In Vitro Antiviral Activity and Molecular Docking Analysis of Dihydropyrimidinone, Chromene and Chalcone Derivatives Against SARS-CoV-2
DOI:
https://doi.org/10.48048/tis.2026.11438Keywords:
Antiviral, Chalcone, Chromene, Dihydropyrimidinone, SARS-CoV-2Abstract
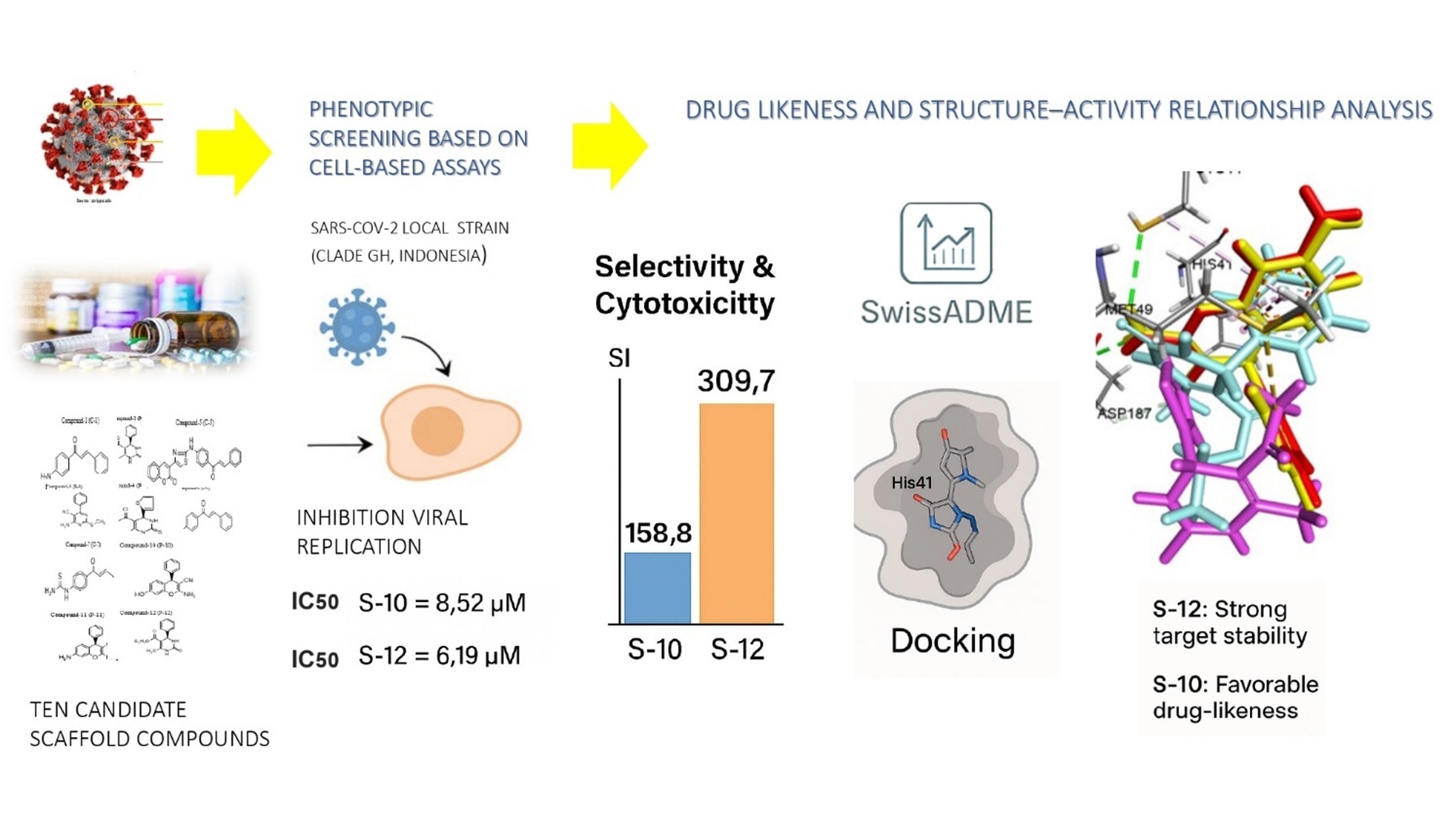
To identify and develop effective antiviral agents against SARS-CoV-2, research is ongoing in the field of drug discovery. Heterocyclic compounds such as Chalcone, Chromene, and Dihydropyrimidinone (DHPM) derivatives are of particular interest because of their structural diversity, easily synthesized and broad pharmacological activities, including antiviral effects. Therefore, we investigated the potential of these derivatives against SARS-CoV-2 using cell-based assays, supported by in silico studies. Ten synthetic compounds, consist of 4 DHPMs, 2 chromenes, and 4 chalcones were tested for cytotoxicity and antiviral activity in Vero cells infected with a locally isolated SARS-CoV-2 strain (Clade GH, GISAID: EPI_ISL_529965), using a phenotypic screening approach. Moreover, in silico analyses were performed to predict their drug-likeness, including absorption, distribution, metabolism, and excretion properties, using SwissADME. Molecular docking was conducted to evaluate the binding affinity and interactions of compounds with the SARS-CoV-2 main protease (Mpro). Among the tested compounds, S-10 (chromene derivative) and S-12 (DHPM derivative) demonstrated exhibited significant antiviral activity compared to the other tested derivatives (p < 0.05), with IC₅₀ values of 8.52 ± 0.28 µM and 6.19 ± 0.41 µM, and selectivity index (SI) of 158.8 and 309.7, respectively, indicating strong efficacy and high safety margins. In silico ADME suggested favorable drug-likeness profiles without major toxicity alerts. Molecular docking further revealed that both compounds established stable interactions with the SARS-CoV-2 main protease (Mpro), particularly at the catalytic residues His-41 and Cys-145, supporting their in vitro activity. These results suggest their potential as lead compounds for further drug development, supporting the use of integrated phenotypic and computational approaches in the discovery of region-specific antiviral agents.
HIGHLIGHTS
- Two novel compounds (S-10 and S-12) were identified with strong anti-SARS-CoV-2 activity and high selectivity index (SI > 100), compared to the other 8 compounds tested, out of 10 Dihydropyrimidinone, Chromene, and Chalcone derivative candidates, suggesting high efficacy and low cytotoxicity.
- Phenotypic screening was performed against a locally isolated SARS-CoV-2 strain (Clade GH, Indonesia) provided, direct biological relevance rather than relying solely on predictive models and supporting region-specific therapeutic development.
- Supplementary in silico analysis (ADME profiling, toxicity prediction, and molecular docking to Mpro) offers, a comprehensive assessment of efficacy, safety, and mechanism of action.
GRAPHICAL ABSTRACT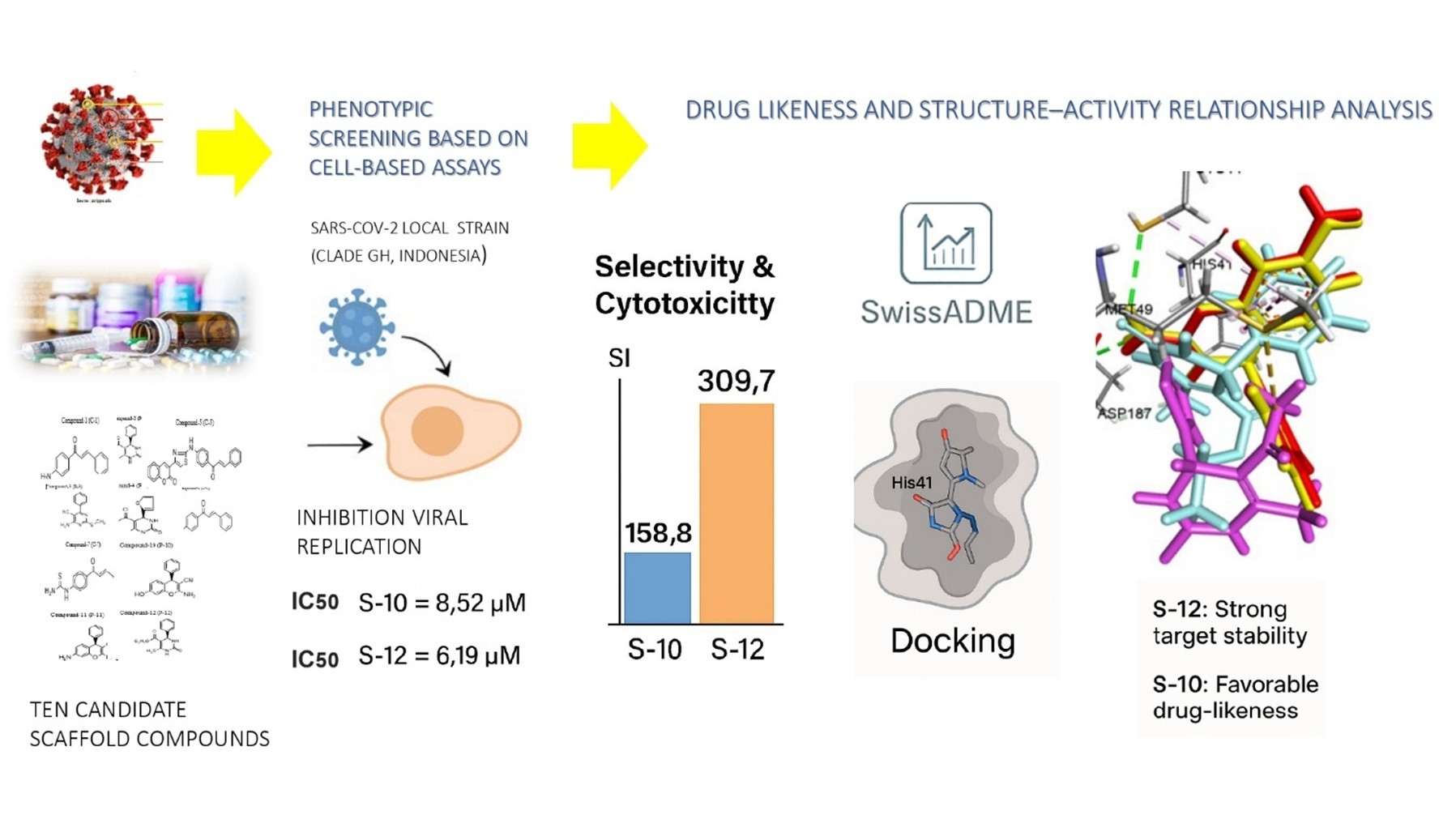
Downloads
References
H Zhu, L Wei and P Niu. The novel coronavirus outbreak in Wuhan, China. Global Health Research and Policy 2020; 5(1), 6.
B Hu, H Guo, P Zhou and ZL Shi. Characteristics of SARS-CoV-2 and COVID-19. Nature Reviews Microbiology 2021; 19(3), 141154.
A Gori, F Fama and C Genovese. Coronavirus disease 19 (COVID-19). In: MCB Raviglione, F Tediosi, S Villa, N Casamitjana and A Plasència (Eds.). Global Health Essentials. Springer, Cham, Switzerland, 2023, p. 137-142.
I Polatoğlu, T Oncu-Oner, I Dalman and S Ozdogan. COVID-19 in early 2023: Structure, replication mechanism, variants of SARS-CoV-2, diagnostic tests, and vaccine & drug development studies. MedComm 2023; 4(2), e228.
Purwati, A Miatmoko, Nasronudin, E Hendrianto, D Karsari, A Dinaryanti, N Ertanti, IS Ihsan, DS Purnama, TP Asmarawati, E Marfiani, Yulistiani, AN Rosyid, PA Wulaningrum, HW Setiawan, I Siswanto and NNT Puspaningsih. An in vitro study of dual drug combinations of anti-viral agents, antibiotics, and/or hydroxychloroquine against the SARS-CoV-2 virus isolated from hospitalized patients in Surabaya, Indonesia. PLoS One 2021; 16(6), e0252302.
H Zhou, WJ Ni, W Huang, Z Wang, M Cai and YC Sun. Advances in pathogenesis, progression, potential targets and targeted therapeutic strategies in SARS-CoV-2-induced COVID-19. Frontiers in Immunology 2022; 13, 834942.
JD Rigg. The Sustainable Development Goals (SDGS). In: The companion to development studies. Routledge, Oxfordshire, 2024, p. 253-257.
EL Berg. The future of phenotypic drug discovery. Cell Chemical Biology 2021; 28(3), 424-430.
F Vincent, A Nueda, J Lee, M Schenone, M Prunotto and M Mercola. Phenotypic drug discovery: Recent successes, lessons learned and new directions. Nature Reviews Drug Discovery 2023; 21(12), 899-914.
S Wang, Z Wang, L Fang, Y Lv and G Du. Advances of the target-based and phenotypic screenings and strategies in drug discovery. Internatioal Journal of Drug Discovery and Pharmacology 2022; 1(1), 5-11.
SE Rhabori, AE Aissouq, S Chtita and F Khalil. QSAR, molecular docking and ADMET studies of quinoline, isoquinoline and quinazoline derivatives against Plasmodium falciparum malaria. Structural Chemistry 2023; 34(2), 585-603.
AM Andrianov, YV Kornoushenko, AD Karpenko, IP Bosko and AV Tuzikov. Computational discovery of small drug-like compounds as potential inhibitors of SARS-CoV-2 main protease. Journal of Biomolecular Structure and Dynamics 2021; 39(15), 5779-5791.
I Amalina, NNT Puspaningsih and H Suwito. In silico analysis of pyrimidine derivatives as potential antibacterial agents. In: Proceedings of the International Conference on Advanced Technology and Multidiscipline, Surabaya, Indonesia. 2023, p. 50004.
LHS Matos, FT Masson, LA Simeoni and M Homem-de-Mello. Biological activity of dihydropyrimidinone (DHPM) derivatives: A systematic review. European Journal of Medical Chemistry 2018; 143, 1779-1789.
AN Kristanti, H Suwito, NS Aminah, KU Haq, HD Hardiyanti, H Anggraeni, N Faiza, RS Anto and S Muharromah. Synthesis of some chalcone derivatives, in vitro and in silico toxicity evaluation. Rasayan Journal of Chemistry 2020; 13(1), 654-662.
A Chaudhary, K Singh, N Verma, S Kumar, D Kumar and PP Sharma. Chromenes - A novel class of heterocyclic compounds: Recent advancements and future directions. Mini Reviews in Medicinal Chemistry 2022; 22(21), 2736-2751.
V Raj and J Lee. 2H/4H-chromenes - A versatile biologically attractive scaffold. Frontiers in Chemistry 2020; 8, 623.
S Khasimbi, F Ali, K Manda, A Sharma, G Chauhan and S Wakode. Dihydropyrimidinones scaffold as a promising nucleus for synthetic profile and various therapeutic targets: A review. Current Organic Synthesis 2020; 18(3), 270-293.
BT Dharmapalan, R Biswas, S Sankaran, B Venkidasamy, M Thiruvengadam, G George, M Rebezov, G Zengin, M Gallo, D Montesano, D Naviglio and MA Shariati. Inhibitory potential of chromene derivatives on structural and non-structural proteins of dengue virus. Viruses 2022; 14(12), 2656.
SM Villa, J Heckman and D Bandyopadhyay. Medicinally privileged natural chalcones: Abundance, mechanisms of action, and clinical trials. International Journal of Molecular Sciences 2024; 25(17), 9623.
SY Ham, H Jeong, J Jung, ES Kim, KU Park, HB Kim, JS Park and KH Song. Performance of STANDARDTM M10 SARS-CoV-2 assay for the diagnosis of COVID-19 from a nasopharyngeal swab. Infection and Chemotherapy 2022; 54(2), 360-363.
Y Shu and J McCauley. GISAID: Global initiative on sharing all influenza data – from vision to reality. Euro Surveillance 2017; 22(13), 30494.
K Rahardjo, AM Nastri, JR Dewantari, RR Prasetya, J Wahyuhadi, G Soegiarto, L Wulandari, RA Setyoningrum, R Yudhawati, YK Shimizu, M Nishimura, Y Mori, Soetjipto, K Shimizu and MI Lusida. GISAID: EpiCoV database. 2020. https://www.epicov.org/epi3/frontend#3280a8.
M Brandolini, F Taddei, MM Marino, L Grumiro, A Scalcione, ME Turba, F Gentilini, M Fantini, S Zannoli, G Dirani and V Sambri. Correlating qRT-PCR, dPCR and viral titration for the identification and quantification of SARS-CoV-2: A new approach for infection management. Viruses 2021; 13(6), 1022.
G McDonnell and DM Hansen. Block’s disinfection, sterilization and preservation. 6th eds. Lippincott Williams and Wilkins, Philadelphia, 2020.
AA Permanasari, C Aoki-Utsubo, TS Wahyuni, L Tumewu, M Adianti, A Widyawaruyanti, H Hotta and AF Hafid. An in vitro study of an Artocarpus heterophyllus substance as a hepatitis C antiviral and its combination with current anti-HCV drugs. BMC Complementary Medicine and Therapies 2021; 21(1), 260.
R Ramadhan, K Ul-Haq, P Phuwapraisirisan, FA Puspitasari and H Suwito. α-glucosidase inhibitory activities and antioxidant properties of depsidone from Garcinia parvifolia. Rasayan Journal of Chemistry 2023; 16(3), 1811-1817.
H Suwito, R Ramadhan, A Abdulloh and KU Haq. Synthesis of dihydrotetrazolopyrimidine derivatives as anticancer agents and inhibitor of α-glucosidase. Rasayan Journal of Chemistry 2023; 16(1), 147-158.
A Purniawan, MI Lusida, RW Pujiyanto, AM Nastri, AA Permanasari, AAH Harsono, NH Oktavia, ST Wicaksono, JR Dewantari, RR Prasetya, K Rahardjo, M Nishimura, Y Mori and K Shimizu. Synthesis and assessment of copper-based nanoparticles as a surface coating agent for antiviral properties against SARS-CoV-2. Scientific Reports 2022; 12(1), 4835.
VNT La, S Nicholson, A Haneef, L Kang and DDL Minh. Inclusion of control data in fits to concentration-response curves improves estimates of half-maximal concentrations. Journal of Medical Chemistry 202; 66(18), 12751-12761.
L Grumiro, M Brandolini, G Gatti, A Scalcione, F Taddei, G Dirani, A Mancini, A Denicolò, M Manera, S Zannoli, MM Marino, M Morotti, V Arfilli, A Battisti, M Cricca and V Sambri. Evaluation of STANDARDTM M10 SARS-CoV-2, a novel cartridge-based real-time PCR assay for the rapid identification of severe acute respiratory syndrome coronavirus 2. Applied Microbiology 2022; 2(4), 873-881.
A Daina, O Michielin and V Zoete. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Scientific Reports 2017; 7, 42717.
S Abu-Melha, MM Edrees, MA Said, SM Riyadh, NS Al-Kaff and SM Gomha. Potential COVID-19 drug candidates based on diazinyl-thiazol-imine moieties: Synthesis and greener pastures biological study. Molecules 2022; 27(2), 488.
CA Lipinski, F Lombardo, BW Dominy and PJ Feeney. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Advanced Drug Delivery Reviews 2001; 46(1-3), 3-25.
R Yadav, M Imran, P Dhamija, DK Chaurasia and S Handu. Virtual screening, ADMET prediction and dynamics simulation of potential compounds targeting the main protease of SARS-CoV-2. Journal of Biomolecular Structure and Dynamics 2021; 39(17), 6617-6632.
L Vegosen and TM Martin. An automated framework for compiling and integrating chemical hazard data. Clean Technologies and Environmental Policy 2020; 22(2), 441-458.
TM Martin. User’s Guide for T.E.S.T. (Toxicity Estimation Software Tool) Version 5.1. 2020, Available at: https://www.epa.gov/chemical-research/toxicity-estimation-software-tool-test, accessed May 2025.
JA Maier, C Martinez, K Kasavajhala, L Wickstrom, KE Hauser and C Simmerling. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. Journal of Chemical Theory and Computation 2015; 11(8), 3696-3713.
S Kumar and S Kumar. Molecular docking: A structure-based approach for drug repurposing. In: K Roy (Ed.). In silico drug design. Academic Press, Massachusetts, 2019, p. 161-189.
MS Coumar. Molecular docking for computer-aided drug design: Fundamentals, Techniques, Resources and Applications. Academic Press, Massachusetts, 2021.
WJ Allen, TE Balius, S Mukherjee, SR Brozell, DT Moustakas, PT Lang, DA Case, ID Kuntz and RC Rizzo. DOCK 6: Impact of new features and current docking performance. Journal of Computational Chemistry 2016; 36(15), 1132-1156.
S Khasimbi, F Ali, K Manda, A Sharma, G Chauhan and S Wakode. Dihydropyrimidinones scaffold as a promising nucleus for synthetic profile and various therapeutic targets: A review. Current Organic Synthesis 2021; 18(3), 270-293.
P Pérez-Ramos, G Biniari, RG Soengas, H Rodríguez-Solla and C Simal. Visible-light photocatalysis for sustainable chromene synthesis and functionalization. Chemistry 2025; 31(29), e202500283.
WW Mar, KU Haq, RA Rusdipoetra, RPFC Samiadji, A Rohman, NNT Puspaningsih and H Suwito. Anticancer activity through inhibition of BCL6BTB of chalcone - Thiourea hybrid compounds: A molecular docking study. AIP Conference Proceedings 2023; 2679(1), 50009.
I Amalina, NNT Puspaningsih and H Suwito. In silico analysis of pyrimidine derivatives as potential antibacterial agents. AIP Conference Proceedings 2023; 2536, 50004.
G Indrayanto, GS Putra and F Suhud. Validation of in-vitro bioassay methods: Application in herbal drug research. Profiles of Drug Substances, Excipients and Related Methodology 2021; 46, 273-307.
AJ Pruijssers, AJ Pruijssers, AS George, A Schäfer, SR Leist, LE Gralinksi, KH Dinnon, BL Yount, ML Agostini, LJ Stevens, JD Chappell, X Lu, TM Hughes, K Gully, DR Martinez, AJ Brown, RL Graham, JK Perry, VD Pont, J Pitts, …, TP Sheahan. Remdesivir inhibits SARS-CoV-2 in human lung cells and chimeric SARS-CoV expressing the SARS-CoV-2 RNA polymerase in mice. Cell Reports 2020; 32(3), 107940.
M Wang, R Cao, L Zhang, X Yang, J Liu, M Xu, Z Shi, Z Hu, W Zhong and G Xiao. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Research 2020; 30(3), 269-271.
B Morak-Młodawska, M Jeleń, E Martula and R Korlacki. Study of lipophilicity and ADME properties of 1,9-diazaphenothiazines with anticancer action. International Journal of Molecular Sciences 2023; 24(8), 6970.
LM Castro-González, JR Alvarez-Idaboy and A Galano. Computationally designed sesamol derivatives proposed as potent antioxidants. ACS Omega 2020; 5(16), 9566-9575.
A Pérez-González, R Castañeda-Arriaga, EG Guzmán-López, LF Hernández-Ayala and A Galano. Chalcone derivatives with a high potential as multifunctional antioxidant neuroprotectors. ACS Omega 2022; 7(43), 38254-38268.
A Fischer, M Sellner, S Neranjan, M Smieško and MA Lill. Potential inhibitors for novel coronavirus protease identified by virtual screening of 606 million compounds. International Journal of Molecular Sciences 2020; 21(10), 3626.
Z Jin, X Du, Y Xu, Y Deng, M Liu, Y Zhao, B Zhang, X Li, L Zhang, C Peng, Y Duan, J Yu, L Wang, K Yang, F Liu, R Jiang, X Yang, T You, X Liu, …, H Yang. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 2020; 582, 289-293.
S Guo, H Xie, Y Lei, B Liu, L Zhang, Y Xu and Z Zuo. Discovery of novel inhibitors against main protease (Mpro) of SARS-CoV-2 via virtual screening and biochemical evaluation. Bioorganic Chemisty 2021; 110, 104767.
B Chatterjee and SS Thakur. Remdesivir and its combination with repurposed drugs as COVID-19 therapeutics. Frontiers in Immunology 2022; 13, 830990.
M Prajapat, Sarma, N Shekhar, P Avti, S Sinha, H Kaur, S Kumar, A Bhattacharyya, H Kumar, S Bansal and B Medhi. Drug targets for corona virus: A systematic review. Indian Journal of Pharmacology 2020; 52(1), 56-65.
A Shirali, V Stebliankin, U Karki, J Shi, P Chapagain and G Narasimhan. A comprehensive survey of scoring functions for protein docking models. BMC Bioinformatics 2025; 26, 25.
A Harmalkar and JJ Gray. Advances to tackle backbone flexibility in protein docking. Current Opinion in Structural Biology 2021; 67, 178-186.
GL Patrick. An introduction to medicinal chemistry. 7th eds. Oxford University Press, Oxford, 2023.
M Hariono, P Hariyono, R Dwiastuti, W Setyani, M Yusuf, N Salin and H Wahab. Potential SARS-CoV-2 3CLpro inhibitors from chromene, flavonoid and hydroxamic acid compound based on FRET assay, docking and pharmacophore studies. Results in Chemistry 2021; 3, 100195.
S Gorai, V Junghare, K Kundu, S Gharui, M Kumar, BS Patro, SK Nayak, S Hazra and S Mula. Synthesis of dihydrobenzofuro[3,2-b]chromenes as potential 3CLpro inhibitors of SARS-CoV-2: A molecular docking and molecular dynamics study. ChemMedChem 2022; 17(8), e202100782.
LH Al-Wahaibi, AM Elshamsy, TFS Ali, BGM Youssif, S Bräse, M Abdel-Aziz and NA El-Koussi. Design and synthesis of new dihydropyrimidine/sulphonamide hybrids as promising anti-inflammatory agents via dual mPGES-1/5-LOX inhibition. Frontiers in Chemistry 2024; 12, 1387923.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






