A Mediated-Approach to Disrupting SpeA-TCR Interactions for the Treatment of Streptococcus pyogenes Infection
DOI:
https://doi.org/10.48048/tis.2025.10393Keywords:
Docking, gRNA, SpeA, Streptococccus, TCR, Docking, gRNA, SpeA, Streptococcus, TCRAbstract
Streptococcus pyrogenic exotoxin A (SpeA) has been implicated in several S. pyogenes illnesses, including invasive and non-invasive infections. The main investigation of this study explores methods to use gRNA SpeA-Cas9 both to disrupt the SpeA-TCR interaction and decrease S. pyogenes pathogenicity. This research aims to develop and verify protective gRNA sequences as well as conduct analyzes of their structural characteristics and thermal behavior while assessing potential therapeutic possibilities. The gRNA SpeA sequence, 2D and 3D structures were predicted using the CRISPRDB, SimRNA 2.0, and RNAfold services, respectively. Geneious Prime was utilized to finish the PCR and clone gRNA-Cas9 into pCRII-TOPO. FUpred was utilized to detect the boundary domains. CGR approaches were used to map a part of the target sequence of gRNA SpeA over the whole S. pyogenes genome. The HDOCK server was used to dock SpeA and TCR molecules. EcoRI is the most suitable restriction enzyme for gRNA SpeA-Cas9 transformation. The target sequence (ACGGAGG) was determined in the A20 strain complete genome map. The optimum biochemical and biophysical properties of gRNA SpeA-Cas9 were estimated. Model 2 is appropriate for positioning the SpeA gRNA between the docking proteins (SpeA-TCR). S. pyogenes loses its virulence and its capacity to induce infection when SpeA begins to attach less strongly to host cell receptors. An effective path toward innovative treatment options is provided by the gRNA-based strategy. There is a chance that gRNA will change how much SpeA and TCR interact molecularly. This study shows CRISPR-Cas9 targeted against SpeA results in a reduction of SpeA’s specificity for TCR receptors which successfully controls immune cell overreactivity. Research reveals that targeting SpeA with CRISPR-Cas9 represents an appealing therapeutic solution to fight S. pyogenes infections as well as decrease their dangerous outcomes.
HIGHLIGHTS
SpeA has been implicated in a range of S. pyogenes diseases. SpeA is a secreted polypeptide, that identifies antigens that escape the immune system. Geneious Prime was utilized to finish the PCR and clone gRNA-Cas9 into pCRII-TOPO. The target sequence (ACGGAGG) was determined in the A20 strain complete genome map.
GRAPHICAL ABSTRACT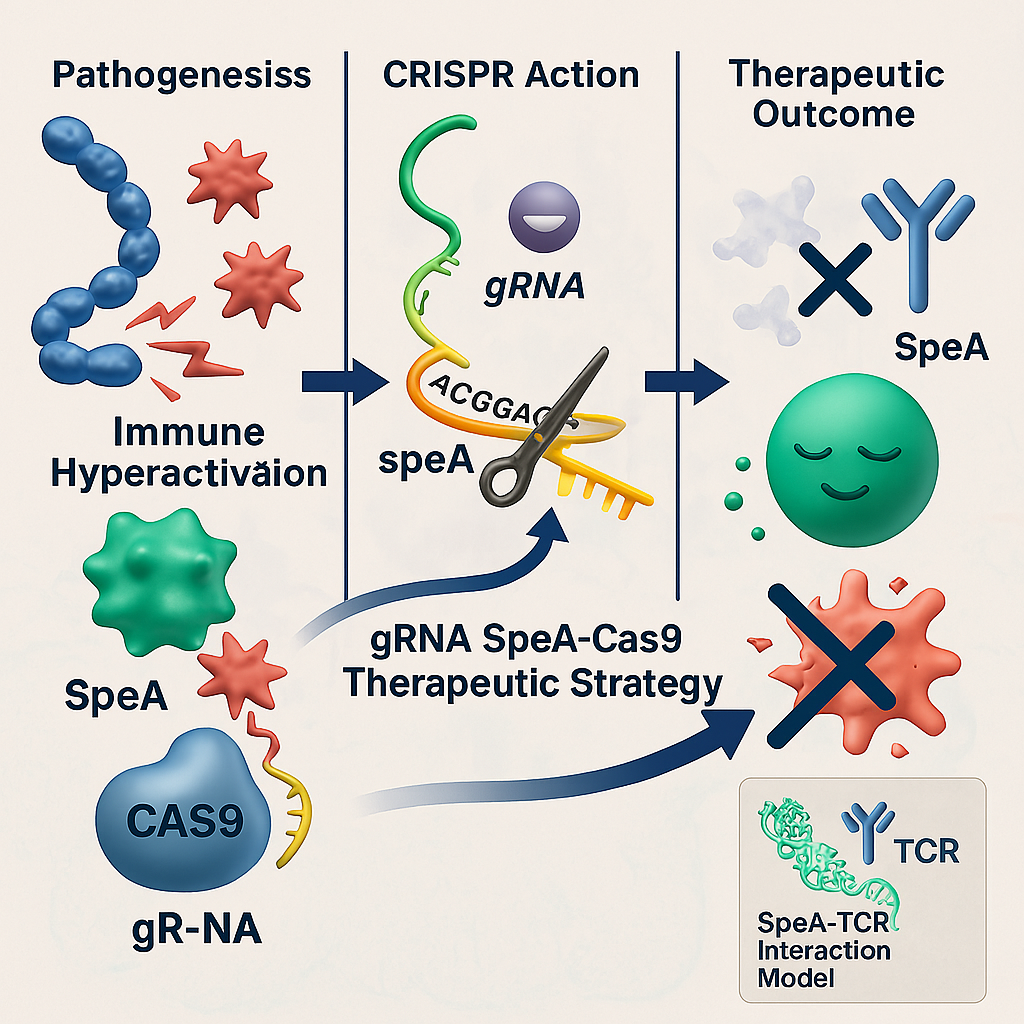
Downloads
References
CC Dominguez-Medina, NL Rash, S Robillard, C Robinson, A Efstratiou, K Broughton, J Parkhill, MTG Holden, MR Lopez-Alvarez, R Paillot and AS Waller. A novel superantigen and its potential as a vaccine adjuvant against strangles. International Journal of Molecular Sciences 2020; 21(12), 4467.
BA Shannon, JK McCormick and PM Schlievert. Toxins and superantigens of group a streptococci. Microbiology Spectrum 2019; 7(1), 10-1128.
C Olive, K Hsien, A Horvath, T Clair, P Yarwood, I Toth and MF Good. Protection against group A streptococcal infection by vaccination with self-adjuvanting lipid core M protein peptides. Vaccine 2005; 23(17-18), 2298-2303.
RG Ulrich. Vaccine based on a ubiquitous cysteinyl protease and streptococcal pyrogenic exotoxin A protects against Streptococcus pyogenes sepsis and toxic shock. Journal of Immune Based Therapies and Vaccines 2008; 31(6), 8.
SF Lee, L Li, N Jalal and SA Halperin. Identification of a thiol-disulfide oxidoreductase (SdbA) catalyzing disulfide bond formation in the superantigen SpeA in Streptococcus pyogenes. Journal of Bacteriology 2021; 203(17), e0015321.
A Popugailo, Z Rotfogel, E Supper, D Hillman and R Kaempfer. Staphylococcal and streptococcal superantigens trigger B7/CD28 costimulatory receptor engagement to hyperinduce inflammatory cytokines. Frontiers in Immunology 2019; 10, 942.
FJ Radcliff, F Clow, M Mahadevan, J Johnston, T Proft, RG Douglas and JD Fraser. A potential role for staphylococcal and streptococcal superantigens in driving skewing of TCR VB subsets in tonsillar hyperplasia. Medical Microbiology and Immunology 2017; 206(4), 337-346.
Y You, MR Davies, M Protani, L McIntyre, MJ Walker and J Zhang. Scarlet fever epidemic in China caused by Streptococcus pyogenes serotype M12: Epidemiologic and molecular analysis. EBioMedicine 2018; 28, 128-135.
F Hille, H Richter, SP Wong, M Bratovič, S Ressel and E Charpentier. The biology of CRISPR-Cas: Backward and forward. Cell 2018; 172(6), 1239-1259.
YS Dagdas, JS Chen, SH Sternberg, JA Doudna and A Yildiz. A conformational checkpoint between DNA binding and cleavage by CRISPR-Cas9. Science Advances 2017; 3(8), eaao0027.
R Li, L Fang, S Tan, M Yu, X Li, S He, Y Wei, G Li, J Jiang and M Wu. Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity. Cell Research 2016; 26(12), 1273-1287.
JW Modell, W Jiang and LA Marraffini. CRISPR-Cas systems exploit viral DNA injection to establish and maintain adaptive immunity. Nature 2017; 544(7648), 101-104.
MM Balla, A Mergani, MEAME Medani, AD Abakar and AM Dafalla. Streptococcal pyrogenic exotoxin genes SpeA and SpeB in isolates of Streptococcus pyogenes from children with pharyngitis, Gezira State, Sudan. American Journal of Molecular Biology 2022; 12, 181-189.
AR Spaulding, W Salgado-Pabón, PL Kohler, AR Horswill, DYM Leung and PM Schlievert. Staphylococcal and streptococcal superantigen exotoxins. Clinical Microbiology Reviews 2013; 26(3), 422-447.
M Roggiani, JA Stoehr, SB Olmsted, YV Matsuka, S Pillai, DH Ohlendorf and PM Schlievert. Toxoids of streptococcal pyrogenic exotoxin A are protective in rabbit models of streptococcal toxic shock syndrome. Infection and Immunity 2000; 68(9), 5011-5017.
AA Dawood. Implementation of immuno-chemoinformatics approaches to construct multi-epitope for vaccine development against Omicron and Delta SARS-CoV-2 variants. Vacunas 2022; 23(S2), S18-S31.
V Ozberk, M Pandey and MF Good. Contribution of cryptic epitopes in designing a group A streptococcal vaccine. Human Vaccines & Immunotherapeutics 2018; 14(8), 2034-2052.
M Pandey, A Calcutt, V Ozberk, Z Chen, M Croxen, J Powell, E Langshaw, JL Mills, FE Jen, J McCluskey, J Robson, GJ Tyrrell and MF Good. Antibodies to the conserved region of the M protein and a streptococcal superantigen cooperatively resolve toxic shock-like syndrome in HLA-humanized mice. Science Advances 2019; 5(9), eaax3013.
M Pandey, V Ozberk, EL Langshaw, A Calcutt, J Powell, MR Batzloff, T Rivera-Hernandez and MF Good. Skin infection boosts memory B-cells specific for a cryptic vaccine epitope of group A streptococcus and broadens the immune response to enhance vaccine efficacy. NPJ Vaccines 2018; 3(1), 15.
A Dawood. Influence of SARS-CoV-2 variants’ spike glycoprotein and RNA-dependent RNA polymerase (Nsp12) mutations on remdesivir docking residues. Medical Immunology (Russia) 2022; 24(3), 617-628.
AJ Zaug and TR Cech. Analysis of the structure of Tetrahymena nuclear RNAs in vivo: Telomerase RNA, the self-splicing rRNA intron, and U2 snRNA. RNA 1995; 1(4), 363-374.

Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






