Sequence Analysis and In Silico Characterization of Lipase from Acinetobacter gerneri CE4.3 and PKY2.2
DOI:
https://doi.org/10.48048/tis.2025.10584Keywords:
Lipase, Lipase genes, Grease traps, Acinetobacter gerneri, In silico, BiotechnologyAbstract
Lipase is an enzyme that catalyzes the breakdown of fats into fatty acids and glycerol. Lipase plays a crucial role in various industries. Bacterial lipases are widely studied and utilized in the biotechnology industry because of their adaptability, efficiency in large-scale production, and abundance in lipid-rich environments. The objectives of this research were to isolate and identify lipase-producing bacteria from grease traps in canteens at the University of Phayao. The bacterial isolates were analyzed for the lipase gene through PCR amplification and sequencing. Characteristics of the lipase protein were predicted through in silico studies. The results revealed that the two isolates were Acinetobacter gerneri (A. gerneri) CE4.3 and PKY2.2, which contain the lipase lip50 and lip66 genes. The nucleotide sequences of the lipase genes and amino acid sequences in both strains showed high similarity to the A. gerneri DSM 14967 (EPR83194.1). The predicted physicochemical properties of Lip50 and Lip66 proteins from both strains indicate that they are slightly basic, thermostable, and hydrophilic. Lip50 and Lip66 proteins of both strains contain conserved domains of the lipase enzyme. The analysis of the phylogenetic tree and multiple sequence alignment of Lip50 and Lip66 proteins from both strains indicated that they belong to lipase family V. In conclusion, Lip50 and Lip66 from A. gerneri CE4.3 and PKY2.2 are the foundations for the study of lipase enzymes for biotechnology.
HIGHLIGHTS
Lipase-producing bacteria isolated from grease traps of canteens at the University of Phayao were identified as Acinetobacter gerneri (A. gerneri) CE4.3 and PKY2.2. Both strains possess lipase genes (lip50 and lip66), which encode proteins (Lip50 and Lip66) containing conserved domains essential for lipase activity. Phylogenetic analysis, together with multiple sequence alignment, indicates that these proteins belong to lipase family V. This study provides fundamental information supporting further research and the potential biotechnological applications of lipase enzymes from A. gerneri.
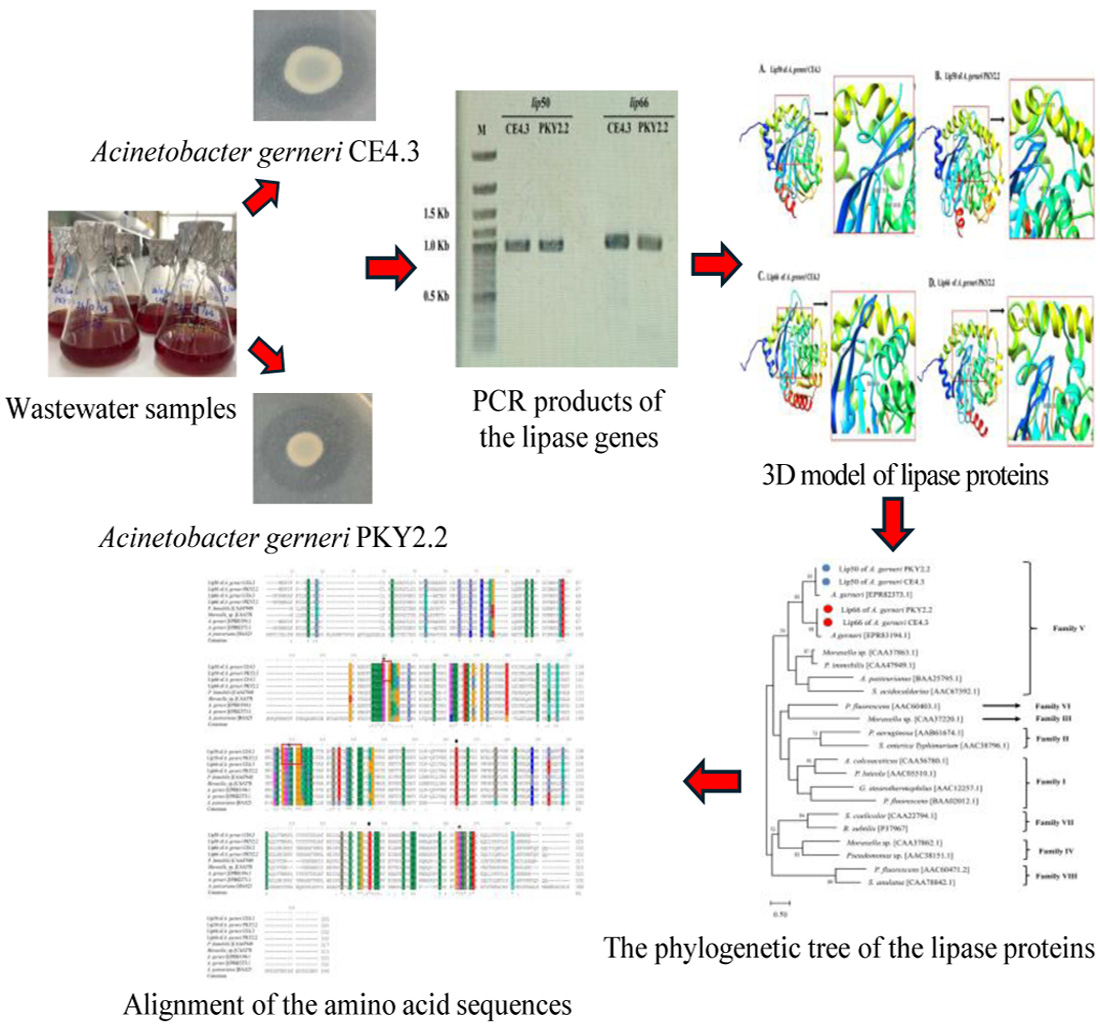
GRAPHICAL ABSTRACT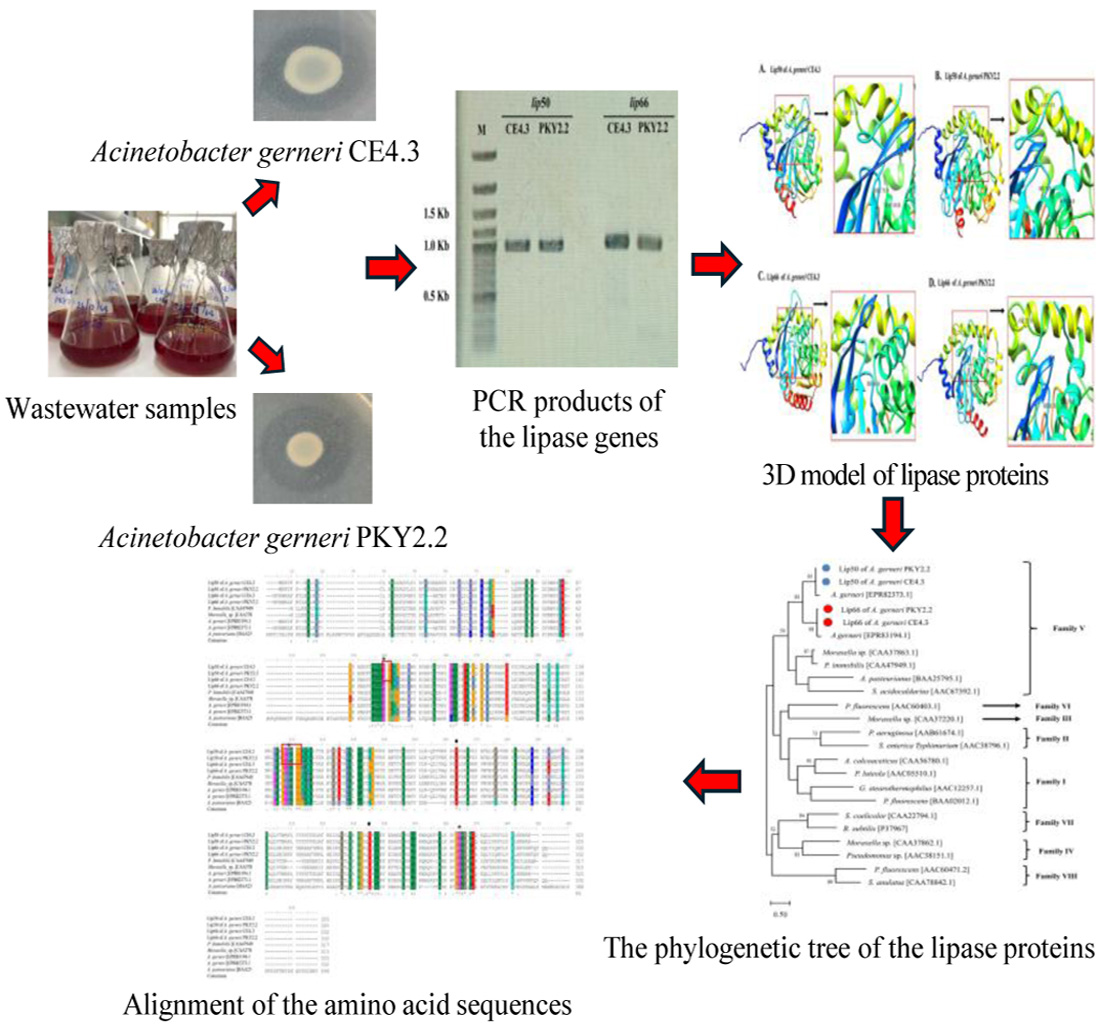
Downloads
References
B Devaraj and G Rajalakshmi. Microbial lipases: An overview of screening, production and purification. Biocatalysis and Agricultural Biotechnology 2019; 22, 101368.
N Patel, D Rai, Shivam, S Shahane and U Mishra. Lipases: Sources, production, purification, and applications. Recent Patents on Biotechnology 2019; 13(1), 45-56.
CA Salgado, CIAD Santos and MCD Vanetti. Microbial lipases: Propitious biocatalysts for the food industry. Food Bioscience 2022; 45, 101509.
T Panyachanakul, V Kitpreechavanich, W Lorliam, T Lomthong and S Krajangsang. Optimization, purification and characterization of lipase from Streptomyces sp. A3301, with application of crude lipase for cooking oily wastewater treatment. Trends in Sciences 2024; 21(11), 8504.
Y Jha. Chapter 11-Bacterial lipases: Biotechnological and industrial applications. Bacterial Secondary Metabolites Synthesis and Applications in Agroecosystem Nanobiotechnology for Plant Protection 2024; 1, 197-209.
P Farrokh, B Yakhchali and AA Karkhane. Cloning and characterization of newly isolated lipase from Enterobacter sp. Bn12. Brazilian Journal of Microbiology 2014; 45(2), 677-687.
VP Sutar, GV Mali, V Upadhye, VK Singh and RP Sinha. Purification of lipase from Pseudomonas aeruginosa VSJK R-9 and its application in combination with the lipolytic consortium for bioremediation of restaurant wastewater. Applied Biochemistry and Biotechnology 2023; 195, 1888-903.
P Ungcharoenwiwat and AH-Kittikun. Purification and characterization of lipase from Burkholderia sp. EQ3 isolated from wastewater from a canned fish factory and its application for the synthesis of wax esters. Journal of Molecular Catalysis B: Enzymatic 2015; 115, 96-104.
T Saengsanga, W Siripornadulsil and S Siripornadulsil. Molecular and enzymatic characterization of alkaline lipase from Bacillus amyloliquefaciens E1PA isolated from lipid-rich food waste. Enzyme and Microbial Technology 2016; 82, 23-33.
G Sukplang, C Narasettiti, O Chunhachart, M Srithaworn, W Lorliam, S Krajangsang and T Panyachanakul. Isolation and identification of lipase-producing bacteria from wastewater treatment pond for the removal of oil and grease in wastewater treatment. Chiang Mai Journal of Science 2024; 51(6), 2024096.
V Sakpuntoon, J Angchuan, W Boontham, P Khunnamwong, C Boonmak and N Srisuk. Grease waste as a reservoir of lipase-producing yeast and description of Limtongella siamensis gen. nov., sp. nov. Microorganisms 2020; 8(1), 27.
K Suwansunthichai, V Kitpreechavanich, L Kraidej and L Chitradon. Rapid method for screening of high lipase producing bacteria. Agriculture and Natural Resources 1991; 25(2), 162-168.
R Noormohamadi, F Tabandeh, P Shariati and M Otadi. Characterization of a lipase from a newly isolated Pseudomonas sp. Iranian Journal of Microbiology 2013; 5(4), 422-427.
A Alawiaha, F Al-Sayied and O El-Hamshary. Detection and molecular characterization of lipase-producing bacteria. Egyptian Pharmaceutical Journal 2023; 22(1), 54-66.
K Wilson. Preparation of genomic DNA from bacteria. Current Protocols in Molecular Biology 2001. https://doi.org/10.1002/0471142727.mb0204s56
LA Kelley, S Mezulis, CM Yates, MN Wass and MJ Sternberg. The Phyre2 web portal for protein modeling, prediction and analysis. Nature Protocols 2015; 10, 845-58.
EF Pettersen, TD Goddard, CC Huang, GS Couch, DM Greenblatt, EC Meng and TE Ferrin. UCSF Chimera--a visualization system for exploratory research and analysis. Journal of Computational Chemistry 2004; 25, 1605-1612.
S Kumar, G Stecher, M Li, C Knyaz and K Tamura. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution 2018; 36(6), 1547-1549.
X Zheng, X Chu, W Zhang, N Wu and Y Fan. A novel cold-adapted lipase from Acinetobacter sp. XMZ-26: Gene cloning and characterisation. Applied Microbiology and Biotechnology 2011; 90, 971-980.
P Anbu, MJ Noh, DH Kim, JS Seo, BK Hur and KH Min. Screening and optimization of extracellular lipases by Acinetobacter species isolated from oil-contaminated soil in South Korea. African Journal of Biotechnology 2011; 10(20), 4147-4156.
K Batumalaie, NA Mahat, MF Edbeib, FZ Huyop and RA Wahab. Characterization of a novel alkaline-stable lipase from Acinetobacter haemolyticus KV1 isolated from an oil palm mill effluent. Malaysian Journal of Microbiology 2018; 14, 640-654.
VD Nimkande, S Sivanesan and A Bafana. Screening, identification, and characterization of lipase-producing halotolerant Bacillus altitudinis Ant19 from Antarctic soil. Archives of Microbiology 2023; 205, 113.
N Lenfant, T Hotelier, E Velluet, Y Bourne, P Marchot and A Chatonnet. ESTHER, the database of the alpha/beta-hydrolase fold superfamily of proteins: tools to explore diversity of functions. Nucleic Acids Research 2012; 41, 423-429.
G Furini, JS Berger, JJM Campos, STV Sand and JC Germani. Production of lipolytic enzymes by bacteria isolated from biological effluent treatment systems. Anais da Academia Brasileira de Ciências 2018; 90, 2955-65.
J Veno, R Rahman, M Masomian, MSM Ali and NHA Kamarudin. Insight into improved thermostability of cold-adapted Staphylococcal lipase by glycine to cysteine mutation. Molecules 2019; 24(17), 3169.
C Guo, R Zheng, R Cai, C Sun and S Wu. Characterization of two unique cold-active lipases derived from a novel deep-sea cold seep bacterium. Microorganisms 2021; 9(4), 802.
H Wang, H Zhong, C Gao, J Zang and D Yang. The distinct properties of the consecutive disordered regions inside or outside protein domains and their functional significance. International Journal of Molecular Sciences 2021; 22(19), 10677.
JT Mindrebo, CM Nartey, Y Seto, MD Burkart and JP Noel. Unveiling the functional diversity of the alpha/beta hydrolase superfamily in the plant kingdom. Current Opinion in Structural Biology 2016; 41, 233-246.
AR Buller and CA Townsend. Intrinsic evolutionary constraints on protease structure, enzyme acylation, and the identity of the catalytic triad. PNAS 2013; 110(8), 653-661.
A Kumari and R Gupta. Phenylalanine to leucine point mutation in oxyanion hole improved catalytic efficiency of Lip12 from Yarrowia lipolytica. Enzyme and Microbial Technology 2013; 53(6-7), 386-390.
A Sahu and R Birner-Gruenberger. Encyclopedia of metalloproteins. In: RH Kretsinger, VN Uversky and EA Permyakov (Eds.). Springer, New York, 2013.
JL Arpigny and KE Jaeger. Bacterial lipolytic enzymes: Classification and properties. Biochemical Journal 1999; 343(1), 177-183.
SH Albayati, M Masomian, SNH Ishak, MSBM Ali, AL Thean, FBM Shariff, NDBM Noor and RNZRA Rahman. Main structural targets for engineering lipase substrate specificity. Catalysts 2020; 10(7), 747.
J Yang, K Kobayashi, Y Iwasaki, H Nakano and T Yamane. In vitro analysis of roles of a disulfide bridge and a calcium binding site in activation of Pseudomonas sp. strain KWI-56 lipase. American Society for Microbiology 2000; 182(2), 295-302.
H Nacke, C Will, S Herzog, B Nowka, M Engelhaupt and R Daniel. Identification of novel lipolytic genes and gene families by screening of metagenomic libraries derived from soil samples of the German biodiversity exploratories. FEMS Microbiology Ecology 2011; 78(1), 188-201.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






