Genomic Characterization of mcr-1-Carrying E. coli Isolated from Food Handlers
DOI:
https://doi.org/10.48048/tis.2025.10562Keywords:
mcr-1, Colistin-resistant E. coli, Food handlers, Asymptomatic carriage, Complete genome sequencing, One HealthAbstract
The emergence of colistin-resistant Escherichai coli harboring the mcr-1 gene poses a significant public health concern, particularly due to colistin’s status as a last-resort antibiotic against multidrug-resistant Gram-negative infections. We provide the first identification of mcr-1-positive E. coli that was isolated from the hands of asymptomatic northern Thai restaurant employees. Twenty E. coli isolates were found from 150 hand swab samples; strain EC16 was shown to be colistin-resistant and to carry the mcr-1 gene. By hybrid-based complete genome sequencing, a 4.76 Mb chromosome and 2 plasmids (2 and 239 kb) were identified, the latter carrying multiple resistance genes including mcr-1. EC16 was shown to be clustered within ST1079 by core genome MLST and pangenome analysis, and it is closely connected to strains that are associated with humans and the environment. A varied resistome and virulome, comprising genes for adhesion and antibiotic inactivation, were revealed by LS-BSR analysis. pEcoEC16b was determined by plasmid typing to be an IncHI1A plasmid that is closely linked to p7PM1-IncHI1-mcr-1. Comparative study verified structural commonalities and shared resistance genes (mcr-1, tetA and cmlA1). These results underline the significance of One Health surveillance by demonstrating the potential of IncHI1A plasmids to spread multidrug resistance across species and environments.
HIGHLIGHTS
- First report of mcr-1 positive E. coli isolated from the hands of restaurant workers
- New insights into potential human-associated reservoirs of colistin resistance.
- Our study highlights a new route of transmission via the hands of food handlers.
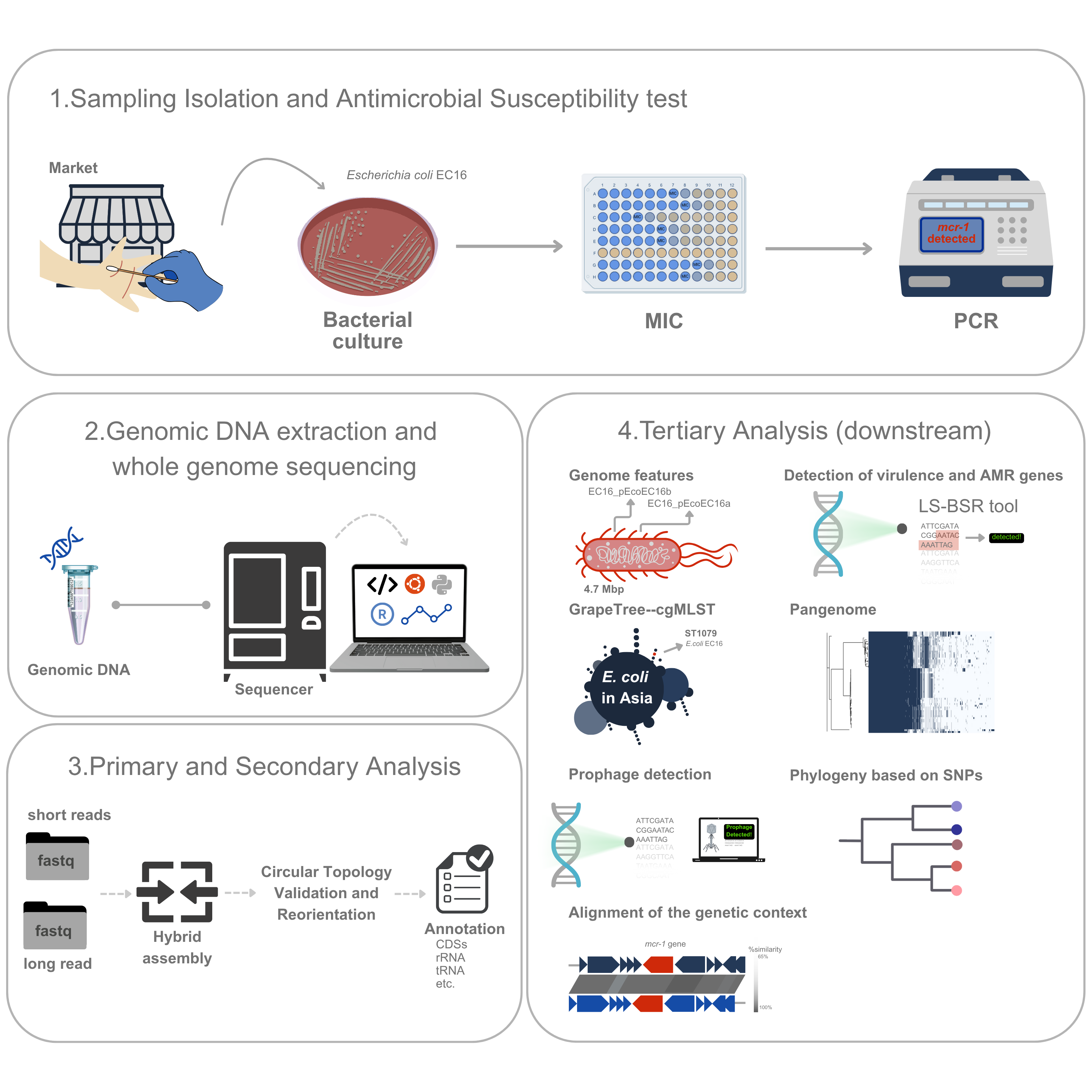
GRAPHICAL ABSTRACT
Downloads
References
GBD 2021 Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance 1990-2021: A systematic analysis with forecasts to 2050. The Lancet 2024; 404, 1199-1226.
S Chautrakarn, W Khumros and P Phutrakool. Self-medication with over-the-counter medicines among the working age population in metropolitan areas of Thailand. Frontiers in Pharmacology 2021; 12, 726643.
FA Lencina, M Bertona, MA Stegmayer, CR Olivero, LS Frizzo, JA Zimmermann, ML Signorini, LP Soto and MV Zbrun. Prevalence of colistin-resistant Escherichia coli in foods and food-producing animals through the food chain: A worldwide systematic review and meta-analysis. Heliyon 2024; 10(5), e26579.
FT Johura, J Tasnim, I Barman, SR Biswas, FT Jubyda, M Sultana, CM George, A Camilli, KD Seed, N Ahmed and M Alam. Colistin-resistant Escherichia coli carrying mcr-1 in food, water, hand rinse, and healthy human gut in Bangladesh. Gut Pathogens 2020; 12, 5.
P Chopjitt, P Boueroy, M Morita, T Iida, Y Akeda, S Hamada and A Kerdsin. Genetic characterization of multidrug-resistant Escherichia coli harboring colistin-resistant gene isolated from food animals in food supply chain. Frontiers in Cellular and Infection Microbiology 2024; 14, 1289134.
A Khanawapee, A Kerdsin, P Chopjitt, P Boueroy, R Hatrongjit, Y Akeda, K Tomono, S Nuanualsuwan and S Hamada. Distribution and molecular characterization of Escherichia coli harboring mcr genes isolated from slaughtered pigs in Thailand. Microb. Microbial Drug Resistance 2021; 27(7), 971-979.
N Thadtapong, S Chaturongakul, S Tangphatsornruang, C Sonthirod, N Ngamwongsatit and R Aunpad. Four new sequence types and molecular characteristics of multidrug-resistant Escherichia coli strains from foods in Thailand. Antibiotics 2024; 13(10), 935.
PC Wu, MF Cheng, WL Chen, WY Hung, JL Wang and CH Hung. Risk factors and prevalence of mcr-1-positive Escherichia coli in fecal carriages among community children in Southern Taiwan. Frontiers in Microbiology 2021; 12, 748525.
HM Martiny, P Munk, C Brinch, J Szarvas, FM Aarestrup and TN Petersen. Global distribution of mcr gene variants in 214K metagenomic samples. mSystems 2022; 7(2), e00105-22.
P Sismova, I Sukkar, N Kolidentsev, J Palkovicova, I Chytilova, J Bardon, M Dolejska and K Nesporova. Plasmid-mediated colistin resistance from fresh meat and slaughtered animals in the Czech Republic: nation-wide surveillance 2020-2021. Microbiology Spectrum 2023; 11(5), e0060923.
Z Lv, Y Shen, W Liu, H Ye, D Liu, J Liu, Y Fu, C Peng, K Chen, X Deng, B Liu, J He, L Yang, C Xu, C Cai, Y Wang, Y Ke and J Shen. Prevalence and risk factors of mcr-1-positive volunteers after colistin banning as animal growth promoter in China: A community-based case-control study. Clinical Microbiology and Infection 2022; 28(2), 267-272.
B Tang, J Wang, X Zheng, J Chang, J Ma, J Wang, X Ji, H Yang and B Ding. Antimicrobial resistance surveillance of Escherichia coli from chickens in the Qinghai Plateau of China. Frontiers in Microbiology 2022; 13, 885132.
H Liao, C Liu, S Zhou, C Liu, DJ Eldridge, C Ai, SW Wilhelm, BK Singh, X Liang, M Radosevich, Q Yang, X Tang, Z Wei, VP Friman, M Gillings, MD Baquerizo and YG Zhu. Prophage-encoded antibiotic resistance genes are enriched in human-impacted environments. Nature Communications 2024; 15(1), 8315.
C Shen, LL Zhong, Y Yang, Y Doi, DL Paterson, N Stoesser, F Ma, MAE El-Sayed Ahmed, S Feng, S Huang, HY Li, X Huang, X Wen, Z Zhao, M Lin, G Chen, W Liang, Y Liang, Y Xia, …, GB Tian. Dynamics of mcr-1 prevalence and mcr-1-positive Escherichia coli after the cessation of colistin use as a feed additive for animals in China: A prospective cross sectional and whole genome sequencing-based molecular epidemiological study. The Lancet Microbe 2020; 1(1), e34-e43.
T An, Y Cai, G Li, S Li, PK Wong, J Guo and H Zhao. Prevalence and transmission risk of colistin and multidrug resistance in long-distance coastal aquaculture. ISME Communications 2023; 3(1), 115.
CH Guo, YQ Liu, Y Li, XX Duan, TY Yang, FY Li, M Zou and BT Liu. High prevalence and genomic characteristics of carbapenem-resistant Enterobacteriaceae and colistin-resistant Enterobacteriaceae from large-scale rivers in China. Environmental Pollution 2023; 331, 121869.
Department of Medical Science. Microbiological quality criteria for food and food contact containers issued by the Department of Medical Science, Ministry of Public Health, Thailand. P2 Design & Print, Bangkok, Thailand, 2017.
AZ Tanzin, C Nath, MRK Nayem, MA Sayeed, SA Khan, RS Magalhaes, JI Alawneh and MM Hassan. Detection and characterisation of colistin-resistant escherichia coli in broiler meats. Microorganisms 2024; 12(12), 2535.
S Andrews. FastQC: A quality control tool for high throughput sequence data. Analytical Biochemistry 2010; 548, 38-43.
P Ewels, M Magnusson, S Lundin and M Käller. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016; 32(19), 3047-3048.
AM Bolger, M Lohse and B Usadel. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014; 30(15), 2114-2120.
W De Coster, S D’Hert, DT Schultz, M Cruts and C Van Broeckhoven. NanoPack: Visualizing and processing long-read sequencing data. Bioinformatics 2018; 34(15), 2666-2669.
RR Wick. Filtlong, Available at: https://github.com/rrwick/Filtlong, accessed January 2025.
RR Wick, LM Judd, CL Gorrie and KE Holt. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Computational Biology 2017; 13(6), e1005595.
M Hunt, ND Silva, TD Otto, J Parkhill, JA Keane and SR Harris. Circlator: Automated circularization of genome assemblies using long sequencing reads. Genome Biology 2015; 16, 294.
RR Wick, MB Schultz, J Zobel and KE Holt. Bandage: Interactive visualization of de novo genome assemblies. Bioinformatics 2015; 31(20), 3350-3352.
T Carver, SR Harris, M Berriman, J Parkhill and JA McQuillan. Artemis: An integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 2012; 28(4), 464-469.
T Seemann. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014; 30(14), 2068-2069.
Z Zhou, NF Alikhan, MJ Sergeant, N Luhmann, C Vaz, AP Francisco, JA Carriço and M Achtman. GrapeTree: Visualization of core genomic relationships among 100,000 bacterial pathogens. Genome Research 2018; 28(9), 1395-1404.
AJ Page, CA Cummins, M Hunt, VK Wong, S Reuter, MT Holden, M Fookes, D Falush, JA Keane and J Parkhill. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015; 31(22), 3691-3693.
JW Sahl, JG Caporaso, DA Rasko and P Keim. The large-scale blast score ratio (LS-BSR) pipeline: A method to rapidly compare genetic content between bacterial genomes. PeerJ 2014; 2, e332.
CE Chandler, AM Horspool, PJ Hill, DJ Wozniak, JW Schertzer, DA Rasko and RK Ernst. Genomic and phenotypic diversity among 10 laboratory isolates of Pseudomonas aeruginosa PAO1. Journal of Bacteriology 2019; 201(5), 10-1128.
DS Wishart, S Han, S Saha, E Oler, H Peters, JR Grant, P Stothard and V Gautam. PHASTEST: Faster than PHASTER, better than PHAST. Nucleic Acids Research 2023; 51(W1), W443-W450.
F Bertels, OK Silander, M Pachkov, PB Rainey and E Van Nimwegen. Automated reconstruction of whole-genome phylogenies from short-sequence reads. Molecular Biology and Evolution 2014; 31(5), 1077-1088.
I Letunic and P Bork. Interactive Tree of Life (iTOL) v6: Recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Research 2024; 52(W1), W78-W82.
MJ Sullivan, NK Petty and SA Beatson. Easyfig: A genome comparison visualizer. Bioinformatics 2011; 27(7), 1009-1010.
Y Li, Y Zhang, X Sun, Y Wu, Z Yan, X Ju, Y Huang, H Zhou, Z Wang, S Wang, R Zhang and R Li. National genomic epidemiology investigation revealed the spread of carbapenem-resistant Escherichia coli in healthy populations and the impact on public health. Genome Medicine 2024; 16(1), 57.
AR Manges, HM Geum, A Guo, TJ Edens, CD Fibke and JDD Pitout. Global extraintestinal pathogenic Escherichia coli (ExPEC) lineages. Clinical Microbiology Reviews 2019; 32(3), 10-1128.
M Touchon, A Perrin, JAM de Sousa, B Vangchhia, S Burn, CL O’Brien, E Denamur, D Gordon and EP Rocha. Phylogenetic background and habitat drive the genetic diversification of Escherichia coli. PLoS Genetics 2020; 16(6), e1008866.
SM Chauhan, O Ardalani, JC Hyun, JM Monk, PV Phaneuf and BO Palsson. Decomposition of the pangenome matrix reveals a structure in gene distribution in the Escherichia coli species. mSphere 2025; 10(1), e00532-24.
HL Her, PT Lin and YW Wu. PangenomeNet: A pan-genome-based network reveals functional modules on antimicrobial resistome for Escherichia coli strains. BMC Bioinformatics 2021; 22, 548.
VS Braz, K Melchior and CG Moreira. Escherichia coli as a multifaceted pathogenic and versatile bacterium. Frontiers in Cellular and Infection Microbiology 2020; 10, 548492.
M Zhuang, Y Achmon, Y Cao, X Liang, L Chen, H Wang, BA Siame and KY Leung. Distribution of antibiotic resistance genes in the environment. Environmental Pollution 2021; 285, 117402.
S Sher, HA Khan, Z Khan, MS Siddique, DA Bukhari and A Rehman. Bacteriophages: Potential candidates for the dissemination of antibiotic resistance genes in the environment. Targets 2025; 3(3), 25.
J Feng, Z Xu, Y Zhuang, M Liu, J Luo, Y Wu, Y Chen and M Chen. The prevalence, diagnosis, and dissemination of mcr-1 in colistin resistance: Progress and challenge. Decoding Infection and Transmission 2023; 1, 100007.
K Zurfluh, J Klumpp, M Nüesch-Inderbinen and R Stephan. Full-length nucleotide sequences of mcr-1-harboring plasmids isolated from extended-spectrum-β-lactamase-producing Escherichia coli isolates of different origins. Antimicrobial Agents and Chemotherapy 2016; 60(9), 5589-5591.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






