GAXGB: A Two-Stage Ensemble Framework Integrating Genetic Algorithms and XGBoost for Anti-HIV Peptide Prediction
DOI:
https://doi.org/10.48048/tis.2026.11717Keywords:
Classification model, Anti-HIV peptides, Genetic algorithms, Ensemble learning methodsAbstract
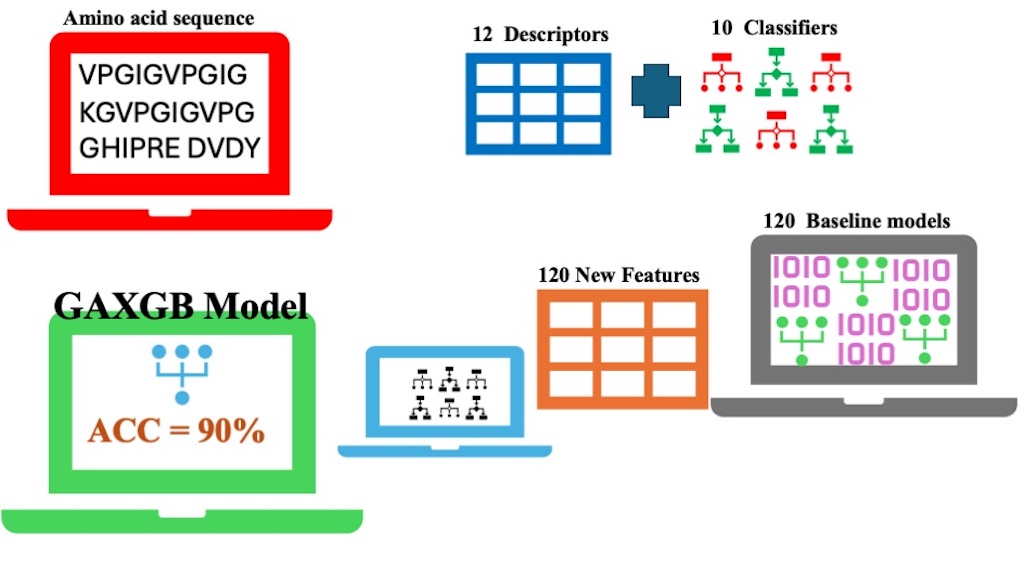
This study applied a computational approach to derive amino acid sequence features relevant to AIDS treatment. A predictive model, GAXGB, was developed to classify anti-HIV peptides based on their amino acid sequence characteristics. The model was built using a 2-stage learning procedure. In the first stage, features were extracted from amino acid sequences using 12 descriptors, and 120 baseline models were constructed with 10 different classifiers. In the second stage, these baseline models generated 120 predictive probability scores, which were used as input features. Various feature selection methods, including chi-square, ANOVA, mutual information, and a genetic algorithm, were employed to identify the most significant features for the final model. Subsequently, 10 classifiers were trained and evaluated. Performance evaluation showed that the GAXGB model, which combines genetic algorithm-based feature selection with an XGBoost classifier, achieved superior predictive accuracy. The model reached an accuracy of 90%, significantly outperforming other models that achieved approximately 80% accuracy. This approach offers a promising tool to accelerate the design and discovery of novel anti-HIV peptides for AIDS treatment.
HIGHLIGHTS
- Developed a 2-stage predictive framework (GAXGB) integrating genetic algorithm and XGBoost for anti-HIV peptide classification.
- Extracted features from amino acid sequences using 12 descriptors and 120 baseline models.
- Applied multiple feature selection methods to identify the most relevant predictive features.
- Achieved superior accuracy (90%), outperforming other models (~80%).
- Provides a promising tool to accelerate the design and discovery of novel anti-HIV peptide.
GRAPHICAL ABSTRACT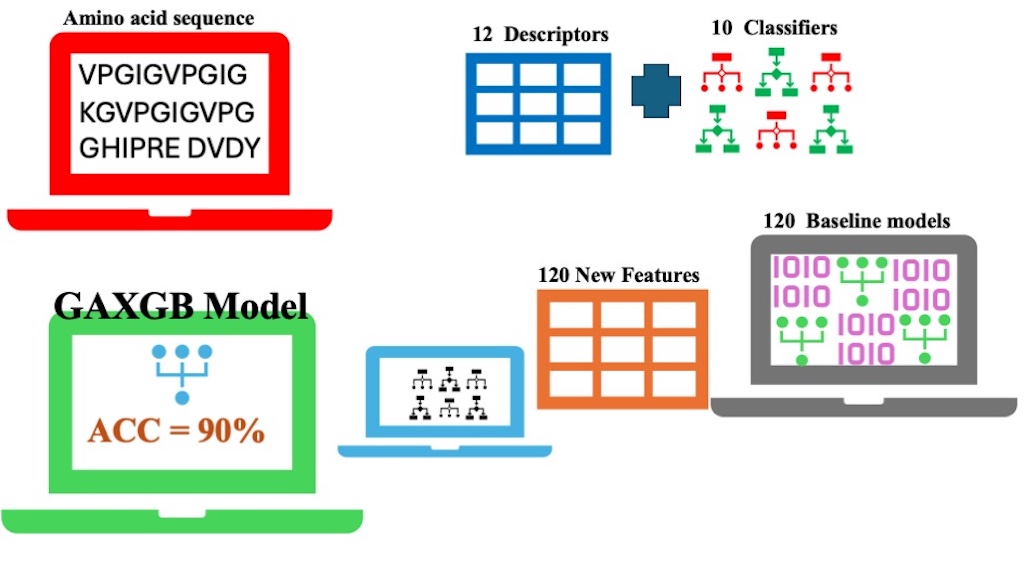
Downloads
References
C Chen, W He, TY Nassirou, A Nsabiyumva, X Dong, YMN Adedze and D Jin. Molecular characterization and genetic diversity of different genotypes of Oryza sativa and Oryza glaberrima. Electronic Journal of Biotechnology 2017; 30, 48-57.
D Zhou, SM Dai and Q Tong. COVID-19: A recommendation to examine the effect of hydroxychloroquine in preventing infection and progression. Journal of Antimicrobial Chemotherapy 2020; 75(7), 1667-1670.
MC Aguilera-Puga, NL Cancelarich, MM Marani, C de la Fuente-Nunez and F Plisson. Accelerating the discovery and design of antimicrobial peptides with artificial intelligence. Methods in Molecular Biology 2024; 2714, 329-352.
S Sachdeva. Peptides as ‘drugs’: The journey so far. International Journal of Peptide Research and Therapeutics 2017; 23(1), 49-60.
C Li, D Sutherland, SA Hammond, C Yang, F Taho, L Bergman, S Houston, RL Warren, T Wong and LM Hoang. AMPlify: Attentive deep learning model for discovery of novel antimicrobial peptides effective against WHO priority pathogens. BMC genomics 2022; 23(1), 77.
N Schaduangrat, C Nantasenamat, V Prachayasittikul and W Shoombuatong. ACPred: A computational tool for the prediction and analysis of anticancer peptides. Molecules 2019; 24(10), 1973.
W Shoombuatong, N Schaduangrat and C Nantasenamat. Unraveling the bioactivity of anticancer peptides as deduced from machine learning. EXCLI journal 2018; 17, 734.
P Charoenkwan, N Anuwongcharoen, C Nantasenamat, MM Hasan and W Shoombuatong. In silico approaches for the prediction and analysis of antiviral peptides: a review. Current pharmaceutical design 2021; 27(18), 2180-2188.
AC Kaushik, A Mehmood, S Peng, YJ Zhang, X Dai and DQ Wei. A-CaMP: A tool for anti-cancer and antimicrobial peptide generation. Journal of Biomolecular Structure and Dynamics 2021; 39(1), 285-293.
Y Pang, L Yao, JH Jhong, Z Wang and TY Lee. AVPIden: A new scheme for identification and functional prediction of antiviral peptides based on machine learning approaches. Briefings in Bioinformatics 2021; 22(6), bbab263.
P Charoenkwan, W Chiangjong, VS Lee, C Nantasenamat, MM Hasan and W Shoombuatong. Improved prediction and characterization of anticancer activities of peptides using a novel flexible scoring card method. Scientific Reports 2021; 11(1), 3017.
P Charoenkwan, N Schaduangrat, B Manavalan and W Shoombuatong. M3S-ALG: Improved and robust prediction of allergenicity of chemical compounds by using a novel multi-step stacking strategy. Future Generation Computer Systems 2024; 162, 107455.
N Thakur, A Qureshi and M Kumar. AVPpred: Collection and prediction of highly effective antiviral peptides. Nucleic Acids Research 2012; 40(W1), W199-W204.
JH Jhong, L Yao, Y Pang, Z Li, CR Chung, R Wang, S Li, W Li, M Luo and R Ma. dbAMP 2.0: Updated resource for antimicrobial peptides with an enhanced scanning method for genomic and proteomic data. Nucleic Acids Research 2022; 50(D1), D460-D470.
A Qureshi, N Thakur, H Tandon and M Kumar. AVPdb: A database of experimentally validated antiviral peptides targeting medically important viruses. Nucleic acids research 2014; 42(D1), D1147-D1153.
S Singh, K Chaudhary, SK Dhanda, S Bhalla, SS Usmani, A Gautam, A Tuknait, P Agrawal, D Mathur and GP Raghava. SATPdb: A database of structurally annotated therapeutic peptides. Nucleic Acids Research 2016; 44(D1), D1119-D1126.
G Shi, X Kang, F Dong, Y Liu, N Zhu, Y Hu, H Xu, X Lao and H Zheng. DRAMP 3.0: An enhanced comprehensive data repository of antimicrobial peptides. Nucleic Acids Research 2022; 50(D1), D488-D496.
M Pirtskhalava, AA Amstrong, M Grigolava, M Chubinidze, E Alimbarashvili, B Vishnepolsky, A Gabrielian, A Rosenthal, DE Hurt and M Tartakovsky. DBAASP v3: Database of antimicrobial/cytotoxic activity and structure of peptides as a resource for development of new therapeutics. Nucleic Acids Research 2021; 49(D1), D288-D297.
A Qureshi, N Thakur and M Kumar. HIPdb: A database of experimentally validated HIV inhibiting peptides. PloS One 2013; 8(1), e54908.
N Poorinmohammad and H Mohabatkar. A comparison of different machine learning algorithms for the prediction of anti-HIV-1 peptides based on their sequence-related properties. International Journal of Peptide Research and Therapeutics 2015; 21(1), 57-62.
N Poorinmohammad, H Mohabatkar, M Behbahani and D Biria. Computational prediction of anti HIV‐1 peptides and in vitro evaluation of anti HIV‐1 activity of HIV‐1 P24‐derived peptides. Journal of peptide science 2015; 21(1), 10-16.
KC Chou. Pseudo amino acid composition and its applications in bioinformatics, proteomics and system biology. Current Proteomics 2009; 6(4), 262-274.
M Esmaeili, H Mohabatkar and S Mohsenzadeh. Using the concept of Chou’s pseudo amino acid composition for risk type prediction of human papillomaviruses. Journal of Theoretical Biology 2010; 263(2), 203-209.
JA Suykens and J Vandewalle. Training multilayer perceptron classifiers based on a modified support vector method. IEEE Transactions on Neural Networks 1999; 10(4), 907-911.
S Suthaharan. Support vector machine. In: S Suthaharan (Ed.). Machine learning models and algorithms for big data classification: Thinking with examples for effective learning. Springer, New York, 2016, p. 207-235.
B Mathiyazhagan, J Liyaskar, AT Azar, HH Inbarani, Y Javed, NA Kamal and KM Fouad. Rough set based classification and feature selection using improved harmony search for peptide analysis and prediction of anti-HIV-1 activities. Applied Sciences 2022; 12(4), 2020.
G Wang, X Li and Z Wang. APD3: The antimicrobial peptide database as a tool for research and education. Nucleic Acids Research 2016; 44(D1), D1087-D1093.
Y Liu, Y Zhu, X Sun, T Ma, X Lao and H Zheng. DRAVP: A Comprehensive database of antiviral peptides and proteins. Viruses 2023; 15(4), 820.
Y Huang, B Niu, Y Gao, L Fu and W Li. CD-HIT Suite: A web server for clustering and comparing biological sequences. Bioinformatics 2010; 26(5), 680-682.
P Charoenkwan, N Schaduangrat and W Shoombuatong. StackTTCA: A stacking ensemble learning-based framework for accurate and high-throughput identification of tumor T cell antigens. BMC Bioinformatics 2023; 24(1), 301.
Z Chen, P Zhao, F Li, A Leier, TT Marquez-Lago, Y Wang, GI Webb, AI Smith, RJ Daly and KC Chou. iFeature: A python package and web server for features extraction and selection from protein and peptide sequences. Bioinformatics 2018; 34(14), 2499-2502.
F Pedregosa, G Varoquaux, A Gramfort, V Michel, B Thirion, O Grisel, M Blondel, P Prettenhofer, R Weiss and V Dubourg. Scikit-learn: Machine learning in Python. the Journal of Machine Learning Research 2011; 12, 2825-2830.
J Kim and S Yoo. Software review: DEAP (distributed evolutionary algorithm in python) library. Genetic Programming and Evolvable Machines 2019; 20(1), 139-142.
D Chicco and G Jurman. The advantages of the Matthews correlation coefficient (MCC) over F1 score and accuracy in binary classification evaluation. BMC Genomics 2020; 21(1), 6.
P Fourie. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Plant Disease 2020; 22, 1658.
TE Creighton. Proteins: Structures and molecular properties. 2nd ed. W. H. Freeman and Company, New York, 1993.
M Fisher. Lehninger principles of biochemistry, 3rd edition; By David L. Nelson and Michael M. Cox. The Chemical Educator 2001; 6, 69-70.
P Charoenkwan, W Chiangjong, C Nantasenamat, MM Hasan, B Manavalan and W Shoombuatong. StackIL6: A stacking ensemble model for improving the prediction of IL-6 inducing peptides. Briefings in Bioinformatics 2021; 22(6), bbab172.
V Laengsri, C Nantasenamat, N Schaduangrat, P Nuchnoi, V Prachayasittikul and W Shoombuatong. TargetAntiAngio: A sequence-based tool for the prediction and analysis of anti-angiogenic peptides. International Journal of Molecular Sciences 2019; 20(12), 2950.
R Barrett, S Jiang and AD White. Classifying antimicrobial and multifunctional peptides with Bayesian network models. Peptide Science 2018; 110(4), e24079.
M Lu and T Gibson. Development of predictive tools for anti-cancer peptide candidates using generative machine learning models. The Journal of Young Investigators 2021; 39(5), 60-64.
J Lin, L Wen, Y Zhou, S Wang, H Ye, J Su, J Li, J Shu, J Huang and P Zhou. PepQSAR: A comprehensive data source and information platform for peptide quantitative structure - activity relationships. Amino Acids 2023; 55(2), 235-242.
Y Wang and C Zhou. Feature selection method based on chi-square test and minimum redundancy. In: Proceedings of the 5th International Conference on Intelligent and Interactive Systems and Application, Shanghai, China. 2020, p. 171-178.
E Asad, A Islam, A Alam and AF Mollah. Univariate feature fitness measures for classification problems: An empirical assessment. In: Proceedings of the 5th International Conference, AMLDA, Tamaulipas, Mexico. 2022, p. 13-26.
B Alhijawi and A Awajan. Genetic algorithms: Theory, genetic operators, solutions, and applications. Evolutionary Intelligence 2024; 17(3), 1245-1256.
T Alam, S Qamar, A Dixit and M Benaida. Genetic algorithm: Reviews, implementations, and applications. International Journal of Engineering Pedagogy 2020; 10(6), 57-77.
D Yang, Z Yu, H Yuan and Y Cui. An improved genetic algorithm and its application in neural network adversarial attack. Plos One 2022; 17(5), e0267970.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






