/10.48048/tis.2026.11952Keywords:
Epigenetic regulation, Gene mutation, Histone demethylase, Histone methyltransferase, Pediatric ALLAbstract
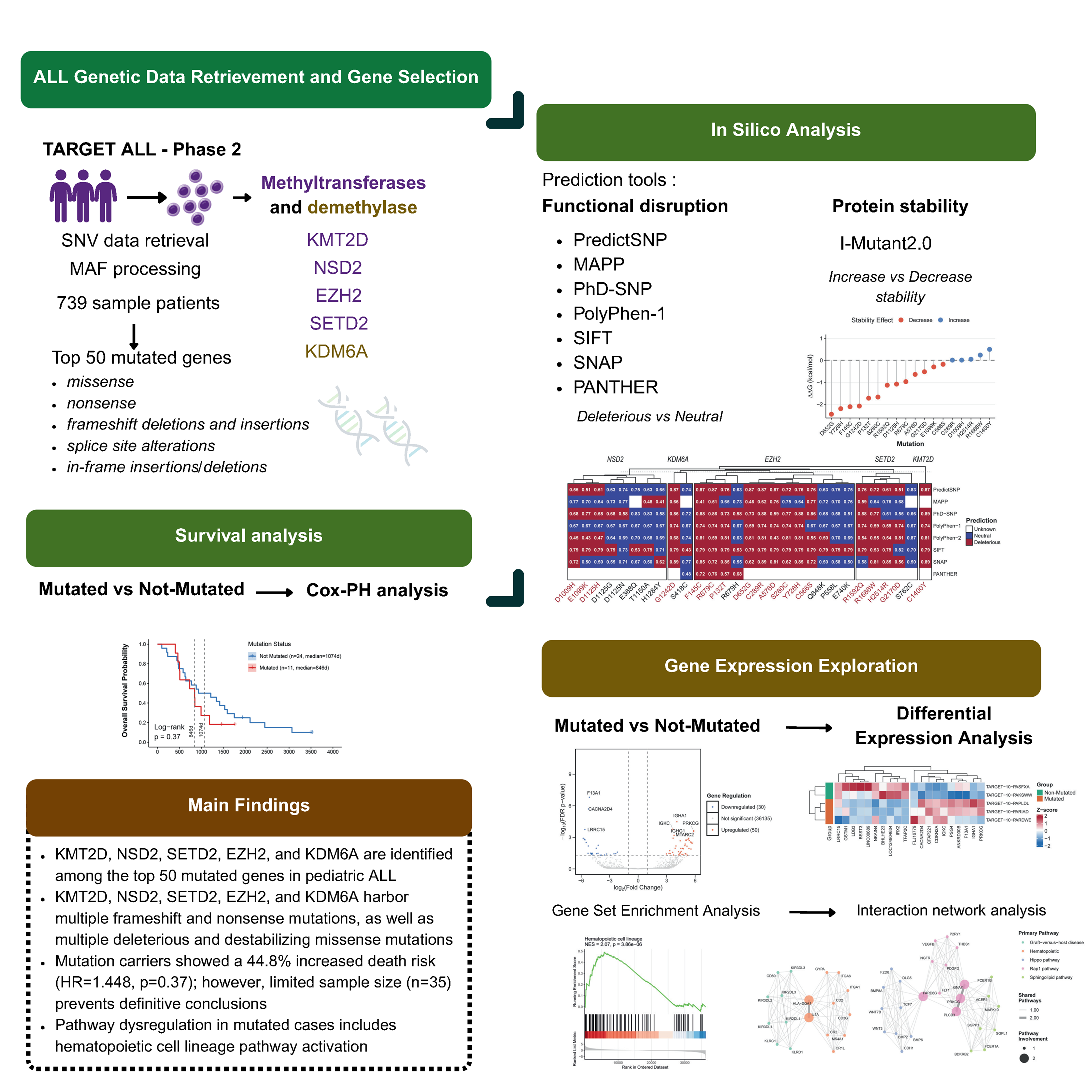
Genetic factors driving the development of pediatric Acute Lymphoblastic Leukemia (ALL) remain incompletely understood. While epigenetic dysregulation through histone methyltransferases and demethylases has emerged as a critical oncogenic mechanism, their specific contributions to pediatric ALL require systematic investigation. In this study, we systematically investigated genetic variants in histone methyltransferases and demethylase found in top 50 mutated genes from TARGET ALL Phase 2 cohort. Single nucleotide variations were functionally annotated using PredictSNP and I-Mutant2.0, followed by survival analysis and pathway enrichment studies comparing mutated versus wild-type patients. KMT2D, NSD2, SETD2, EZH2 (methyltransferases), and KDM6A (demethylase), ranked among the most frequently mutated genes. Multiple frameshift and nonsense mutations were identified in these genes, likely resulting in truncated proteins and loss of function. Critically, missense variants in EZH2 (F145C, R679C, P132T, D652G, A576D, S280C, Y728H, C566S), KDM6A (G1242D), NSD2 (E1099K, D1125H), and SETD2 (G2170D, R1592Q) were computationally predicted as deleterious and destabilizing. Although survival analysis revealed no statistically significant differences (p = 0.37), mutation carriers exhibited a 44% increased mortality risk, suggesting clinical relevance despite limited sample size (n = 35). Gene set enrichment analysis of differentially expressed genes revealed significant activation of hematopoietic cell lineage pathways in mutated patients. Our findings underscore the importance of histone methyltransferase and demethylase gene variants in pediatric ALL, particularly novel mutations affecting protein function and stability that remain poorly characterized in the literature. This study identifies candidate mutations that warrant further functional and clinical investigation for their role in disease progression and therapeutic targeting.
HIGHLIGHTS
- KMT2D, NSD2, SETD2, EZH2, and KDM6A are identified among the top 50 most frequently mutated genes in pediatric ALL
- KMT2D exhibits the highest mutational burden in pediatric ALL, predominantly harboring frameshift and nonsense alterations
- KMT2D, NSD2, SETD2, EZH2, and KDM6A genes show consensus deleterious predictions across multiple in silico tools, with 18 mutations representing high-priority candidates for experimental validation
- Structural stability analysis reveals critical destabilizing mutations in EZH2 (F145C, R679C, P132T, D652G, A576D, S280C, Y728H, C566S), KDM6A (G1242D), NSD2 (E1099K, D1125H), and SETD2 (G2170D, R1592Q)
GRAPHICAL ABSTRACT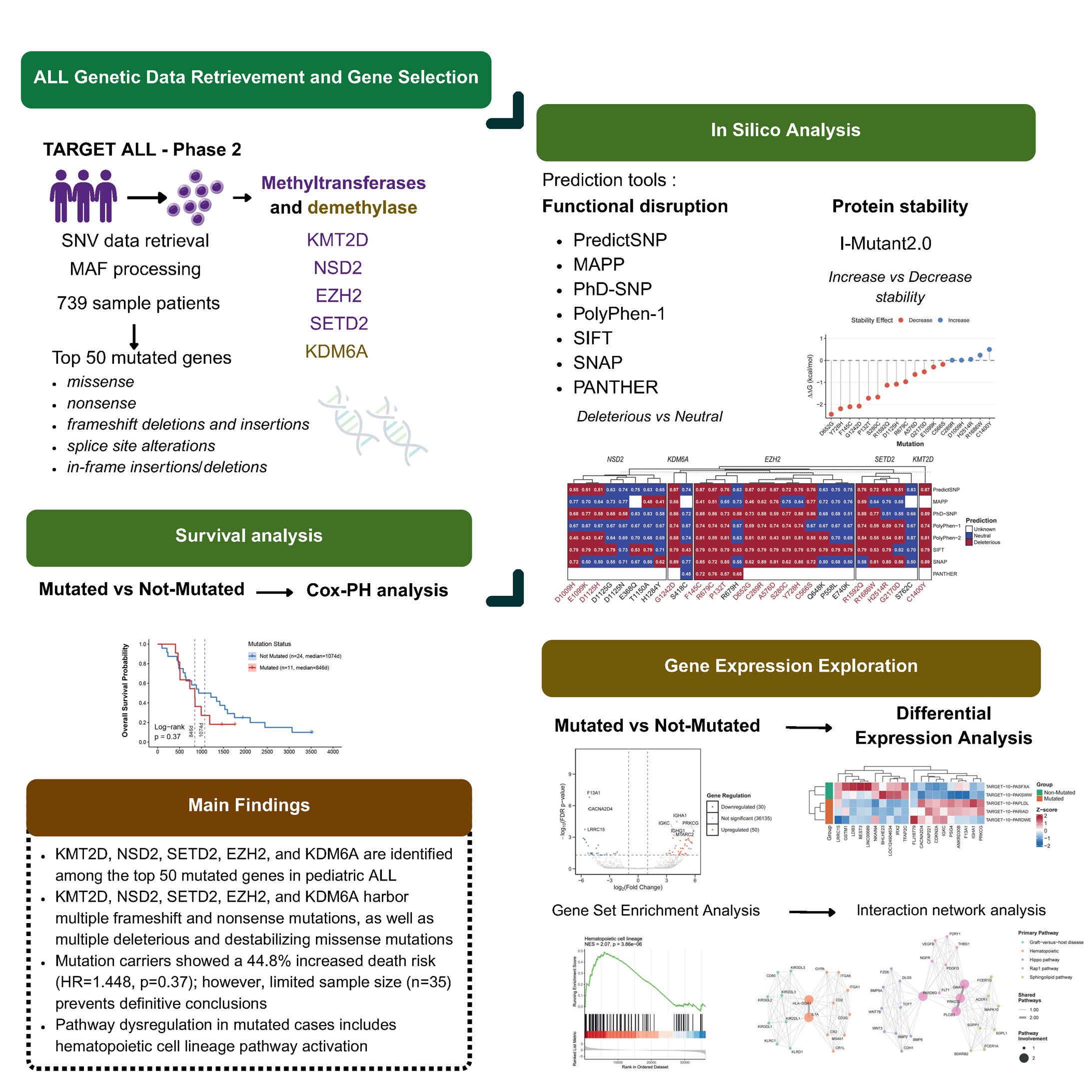
Downloads
References
H Inaba, D Teachey, C Annesley, S Batra, J Beck, S Colace, S Cooper, M Dallas, SD Oliveira, K Kelly, C Kitko, M Kohorst, M Kutny, N Lacayo, C Lee-Miller, K Ludwig, L Madden, K Maloney, D Mangum, …, K Stehman. Pediatric acute lymphoblastic leukemia, version 2.2025, NCCN clinical practice guidelines in oncology. Journal of the National Comprehensive Cancer Network 2025; 23(2), 41-62.
A Mohammadian-Hafshejani, IM Farber and S Kheiri. Global incidence and mortality of childhood leukemia and its relationship with the Human Development Index. PLoS One 2024; 19(7), e0304354.
MS Lajevardi, M Ashrafpour, SMH Mubarak, B Rafieyan, A Kiani, E Noori, M Roayaei Ardakani, M Montazeri, N Kouhi Esfahani, N Asadimanesh, S Khalili and Z Payandeh. Dual roles of extracellular vesicles in acute lymphoblastic leukemia: Implications for disease progression and theranostic strategies. Medical Oncology 2024; 42(1), 11.
H Kantarjian and E Jabbour. Adult acute lymphoblastic leukemia: 2025 update on diagnosis, therapy, and monitoring. American Journal of Hematology 2025; 100(7), 1205-1231.
QL Ekpa, PC Akahara, AM Anderson, OO Adekoya, OO Ajayi, PO Alabi, OE Okobi, O Jaiyeola and MS Ekanem. A review of Acute Lymphocytic Leukemia (ALL) in the pediatric population: Evaluating current trends and changes in guidelines in the past decade. Cureus 2023; 15(12), e49930.
R Salvaris and PL Fedele. Targeted therapy in acute lymphoblastic leukaemia. Journal of Personalized Medicine 2021; 11(8), 715.
J Liu, D Tran, L Xue, BJ Wiley, C Vlasschaert, CJ Watson, HAJ MacGregor, X Zong, ICC Chan, I Das, MM Uddin, A Niroula, G Griffin, BL Ebert, T Mack, Y Pershad, B Sharber, M Berger, A Zehir, …, KL Bolton. Germline genetic variation impacts clonal hematopoiesis landscape and progression to malignancy. Nature Genetics 2025; 57(8), 1872-1880.
F El Chaer, M Keng and KK Ballen. MLL-rearranged acute lymphoblastic leukemia. Current Hematologic Malignancy Reports 2020; 15(2), 83-89.
N Aziz, YH Hong, HG Kim, JH Kim and JY Cho. Tumor-suppressive functions of protein lysine methyltransferases. Experimental & Molecular Medicine 2023; 55(12), 2475-2497.
L Reed, J Abraham, S Patel and SS Dhar. Epigenetic modifiers: Exploring the roles of histone methyltransferases and demethylases in cancer and neurodegeneration. Biology 2024; 13(12), 1008.
M Górecki, I Kozioł, A Kopystecka, J Budzyńska, J Zawitkowska and M Lejman. Updates in KMT2A Gene rearrangement in pediatric acute lymphoblastic leukemia. Biomedicines 2023; 11(3), 821.
C Meyer, P Larghero, BALopes, T Burmeister, D Gröger, R Sutton, NC Venn, G Cazzaniga, LC Abascal, G Tsaur, L Fechina, M Emerenciano, MS Pombo-de-Oliveira, T Lund-Aho, T Lundán, M Montonen, V Juvonen, J Zuna, J Trka, …, R Marschalek. The KMT2A recombinome of acute leukemias in 2023. Leukemia 2023; 37(5), 988-1005.
L Oksa, A Mäkinen, A Nikkilä, N Hyvärinen, S Laukkanen, A Rokka, P Haapaniemi, M Seki, J Takita, O Kauko, M Heinäniemi and O Lohi. Arginine methyltransferase PRMT7 deregulates expression of RUNX1 target genes in T-cell acute lymphoblastic leukemia. Cancers 2022; 14(9), 2169.
A Montanaro, S Kitara, E Cerretani, M Marchesini, C Rompietti, L Pagliaro, A Gherli, A Su, ML Minchillo, M Caputi, R Fioretzaki, B Lorusso, L Ross, G Alexe, E Masselli, M Marozzi, FMA Rizzi, RL Starza, C Mecucci, …, G Roti. Identification of an Epi-metabolic dependency on EHMT2/G9a in T-cell acute lymphoblastic leukemia. Cell Death & Disease 2022; 13(6), 551
M Khodadoust and O Silva. Hepatosplenic T-cell lymphoma with STAT5B and SETD2 mutations recurring as cells with NK-cell immunophenotype. Blood, The Journal of the American Society of Hematology 2023; 141(5), 555.
J Li, J Hlavka-Zhang, JH Shrimp, C Piper, D Dupéré-Richér, JS Roth, D Jing, HLC Román, C Troche, A Swaroop, M Kulis, JA Oyer, CM Will, M Shen, A Riva, RL Bennett, AA Ferrando, MD Hall, RB Lock and JD Licht. PRC2 inhibitors overcome glucocorticoid resistance driven by NSD2 mutation in pediatric acute lymphoblastic leukemia. Cancer Discovery 2022; 12(1), 186-203.
DY Nie, JR Tabor, J Li, M Kutera, J St-Germain, RP Hanley, E Wolf, E Paulakonis, TMG Kenney, S Duan, S Shrestha, DDG Owens, MER Maitland, A Pon, M Szewczyk, AJ Lamberto, M Menes, F Li, LZ Penn, …., CH Arrowsmith. Recruitment of FBXO22 for targeted degradation of NSD2. Nature Chemical Biology 2024; 20(12), 1597-1607.
SW Brady, KG Roberts, Z Gu, L Shi, S Pounds, D Pei, C Cheng, Y Dai, M Devidas, C Qu, AN Hill, D Payne-Turner, X Ma, I Iacobucci, P Baviskar, L Wei, S Arunachalam, K Hagiwara, Y Liu, …, CG Mullighan. The genomic landscape of pediatric acute lymphoblastic leukemia. Nature Genetics 2022; 54(9), 1376-1389.
M Mounir, M Lucchetta, TC Silva, C Olsen, G Bontempi, X Chen, H Noushmehr, A Colaprico and E Papaleo. New functionalities in the TCGAbiolinks package for the study and integration of cancer data from GDC and GTEx. PLoS Computational Biology 2019; 15(3), e1006701.
D Long, Y Xue, X Yu, X Qin, J Chen, J Luo, K Ma, L Wei and X Li. Integrative multi-omics analysis reveals the molecular characteristics, tumor microenvironment, and clinical significance of ubiquitination mechanisms in lung adenocarcinoma. International Journal of Molecular Sciences 2025; 26(13), 6501.
E Capriotti, P Fariselli and R Casadio. I-Mutant2.0: Predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Research 2005; 33(S2), W306-W310.
DK Sarker, P Ray, FBA Salam and SJ Uddin. Exploring the impact of deleterious missense nonsynonymous single nucleotide polymorphisms in the DRD4 gene using computational approaches. Scientific Reports 2025; 15(1), 3150.
MT Terry. A package for survival analysis in R. R package version 3.8-3, Available at https://CRAN.R-project.org/package=survival, accessed June 2025.
Y Chen, L Chen, ATL Lun, PL Baldoni and GK Smyth. edgeR v4: Powerful differential analysis of sequencing data with expanded functionality and improved support for small counts and larger datasets. Nucleic Acids Research 2025; 53(2), gkaf018.
C W Law, Y Chen, W Shi and GK Smyth. Voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biology 2014; 15(2), R29.
M Shahjaman, MMH Mollah, MR Rahman, SMS Islam and MNH Mollah. Robust identification of differentially expressed genes from RNA-seq data. Genomics 2020; 112(2), 2000-2010.
Z Gu. Complex heatmap visualization. iMeta 2022; 1(3), e43.
S Xu, E Hu, Y Cai, Z Xie, X Luo, L Zhan, W Tang, Q Wang, B Liu, R Wang, W Xie, T Wu, L Xie and G Yu. Using clusterProfiler to characterize multiomics data. Nature Protocols 2024; 19(11), 3292-3320.
J Bendl, J Stourac, O Salanda, A Pavelka, ED Wieben, J Zendulka, J Brezovsky and J Damborsky. PredictSNP: robust and accurate consensus classifier for prediction of disease-related mutations. PLoS Computational Biology 2014; 10(1), e1003440.
R Reshmi and D Kaur. In silico analysis of non-synonymous single nucleotide polymorphisms of human ABCD1 gene associated with adrenoleukodystrophy. Egyptian Journal of Medical Human Genetics 2025; 26(1), 144.
EA Stone and A Sidow. Physicochemical constraint violation by missense substitutions mediates impairment of protein function and disease severity. Genome research 2005; 15(7), 978-986.
E Capriotti and P Fariselli. PhD-SNPg: Updating a webserver and lightweight tool for scoring nucleotide variants. Nucleic Acids Research 2023; 51(W1), W451-W458.
I Adzhubei, DM Jordan and SR Sunyaev. Predicting functional effect of human missense mutations using PolyPhen-2. Current Protocols in Human Genetics 2013; 76(1), 7-20.
PC Ng and S Henikoff. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Research 2003; 31(13), 3812-3814.
NL Sim, P Kumar, J Hu, S Henikoff, G Schneider and PC Ng. SIFT web server: Predicting effects of amino acid substitutions on proteins. Nucleic Acids Research 2012; 40(W1), W452-W457.
Y Bromberg, G Yachdav and B Rost. SNAP predicts effect of mutations on protein function. Bioinformatics 2008; 24(20), 2397-2398.
H Tang and PD Thomas. PANTHER-PSEP: Predicting disease-causing genetic variants using position-specific evolutionary preservation. Bioinformatics 2016; 32(14), 2230-2232.
J Ma, Y Zhang, J Li, Y Dang and D Hu. Regulation of histone H3K27 methylation in inflammation and cancer. Molecular Biomedicine 2025; 6(1), 14.
H Chen, S Wang, R Dong, P Yu, T Li, L Hu, M Wang, Z Qian, H Zhou, X Yue, L Wang and H Xiao. KDM6A deficiency induces myeloid bias and promotes CMML-like disease through JAK/STAT3 activation by repressing SOCS3. Advanced Science 2025; 12(21), 2413091.
H Chen, H Huang and H Xiao. Kdm6a modulates hematopoiesis and leukemogenesis via demethylase-dependent epigenetic programming. Blood 2023; 142(S1), 1372-1372.
S Chen, D Liu, B Chen, Z Li, B Chang, C Xu, N Li, C Feng, X Hu, W Wang, Y Zhang, Y Xie, Q Huang, Y Wang, S D Nimer, S Chen, Z Chen, L Wang and X Sun. Catalytic activity of Setd2 is essential for embryonic development in mice: Establishment of a mouse model harboring patient-derived Setd2 mutation. Frontiers of Medicine 2024; 18(5), 831-849.
RM Chavez, DR Powell, K Lakhani, J Attelah, E Flynt, T Connolly, M Hamilton, G Mulligan, D Auclair, J Keats, PM Vertino, LH Boise, S Lonial, KN Conneely and BG Barwick. Ectopic NSD2 remodels H3K36me2 and DNA methylation to promote oncogenic gene expression in multiple myeloma. Blood 2024; 144(S1), 1359.
PSY Chong, JY Chooi, JSL Lim, SHM Toh, TZ Tan and WJ Chng. SMARCA2 is a novel interactor of NSD2 and regulates prometastatic PTP4A3 through chromatin remodeling in t(4;14) multiple myeloma. Cancer Research 2021; 81(9), 2332-2344.
K Wang, F Zhan, X Yang, M Jiao, P Wang, H Zhang, W Shang, J Deng and L Wang. KMT2D: A key emerging epigenetic regulator in head and neck diseases and tumors. Life Sciences 2025; 12(21), 2413091.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






