Evaluation of SARS-CoV-2 Detection Efficiency by qRT-PCR Assays, Mainly on New Variants in East Java
DOI:
https://doi.org/10.48048/tis.2025.10253Keywords:
COVID-19, In silico, Molecular diagnostic, Mutations, Primer and probe mismatches, qRT-PCR, SARS-CoV-2, VariantsAbstract
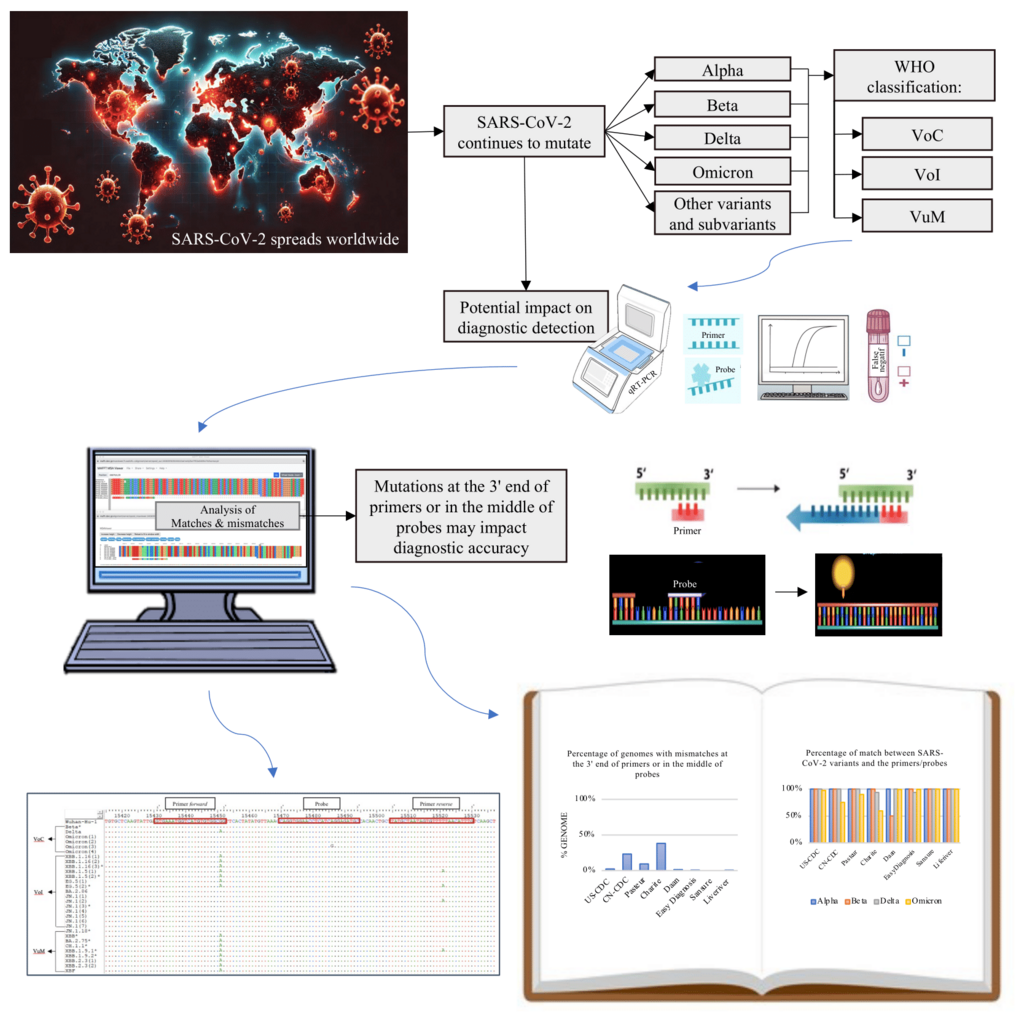
Coronavirus disease 2019 (COVID-19) is an infectious disease caused by the novel coronavirus, SARS-CoV-2, which led to a global pandemic. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) has undergone mutations, leading to the emergence of new variants worldwide, which may affect the accuracy of the diagnostic methods. This study aimed to identify the distribution of SARS-CoV-2 variants in East Java from 2021 to 2024 and analyze the mutation positions in the primer/probe targets of quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR). This study assessed the mismatches in WHO-recommended and other kits binding regions qRT-PCR assays against SARS-CoV-2 genome sequences from East Java through in-silico bioinformatics analysis. The results indicate that from 2021 to 2024, the distribution of SARS-CoV-2 variants in East Java has changed over time, including B.1.466.2, Alpha, Beta, Delta, and Omicron, along with various Omicron lineages. Primer and probe sequences from Sansure, Liferiver, US-CDC, EasyDiagnosis, and Pasteur displayed accurate matches (> 90%) with the SARS-CoV-2 sequences from East Java. On the other hand, many Omicron subvariants in East Java had high mismatches with the primer sequences from Charité-RdRp and CN-CDC-N. These findings highlight the importance of continuous surveillance of circulating variants and their mutations to ensure the effectiveness of diagnostics and response actions for emerging and re-emerging variants in the future.
HIGHLIGHTS
- Comprehensive analysis of SARS-CoV-2 variants in East Java, Indonesia.
- Several analyzed SARS-CoV-2 variants have mutations in critical regions, particularly at the 3' end of the primer or the middle of the probe.
- Mutations that may impact the result of qRT-PCR, especially in new Omicron lineages, highlight the necessity for continuous surveillance.
- By identifying mismatches, this study provides essential data to enhance test accuracy.
- The findings can serve as a guide for managing emerging and re-emerging variants in the future.
GRAPHICAL ABSTRACT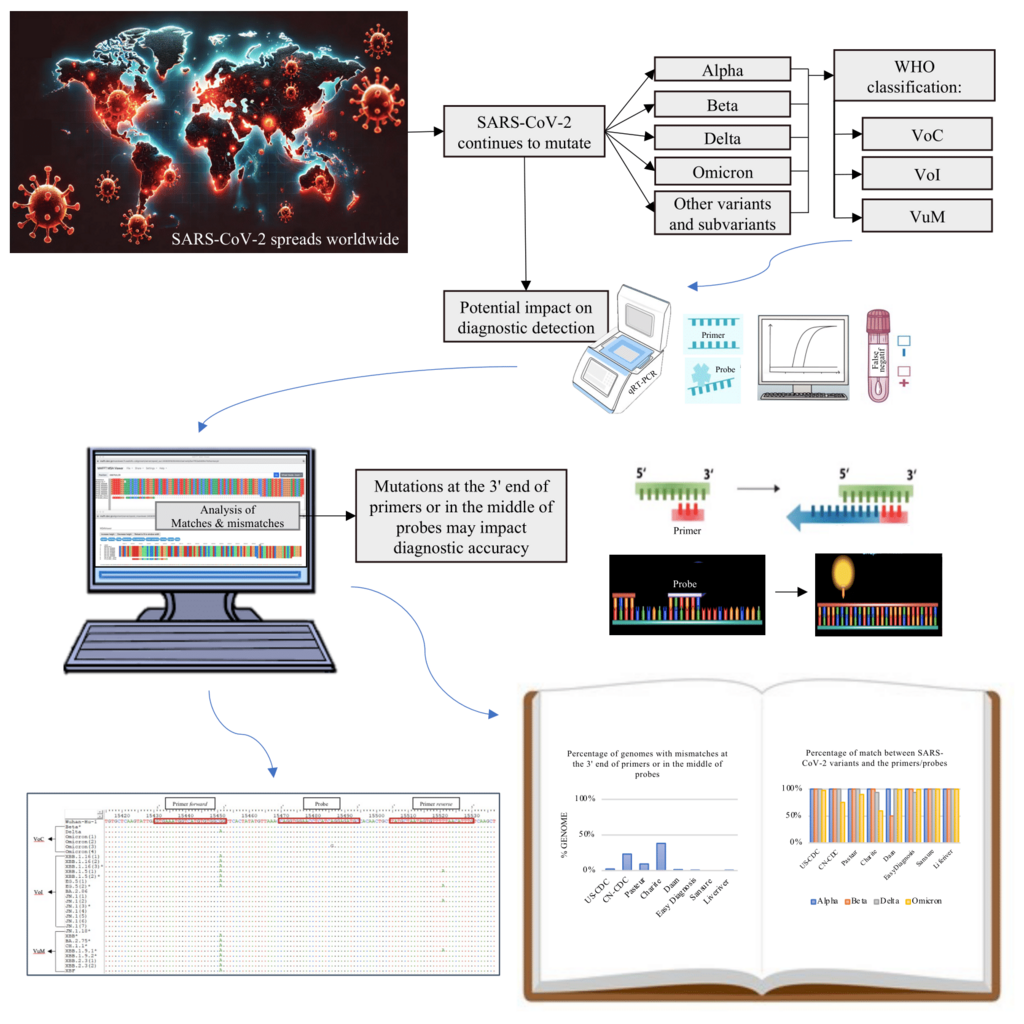
Downloads
References
World Health Organization, WHO COVID-19 cases, Available at: https://data.who.int/dashboards/covid19/cases, accessed February 2025.
C Chhoung, K Ko, S Ouoba, Z Phyo, GA Akuffo, A Sugiyama, T Akita, H Sasaki, T Yamamoto, K Takahashi and J Tanaka. Sustained applicability of SARS-CoV-2 variants identification by Sanger Sequencing Strategy on emerging various SARS-CoV-2 Omicron variants in Hiroshima, Japan. BMC Genomics 2024; 25(1), 1063.
S Bhattacharyya. Analysis of mutational profiles of SARS-CoV-2 structural and non-structural proteins with emphasis on spike protein variants. Trends in Sciences 2022; 19(17), 1-9.
World Health Organization, Tracking SARS-CoV-2 variants, Available at: https://www.who.int/activities/tracking-SARS-CoV-2-variants, accesed March 2025.
SD Thepade and H Jha. Covid-19 identification using machine learning classifiers with GLCM features of chest x-ray images. Trends in Sciences 2021; 18(23), 46.
World Health Organization, Molecular Assays to diagnose COVID-19, Available at: https://www.who.int/docs/defaultsource/coronaviruse/whoinhouseassays.pdf, accesed February 2025.
A Marchini, M Petrillo, A Parrish, G Buttinger, S Tavazzi, M Querci, F Betsou, G Elsinga, G Medema, T Abdelrahman, B Gawlik and P Corbisier. New RT-PCR assay for the detection of current and future SARS-CoV-2 variants. Viruses 2023; 15(1), 206.
Y Chen, Y Han, J Yang, Y Ma, J Li and R Zhang. Impact of SARS-CoV-2 variants on the analytical sensitivity of rRT-PCR assays. Journal of Clinical Microbiology 2022; 60(4), e0237421.
Food and drug administration (FDA). SARS-CoV-2 viral mutations: Impact on COVID-19 tests, Available at: https://www.fda.gov/medical-devices/coronavirus-covid-19-and-medical-devices/sars-cov-2-viral-mutations-impact-covid-19-tests, accesed March 2025.
K Mani, K Thirumalmuthu, DS Kathiresan, S Ramalingam, R Sankaran and S Jeyaraj. In-silico analysis of Covid-19 genome sequences of Indian origin: Impact of mutations in identification of SARS-Co-V2. Molecular and Cellular Probes 2021; 58(6), 101748.
M Alkhatib, L Carioti, S D’Anna, F Ceccherini-Silberstein, V Svicher and R Salpini. SARS-CoV-2 mutations and variants may muddle the sensitivity of COVID-19 diagnostic assays. Microorganisms 2022; 10(8), 1559.
MMH Sarkar, SR Naser, SF Chowdhury, MS Khan, MA Habib, S Akter, TA Banu, B Goswami, I Jahan, MR Nayem, MA Hassan, I Khan, MFA Rabbi, CR Ahsan, MI Miah, A Nessa, SMRU Islam, MA Rahman, MAA Shaikh and MS Ahmed. M gene targeted qRT-PCR approach for SARS-CoV-2 virus detection. Scientific Reports 2023; 13(1), 16659.
Y Shu and J McCauley. GISAID: Global initiative on sharing all influenza data - from vision to reality. Eurosurveillance 2017; 22(13), 30494.
V Setiawaty, H Kosasih, Y Mardian, E Ajis, EB Prasetyowati, Siswanto and M Karyana. The identification of first COVID-19 cluster in Indonesia. American Journal of Tropical Medicine and Hygiene 2020; 103(6), 2339-2342.
C Trobajo-Sanmartín, I Martínez-Baz, A Miqueleiz, M Fernández-Huerta, C Burgui, I Casado, F Baigorría, A Navascués, J Castilla and C Ezpeleta. Differences in transmission between SARS-CoV-2 Alpha (B.1.1.7) and delta (B.1.617.2) variants. Microbiology Spectrum 2022; 10(2), e0000822.
FA Alsuwairi, AN Alsaleh, DA Obeid, AA Al-Qahtani, RS Almaghrabi, BM Alahideb, MA AlAbdulkareem, MS Alsanea, LA Alharbi, SI Althawadi, SA Altamimi, AN Alshukairi and FS Alhamlan. Genomic surveillance and mutation analysis of SARS-CoV-2 variants among patients in Saudi Arabia. Microorganisms 2024; 12(3), 467.
I Cahyani, EW Putro, AM Ridwanuloh, S Wibowo, H Hariyatun, G Syahputra, G Akbariani, AR Utomo, M Ilyas, M Loose, W Kusharyoto and S Susanti. Genome Profiling of SARS-CoV-2 in Indonesia, ASEAN and the neighbouring east Asian countries: Features, challenges and achievements. Viruses 2022; 14(4), 778.
MN Massi, R Sjahril, H Halik, G Vita Soraya, N Hidayah, MY Pratama, M Faruk, I Handayani, FM Gazali, MS Hakim and T Wibawa. Sequence analysis of SARS-CoV-2 Delta variant isolated from Makassar, South Sulawesi, Indonesia. Heliyon 2023; 9(2), e13382.
R Bestari, IRA Nainggolan, M Hasibuan, R Ratnanggana, K Rahardjo, AM Nastri, JR Dewantari, S Soetjipto, MI Lusida, Y Mori, K Shimizu, RL Kusumawati, M Ichwan and IND Lubis. SARS-CoV-2 lineages circulating during the first wave of the pandemic in North Sumatra, Indonesia. IJID Regions 2023; 8, S1-S7.
L Linosefa, H Hasmiwati, J Jamsari and AE Putra. Genomic analysis of three SARS-CoV-2 waves in west Sumatra: Evolutionary dynamics and variant clustering. Heliyon 2024; 10(14), e34365.
K Dhama, F Nainu, A Frediansyah, MI Yatoo, RK Mohapatra, S Chakraborty, H Zhou, MR Islam, SS Mamada, HI Kusuma, AA Rabaan, S Alhumaid, AA Mutair, M Iqhrammullah, JA Al-Tawfiq, MA Mohaini, AJ Alsalman, HS Tuli, C Chakraborty and H Harapan. Global emerging Omicron variant of SARS-CoV-2: Impacts, challenges and strategies. Journal of Infection and Public Health 2023; 16(1), 4-14.
D Planas, I Staropoli, V Michel, F Lemoine, F Donati, M Prot, F Porrot, F Guivel-Benhassine, B Jeyarajah, A Brisebarre, O Dehan, L Avon, WH Bolland, M Hubert, J Buchrieser, T Vanhoucke, P Rosenbaum, D Veyer, H Péré, B Lina, …, O Schwartz. Distinct evolution of SARS-CoV-2 Omicron XBB and BA.2.86/JN.1 lineages combining increased fitness and antibody evasion. Nature Communications 2024; 15(1), 2254.
S Zhou, P Lv, M Li, Z Chen, H Xin, S Reilly and X Zhang. SARS-CoV-2 E protein: Pathogenesis and potential therapeutic development. Biomedicine and Pharmacotherapy 2023; 159, 114242.
D Eskier, A Suner, G Karakülah and Y Oktay. Mutation density changes in SARS-CoV-2 are related to the pandemic stage but to a lesser extent in the dominant strain with mutations in spike and RdRp. PeerJ 2020; 8, e9703.
R Wang, Y Hozumi, C Yin and G Wei. Mutations on COVID-19 diagnostic targets. Genomics 2020; 112(6), 5204-5213.
S Islam, T Islam and MR Islam. New coronavirus variants are creating more challenges to global healthcare system: A brief report on the current knowledge. Clinical Pathology 2022; 15, 2632010X221075584.
R Stadhouders, SD Pas, J Anber, J Voermans, THM Mes and M Schutten. The effect of primer-template mismatches on the detection and quantification of nucleic acids using the 5′ nuclease assay. Journal of Molecular Diagnostics 2010; 12(1), 109-117.
A Mentes, K Papp, D Visontai, J Stéger, VEO Technical Working Group, I Csabai, A Medgyes-Horváth and OA Pipek. Identification of mutations in SARS-CoV-2 PCR primer regions. Scientific Reports 2022; 12(1), 18651.
KA Khan and P Cheung. Presence of mismatches between diagnostic PCR assays and coronavirus SARS-CoV-2 genome: Sequence mismatches in SARS-CoV-2 PCR. Royal Society Open Science 2020; 7(6), 200636.
S Bustin, S Kirvell, JF Huggett and T Nolan. RT-qPCR diagnostics: The “drosten” SARS-CoV-2 assay paradigm. International Journal of Molecular Sciences 2021; 22(16), 8702.
T Naiser, O Ehler, J Kayser, T Mai, W Michel and A Ott. Impact of point-mutations on the hybridization affinity of surface-bound DNA/DNA and RNA/DNA oligonucleotide-duplexes: Comparison of single base mismatches and base bulges. BMC Biotechnology 2008; 8, 48.
RR Shirima, EN Wosula, AA Hamza, NA Mohammed, H Mouigni, S Nouhou, NM Mchinda, G Ceasar, M Amour, E Njukwe and JP Legg. Epidemiological analysis of cassava mosaic and brown streak diseases, and Bemisia tabaci in the Comoros Islands. Viruses 2022; 14(10), 2156.
JP Cuff, JJN Kitson, D Hemprich-Bennett, MPTG Tercel, SS Browett and DM Evans. The predator problem and PCR primers in molecular dietary analysis: Swamped or silenced; depth or breadth?. Molecular Ecology Resources 2023; 23(1), 41-51.
MT Dorak. Real-time PCR. Taylor & Francis, London, 2007, p. 333.
C Basu. PCR primer design. 2nd Eds., Humana Press, New York, 2015, p. 276.
S Liu, X Chai, C Liu, J Bai, J Meng, H Tian, X Han, G Han, Q Li and X Xu. Sensitivity analysis of RT-qPCR and RT-ddPCR for SARS-CoV-2 detection with mutations on N1 and E primer-probe region. Microbiology Spectrum 2024; 12(8), e04292-23.
P Gupta, V Gupta, CM Singh and L Singhal. Emergence of COVID-19 Variants: An update. Cureus 2023; 15(7), e41295.
M Vanaerschot, SA Mann, JT Webber, J Kamm, SM Bell, J Bell, SN Hong, MP Nguyen, LY Chan, KD Bhatt, M Tan, AM Detweiler, A Espinosa, W Wu, J Batson, D Dynerman, DA Wadford, AS Puschnik, N Neff, V Ahyong, S Miller, P Ayscue, CM Tato, S Paul, AL Kistler, JL DeRisi and ED Crawford. Identification of a polymorphism in the N Gene of SARS-CoV-2 that adversely impacts detection by reverse transcription- PCR. Journal of Clinical Microbiology 2020; 59(1), e02369-20.
C Moore, L Davies, R Rees, L Gifford, H Lewis, A Plimmer, A Barratt, N Pacchiarini, J Southgate, MJ Bull, J Watkins, S Corden and TR Connor. Localised community circulation of SARS-CoV-2 viruses with an increased accumulation of single nucleotide polymorphisms that adversely affect the sensitivity of real-time reverse transcription assays targeting Nucleocapsid protein, Available at: https://doi.org/10.1101/2021.03.22.21254006, accessed on January 2025.
S Fox-Lewis, A Fox-Lewis, J Harrower, R Chen, J Wang, JD Ligt, G McAuliffe, S Taylor and E Smit. Lack of N2-gene amplification on the Cepheid Xpert Xpress SARS-CoV-2 assay and potential novel causative mutations: A case series from Auckland, New Zealand. IDCases 2021; 25(39), e01233.
D Sharma, KI Notarte, RA Fernandez, G Lippi, MM Gromiha and BM Henry. In silico evaluation of the impact of Omicron variant of concern sublineage BA.4 and BA.5 on the sensitivity of RT-qPCR assays for SARS-CoV-2 detection using whole genome sequencing. Journal of Medical Virology 2023; 95(1), e28241.
M Artesi, S Bontems, P Göbbels, M Franckh, P Maes, R Boreux, C Meex, P Melin, M Hayette, V Bours and K Durkin. A recurrent mutation at position 26340 of SARS-CoV-2 is associated with failure of the e gene quantitative reverse transcription-pcr utilized in a commercial dual-target diagnostic assay. Journal of Clinical Microbiology 2020; 58(10), e01598.
K Ziegler, P Steininger, R Ziegler, J Steinmann, K Korn and A Ensser. SARS-CoV-2 samples may escape detection because of a single point mutation in the N gene. Eurosurveillance 2020; 25(39), 2001650.
KKK Ko, N Binte A Rahman, SYL Tan, KXL Chan, SS Goh, JHC Sim, KL Lim, WL Tan, KS Chan and LLE Oon. SARS-CoV-2 N gene G29195T point mutation may affect diagnostic reverse transcription-PCR detection. Microbiology Spectrum 2022; 10(1), e02223-21.
GR McCracken, D Gaston, J Pettipas, A Loder, A Majer, E Grudeski, G Labbé, BK Joy, G Patriquin and JJ LeBlanc. Neglected SARS-CoV-2 variants and potential concerns for molecular diagnostics: A framework for nucleic acid amplification test target site quality assurance. Microbiology Spectrum 2023; 11(6), e0076123.
F Roy, J Beirnes, JJ Leblanc, S Gibbons, A Sivro and A Severini. Surveillance for Chlamydia trachomatis variants escaping detection with the Aptima Combo 2 assay in Canada from 2019 to 2021. Microbiology Spectrum 2025; 13(3), e0206224.
B Ashwood, MS Jones, Y Lee, JR Sachleben, AL Ferguson and A Tokmakoff. Molecular insight into how the position of an abasic site modifies DNA duplex stability and dynamics. Biophysical Journal 2024; 123(2), 118-133.
A Bulut, OZYesilel, N Dege, H Icbudak, H Olmez and O Buyukgungor. Dinicotinamidium squarate. Acta Crystallographica Section C: Structural Chemistry 2003; 59(12), o727-o729.
S Genç, N Dege, A Çetin, A Cansiz, M Şekerci and M Dinçer. 3-(2-Hydroxyphenyl)-4-phenyl-1 H-1,2,4-triazole-5(4H)-thione. Acta Crystallographica Section E: Crystallographic Communications 2004; 60(9), 1580-1582.
N Dege, H Içbudak and E Adiyaman. Bis(acesulfamato-κO,N)bis(3-methylpyridine)copper(II). Acta Crystallographica Section C: Structural Chemistry 2006; 62(P9), 401-403.
MN Tahir, M Ashfaq, M Feizi-Dehnayebi, KS Munawar, S Atalay, N Dege, N Guliyeva and A Sultan. Crystal structure, Hirshfeld surface analysis, computational study and molecular docking simulation of 4-aminoantipyrine derivative. Journal of Molecular Structure 2025; 1320,139747.
ATS Bodapati, RS Reddy, K Lavanya, SR Madku and BK Sahoo. Minor groove binding of antihistamine drug bilastine with calf thymus DNA: A molecular perspective with thermodynamics using experimental and theoretical methods. Journal of Molecular Structure 2024; 1301, 137385.
N Arumugam, AI Almansour, RS Kumar, VS Krishna, D Sriram and N Dege. Stereoselective synthesis and discovery of novel spirooxindolopyrrolidine engrafted indandione heterocyclic hybrids as antimycobacterial agents. Bioorganic Chemistry 2021; 110, 104798.
J Kashyap and D Datta. Drug repurposing for SARS-CoV-2: A high-throughput molecular docking, molecular dynamics, machine learning, and DFT study. Journal of Materials Science 2022; 57(23), 10780-10802.
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.






