DFT Analysis on Electronic Properties, Reactivity and Adsorption Mechanism of Favipiravir/XTiO4H2 (X = Pt, Zr, Zn) Nanocomplexes and their Biological Evaluations
DOI:
https://doi.org/10.48048/tis.2025.10384Keywords:
Density functional theory, Favipiravir, TiO2 nanocomplexes, Reactivity, Charge transfer, Protein-ligand interactions, Density functional theory, Favipiravir, TiO2 nanocomplexes, Reactivity, Charge transfer, Protein-ligand interactionsAbstract
The objective of this study is to explore quantum chemical properties of favipiravir- loaded XTiO4H2 nanostructures (X = Pt, Zr, Zn) using density functional theory and to evaluate their toxicity profiling, drug-likeness and binding efficacy to the target proteins. Favipiravir interacted strongly with XTiO4H2 forming the stable nanocomplexes (NCs) having adsorption energy in the range of −2.49 to −4.70 eV in gas phase. The HOMO-LUMO energy gap of favipiravir was 3.56 eV whereas Fav + PtTiO4H2 (NC1), Fav + ZrTiO4H2 (NC2), and Fav + ZnTiO4H2 (NC3) exhibited lower energy gaps of 1.23, 1.43 and 0.65 eV in gas phase, respectively, showing better reactivity of these NCs. The results of dipole moment, global reactivity descriptor, MEP surface, NBO, QTAIM and NCI of the proposed NCs were more favorable than that of favipiravir. The functionalized TiO2 NCs exhibited better docking scores, among which NC2 showed the strongest inhibition of SARS-CoV-2 main protease and spike protein. NC3 having the predicted LD50 of 3,000 mg/kg in toxicity class V appears to be less toxic than favipiravir having LD50 of 1,717 mg/kg in the class IV. This study confirms the possibility of using transition metal-doped titania nanostructures for targeted drug delivery, antiviral therapy and photocatalytic degradation of the drugs.
HIGHLIGHTS
- Favipiravir with XTiO4H2 (X = Pt, Zr, Zn) nanocomplexes exhibited significant adsorption energies ranging from -1.18 to -4.70 eV.
- The HOMO-LUMO energy gaps of the nanocomplexes (0.65-1.23 eV) were lower than that of favipiravir (3.56 eV), enhancing electronic reactivity.
- Quantum chemical parameters like dipole moment, global reactivity descriptors, MEP surface, NBO, QTAIM and NCI analyzed the physicochemical properties of the proposed nanocomplexes.
- Fav+ZrTiO4H2 showed the strongest binding score with SARS-CoV-2 main protease and spike protein, while Fav+ZnTiO4H2 exhibited the least toxicity (LD50 = 3000 mg/kg, class V)
GRAPHICAL ABSTRACT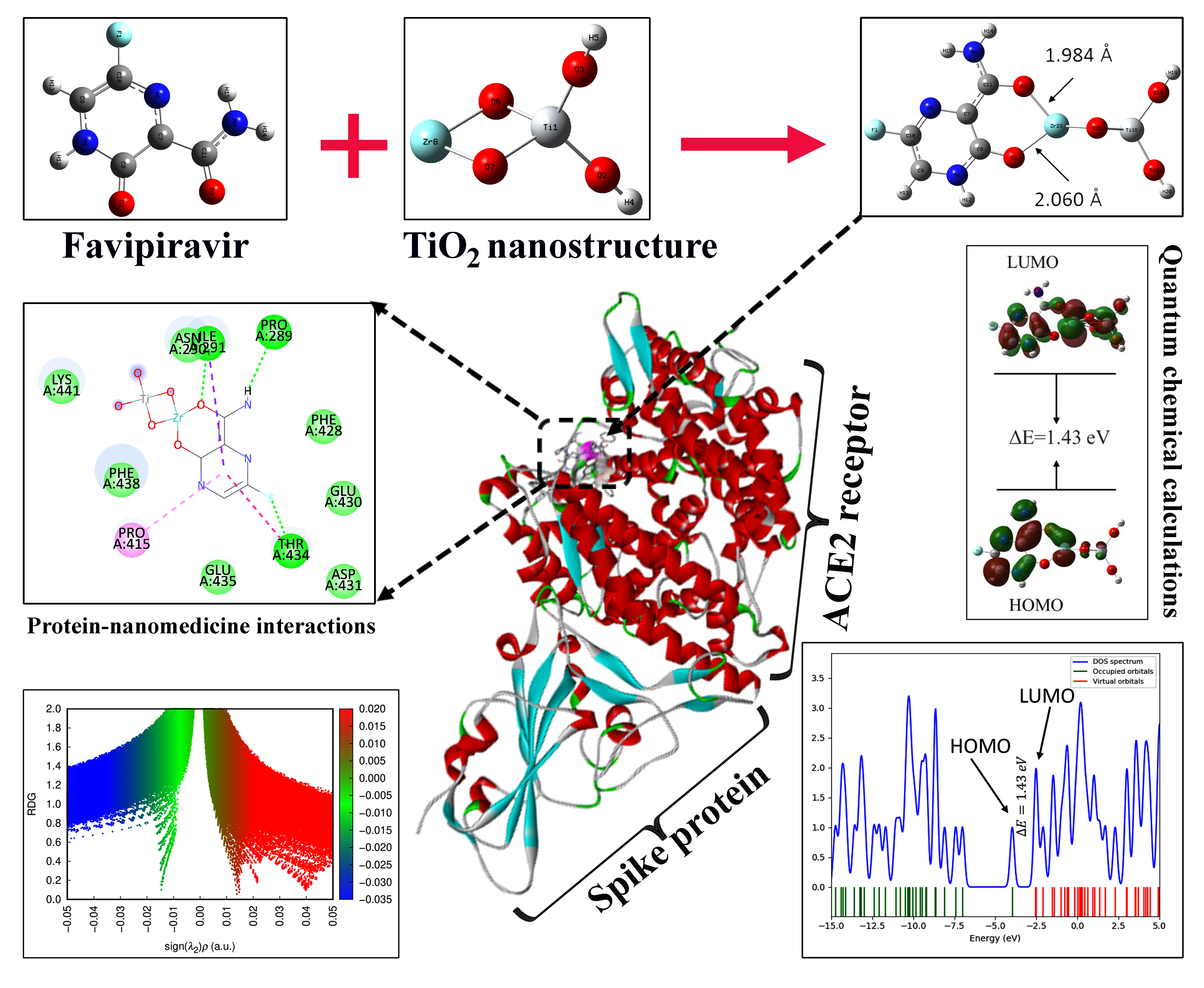
Downloads
References
S Ludwig and A Zarbock. Coronaviruses and SARS-CoV-2: A brief overview. Anesthesia & Analgesia 2020; 131(1), 93-96.
FH Haghighi, M Mercurio, S Cerra, TA Salamone, R Bianymotlagh, C Palocci, VR Spica and I Fratoddi. Surface modification of TiO₂ nanoparticles with organic molecules and their biological applications. Journal of Materials Chemistry B 2023; 11, 2334-2366.
A Ul-Hamid, N Baig, A Haider, AS Hakeem and M Ikram. Using biologically synthesized TiO₂ nanoparticles as potential remedy against multiple drug resistant Staphylococcus aureus of bovine mastitis. Scientific Reports 2023; 13, 18785.
S Chibber and I Ahmad. Molecular docking, a tool to determine interaction of CuO and TiO₂ nanoparticles with human serum albumin. Biochemistry and Biophysics Reports 2016; 6, 63-67.
M Cho, H Chung, W Choi and J Yoon. Different inactivation behaviors of MS-2 phage and Escherichia coli in TiO₂ photocatalytic disinfection. Applied and Environmental Microbiology 2005; 71(1), 270-275.
M Ikram, J Hassan, A Raza, A Haider, S Naz, A Ul-Hamid, J Haider, I Shahzadi, U Qamar and S Ali. Photocatalytic and bactericidal properties and molecular docking analysis of TiO₂ nanoparticles conjugated with Zr for environmental remediation. RSC Advances 2020; 10, 30007-30024.
V Chandrakala, V Aruna and G Angajala. Review on metal nanoparticles as nanocarriers: Current challenges and perspectives in drug delivery systems. Emergent Materials 2022; 5, 1593-1615.
KY Kumar, MK Prashanth, OK Alduaij, TA Yousef, KM Abualnaja and MS Raghu. Mentha arvensis mediated green synthesis of platinum doped TiO₂ nanocomposite for enhanced anticancer and photocatalytic degradation activity: Insights from molecular docking and DFT studies. Inorganic Chemistry Communications 2021; 134, 108987.
G Leon-Gutierrez, JE Elste, C Cabello-Gutierrez, C Millan-Pacheco, MH Martinez-Gomez, R Mejia-Alvare, V Tiwari and A Mejia. A potent virucidal activity of functionalized TiO₂ nanoparticles adsorbed with flavonoids against SARS-CoV-2. Applied Microbiology and Biotechnology 2022; 106(18), 5987-6002.
NA Mazurkova, YE Spitsyna, NV Shikina, ZR Ismagilov, SN Zagrebel’nyi and EI Ryabchikova. Interaction of titanium dioxide nanoparticles with influenza virus. Nanotechnologies in Russia 2010; 5, 417-420.
S Jafari, B Mahyad, H Hashemzadeh, S Janfaza, T Gholikhani and L Tayebi. Biomedical applications of TiO₂ nanostructures: Recent advances. International Journal of Nanomedicine 2020; 15, 3447-3470.
Q Wang, JY Huang, HQ Li, AZJ Zhao, Y Wang, KQ Zhang, HT Sun and YK Lai. Recent advances on smart TiO₂ nanotube platforms for sustainable drug delivery applications. International Journal of Nanomedicine 2016; 2017(12), 151-165.
RH Aguilera-del-Toro, F Aguilera-Granja and EE Vogel. Structural and electronic properties of (TiO₂)₁₀ clusters with impurities: A density functional theory investigation. Journal of Physics and Chemistry of Solids 2019; 135, 109107.
CA Lipinski, F Lombardo, BW Dominy and PJ Feeney. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Advanced Drug Delivery Reviews 1997; 23(1-3), 3-25.
M Frisch, G Trucks, HB Schlegel, GE Scuseria, M Robb, JR Cheeseman, G Scalmani, V Barone, GA Petersson, H Nakatsuji, X Li, M Caricato, A Marenich, J Bloino, B Janesko, R Gomperts, B MENNUCCI, H Hratchian, JV Ortiz, A izmaylov, ..., DJ Fox. Gaussian 16, revision A.03. Gaussian, Inc., Wallingford, 2016.
AD Becke. Density-functional exchange-energy approximation with correct asymptotic behavior. Physical Review A 1988; 38, 3098.
C Lee, W Yang and RG Parr. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Physical Review B 1988; 37, 785.
PJ Hay and WR Wadt. Ab initio effective core potentials for molecular calculations. Potentials for the transition metal atoms Sc to Hg. The Journal of Chemical Physics 1985; 82(1), 270-283.
R Dennington, TA Keith and JM Millam. GaussView, version 6.0.16. Semichem, Inc., Shawnee Mission, 2016.
NM O’Boyle, AL Tenderholt and KM Langner. Cclib: A library for package-independent computational chemistry algorithms. Journal of Computational Chemistry 2008; 29(5), 839-845.
ED Glendening, AE Reed, JE Carpenter and F Weinhold. NBO 3.0 program manual. CCL, Florida, 1990.
T Lu and F Chen. Multiwfn: A multifunctional wavefunction analyzer. Journal of Computational Chemistry 2012; 33(5), 580-592.
Z Jin, X Du, Y Xu, Y Deng, M Liu, Y Zhao, B Zhang, X Li, L Zhang, C Peng, Y Duan, J Yu, L Wang, K Yang, F Liu, R Jiang, X Yang, T You, X Liu, X Yang, ..., H Yang. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 2020; 582(7811), 289-293.
J Lan, J Ge, J Yu, S Shan, H Zhou, S Fan, Q Zhang, X Shi, Q Wang, L Zhang and X Wang. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020; 581(7807), 215-220.
PW Rose, A Prlic, A Altunkaya, C Bi, AR Bradley, CH Christie, LD Costanzo, JM Duarte, S Dutta, Z Feng, RK Green, DS Goodsell, B Hudson, T Kalro, R Lowe, E Peisach, C Randle, AS Rose, C Shao, YP Tao, ..., SK Burley. The RCSB protein data bank: Integrative view of protein, gene and 3D structural information. Nucleic Acids Research 2017; 45(1), 271-281.
A Douangamath, D Fearon, P Gehrtz, T Krojer, P Lukacik, CD Owen, E Resnick, C Strain-Damerell, A Aimon, P Abranyi-Balogh, J Brandão-Neto, A Carbery, G Davison, A Dias, TD Downes, L Dunnett, M Fairhead, JD Firth, SP Jones, A Keeley, GM Keserü, ..., MA Walsh. Crystallographic and electrophilic fragment screening of the SARS-CoV-2 main protease. Nature Communications 2020; 11, 5047.
J Medina-Barandica, N Contreras-Puentes, A Taron-Dunoyer, M Duran-Lengua, A Alviz-Amador. In-silico study for the identification of potential destabilizers between the spike protein of SARS-CoV-2 and human ACE-2. Informatics in Medicine Unlocked 2023; 40, 101278.
A Waterhouse, M Bertoni, S Bienert, G Studer, G Tauriello, R Gumienny, FT Heer, TAP de Beer, C Rempfer, L Bordoli, R Lepore and T Schwede. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Research 2018; 46(1), 296-303.
GM Morris, R Huey, W Lindstrom, MF Sanner, RK Belew, DS Goodsell and AJ Olson. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of Computational Chemistry 2009; 30(16), 2785-2791.
Dassault Systèmes. Discovery studio modeling environment. Dassault Systèmes, France, 2017.
R Maharjan, K Gyawali, A Acharya, M Khanal, MP Ghimire and TR Lamichhane. Artemisinin derivatives as potential drug candidates against Mycobacterium tuberculosis: Insights from molecular docking, MD simulations, PCA, MM/GBSA, and ADMET analysis. Molecular Simulation 2024; 50(11), 717-728.
A Acharya, M Khanal, R Maharjan, K Gyawali, BR Luitel, R Adhikari, DD Mulmi, TR Lamichhane and HP Lamichhane. Quantum chemical calculations on calcium oxalate and dolichin A and their binding efficacy to lactoferrin: An in silico study using DFT, molecular docking, and molecular dynamics simulations. AIMS Biophys. 2024; 11(2), 142-165.
A Daina, O Michielin and V Zoete. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Scientific Reports 2017; 7, 42717.
P Banerjee, AO Eckert, AK Schrey and R Preissner. ProTox-II: A webserver for the prediction of toxicity of chemicals. Nucleic Acids Research 2018; 46, 257-263.
R Meyer and AW Hauser. Geometry optimization using Gaussian process regression in internal coordinate systems. Journal of Chemical Physics 2020; 152, 084112.
O Noureddine, N Issaoui and OD Al-Dossary. DFT and molecular docking study of chloroquine derivatives as antiviral to coronavirus COVID-19. Journal of King Saud University - Science 2021; 33(1), 101248.
N Issaoui, H Ghalla, S Muthu, H Flakus and B Oujia. Molecular structure, vibrational spectra, AIM, HOMO–LUMO, NBO, UV, first order hyperpolarizability, analysis of 3-thiophenecarboxylic acid monomer and dimer by Hartree–Fock and density functional theory. Spectrochim. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy 2015; 136, 1227-1242.
B Borah and TG Devi. Characterization of Zn (L-Proline) 2 complex using spectroscopic techniques and DFT analysis. Journal of Molecular Structure 2020; 1210, 128022.
E Barim and F Akman. Synthesis, characterization and spectroscopic investigation of N-(2-acetylbenzofuran-3-yl) acrylamide monomer: Molecular structure, HOMO–LUMO study, TD-DFT and MEP analysis. Journal of Molecular Structure 2019; 1195, 506-513.
P Chattaraj, S Nath and B Maiti. Reactivity descriptors. Computer-Aided Drug Design and Drug Discovery 2003; 11, 295.
RG Pearson. Absolute electronegativity and hardness: Applications to organic chemistry. The Journal of Organic Chemistry 1989; 54(6), 1423-1430.
N GF, V Vetrivelan and M Prasath. Enhancing therapeutic efficacy and safety profile of Favipiravir through noble metal loaded silica nanocomposites: A quantum chemical and molecular docking study for COVID-19 treatment. Journal of Organometallic Chemistry 2024; 1017, 123291.
T Koopmans. Über die Zuordnung von Wellenfunktionen und Eigenwerten zu den einzelnen Elektronen eines Atoms. Physica 1934; 1(1-6), 104-113.
GF Nivetha, V Vetrivelan, S Muthu and M Prasath. Adsorption behavior, different green solvent effect and surface enhanced Raman spectra (SERS) investigation on inhibition of SARS-CoV-2 by antineoplastic drug Carmofur with silver/gold/platinum loaded silica nanocomposites: A combined computational analysis and molecular modelling approach. Results in Chemistry 2023; 6, 101096.
M Miar, A Shiroudi, K Pourshamsian, AR Oliaey and F Hatamjafari. Theoretical investigations on the HOMO–LUMO gap and global reactivity descriptor studies, natural bond orbital, and nucleus-independent chemical shifts analyses of 3-phenylbenzo [d] thiazole-2 (3 H)-imine and its para-substituted derivatives: Solvent and substituent effects. Journal of Chemical Research 2021; 45, 147-158.
P Sjoberg, JS Murray, T Brinck, P Politzer. Average local ionization energies on the molecular surfaces of aromatic systems as guides to chemical reactivity. Canadian Journal of Chemistry 1990; 68(8), 1440-1443.
E Scrocco and J Tomasi. Advances in quantum chemistry. E. Hoggan - Institut Pascal, France, 1978.
R Abel, T Young, R Farid, BJ Berne and RA Friesner. Role of the active-site solvent in the thermodynamics of factor Xa ligand binding. Journal of the American Chemical Society 2008; 130(9), 2817-2831.
H AlRabiah, S Muthu, F Al-Omary, AM Al-Tamimi, M Raja, RR Muhamed and AAR El-Emam. Molecular structure, vibrational spectra, NBO, Fukui function, HOMO-LUMO analysis and molecular docking study of 6-[(2-methylphenyl) sulfanyl]-5 propylpyrimidine-2, 4 (1H, 3H)-dione. Macedonian Journal of Chemistry and Chemical Engineering 2017; 36(1), 59-80.
C Cassidy, JM Halbout, W Donaldson and CL Tang. Nonlinear optical properties of urea. Optics Communications 1979; 29(2), 243-246.
K Fukui. Role of frontier orbitals in chemical reactions. Science 1982; 218(4547), 747-754.
RG Parr and W Yang. Density functional approach to the frontier-electron theory of chemical reactivity. Journal of the American Chemical Society 1984; 106(14), 4049-4050.
M Vimala, SS Mary, R Ramalakshmi and S Muthu. Theoretical description of green solvents effect on electronic property and reactivity of Tert-butyl 4-formylpiperidine-1-carboxylate. Computational and Theoretical Chemistry 2021; 1201, 113255.
C Morell, A Grand and A Toro-Labbe. New dual descriptor for chemical reactivity. The Journal of Physical Chemistry A 2005; 109(1), 205-212.
M Kurt, PC Babu, N Sundaraganesan, M Cinar and M Karabacak. Molecular structure, vibrational, UV and NBO analysis of 4-chloro-7-nitrobenzofurazan by DFT calculations. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy 2011; 79(5), 1162-1170.
ME Lestard, DM Gil, O Estevez-Hernandez, MF Erben and J Duque. Structural, vibrational and electronic characterization of 1-benzyl-3-furoyl-1-phenylthiourea: An experimental and theoretical study. New Journal of Chemistry 2015; 39(9), 7459-7471.
F Weinhold and CR Landis. Valency and bonding: A natural bond orbital donor-acceptor perspective. Cambridge University Press, Cambridge, 2005.
C James, AA Raj, R Reghunathan, V Jayakumar and IH Joe. Structural conformation and vibrational spectroscopic studies of 2,6-bis(p-N,N-dimethyl benzylidene) cyclohexanone using density functional theory. Journal of Raman Spectroscopy 2006; 37(12), 1381-1392.
ER Johnson, S Keinan, P Mori-Sanchez, J Contreras-Garcia, AJ Cohen and W Yang. Revealing noncovalent interactions. Journal of the American Chemical Society 2010; 132(18), 6498-6506.
RA Boto, JP Piquemal and J Contreras-Garcia. Revealing strong interactions with the reduced density gradient: A benchmark for covalent, ionic and charge-shift bonds. Theoretical Chemistry Accounts 2017; 136(139), 1-9.
F Giordanetto, C Tyrchan and J Ulander. Intramolecular hydrogen bond expectations in medicinal chemistry. ACS Medicinal Chemistry Letters 2017; 8(2), 139-142.
I Rozas, I Alkorta and J Elguero. Behavior of ylides containing N, O and C atoms as hydrogen bond acceptors. Journal of the American Chemical Society 2000; 122(45), 11154-11161.
X Wang, J Lan, J Ge, J Yu and S Shan. Crystal structure of SARS-CoV-2 spike receptor binding domain bound with ACE2. Protein Data Bank 2020. https://doi.org/10.2210/pdb6M0J/pdb
M Khanal, A Acharya, R Maharjan, K Gyawali, R Adhikari, DD Mulmi, TR Lamichhane and HP Lamichhane. Identification of potent inhibitors of HDAC2 from herbal products for the treatment of colon cancer: Molecular docking, molecular dynamics simulation, MM/GBSA calculations, DFT studies, and pharmacokinetic analysis. PLOS One 2024; 19(7), 0307501.
K Gyawali, R Maharjan, A Acharya, M Khanal, MP Ghimire and TR Lamichhane. Identification of catechin as main protease inhibitor of SARS-CoV-2 Omicron variant using molecular docking, molecular dynamics, PCA, DCCM, MM/GBSA and ADMET profiling. Natural Product Research 2024; 2, 1-8.
A Ali, GJ Mir, A Ayaz, I Maqbool, SB Ahmad, S Mushtaq, A Khan, TM Mir and MU Rehman. In silico analysis and molecular docking studies of natural compounds of Withania somnifera against bovine NLRP9. Journal of Molecular Modeling 2023; 29, 171.

Additional Files
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






