Preliminary Computational Analysis of Pyrazinamide-Based Derivatives Reveals Possible Inhibition of SARS-CoV-2 RNA-Dependent RNA Polymerase, and Their Possible Use as Antiviral Agents
DOI:
https://doi.org/10.48048/tis.2022.3918Keywords:
Pyrazinamide, Favipiravir, In silico analysis, COVID-19, SARS-CoV-2, RNA-dependent RNA polymeraseAbstract
Pyrazinamide is a pyrazine analog currently used to treat tuberculosis. It shares a core structure similar to that of favipiravir, which is a promising drug candidate that may inhibit the SARS-CoV-2 RNA-dependent RNA polymerase. This feature could be an opportunity for further drug development of anti-COVID-19 medication starting from a pyrazinamide core structure. This study aimed to determine and predict the most effective pyrazinamide-based analogs against the SARS-CoV-2 RNA-dependent RNA polymerase by using combined ligand- and structure-based computational analysis. This study performed a rational in silico study to screen pyrazinamide-like molecules from a commercially available ZINC database, with a similarity score higher than 0.40, and then these screened for acceptable pharmacokinetic properties, and then to further conduct molecular docking analysis with SARS-CoV-2 RNA-dependent RNA polymerase. The results showed that compound 12, having a dichloropyrimidine core structure had a similarity score of 0.446. It exerted the most binding affinity with RNA-dependent RNA polymerase, with estimated docking scores of −5.72, −5.25, −7.06, −7.00 and −4.63 kcal/mol in intact, ribosylated, mono-phosphoribosylated, di-phosphoribosylated and tri-phosphoribosylated forms, respectively. Watson-Crick base-pairing of compound 12 indicated that it favored binding with the uracil nucleoside of the RNA template. Compound 12 was confirmed as the lead compound, being a pyrazinamide-like molecule, and so might be a most promising candidate molecule, as an adenine analog RNA-dependent RNA polymerase inhibitor. It is suggested that the antiviral effect of this lead compound should be studied further as part of a drug discovery and development process.
HIGHLIGHTS
- In silico analysis was performed to screen pyrazinamide-like compounds against the RNA-dependent RNA polymerase
- Pharmacokinetics and drug-likeliness properties of fourteen preselected compounds were assessed by using SwissADME with favorable results
- Among these preselected compounds, compound 12 showed the best docking score in almost all of its phosphoribosylated forms
GRAPHICAL ABSTRACT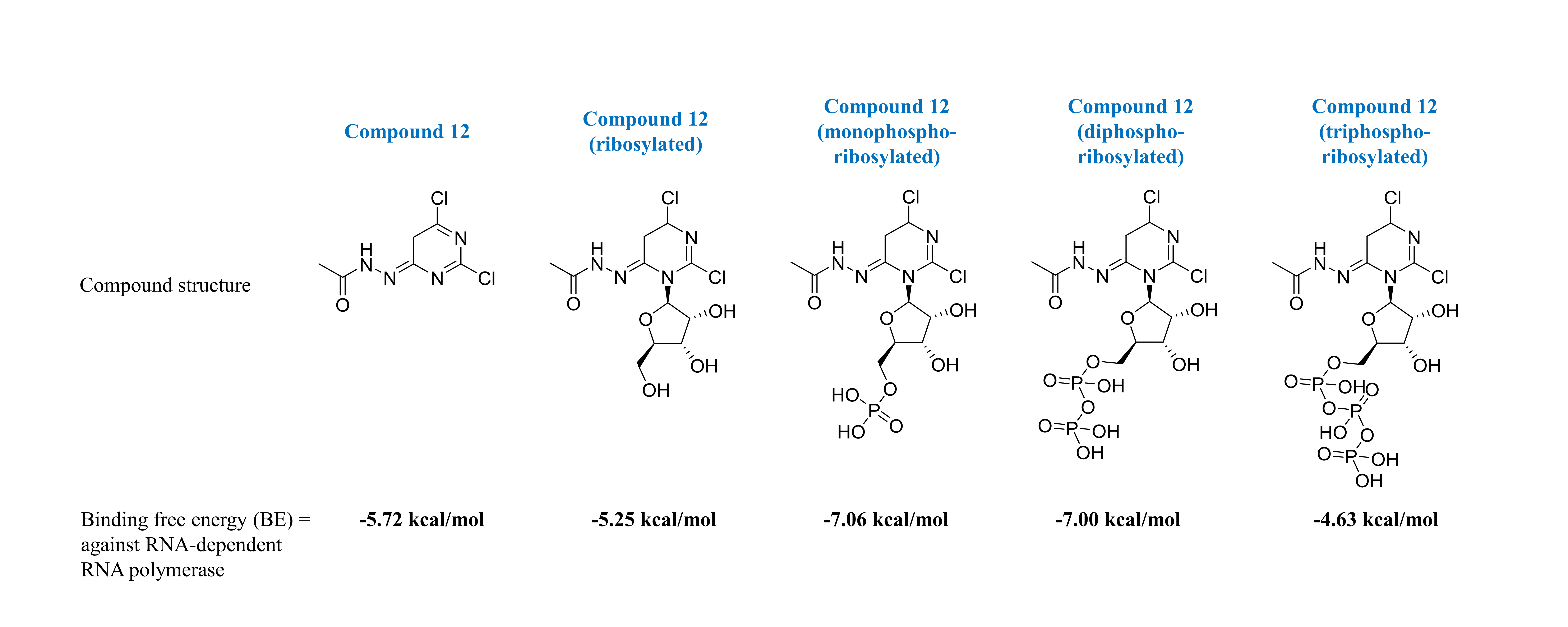
Downloads
References
World Health Organization. Therapeutics and COVID-19: Living guideline. World Health Organization, Geneva, 2020, p. 16-56.
SSA Karim and QA Karim. Omicron SARS-CoV-2 variant: A new chapter in the COVID-19 pandemic. Lancet 2021; 398, 2126-8.
P V’Kovski, A Kratzel, S Steiner, H Stalder and V Thiel. Coronavirus biology and replication: Implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021; 19, 155-70.
WF Zhang, P Stephen, JF Thériault, R Wang and SX Lin. Novel coronavirus polymerase and nucleotidyl-transferase structures: Potential to target new outbreaks. J. Phys. Chem. Lett. 2020; 11, 4430-5.
SO Aftab, MZ Ghouri, MU Masood, Z Haider, Z Khan, A Ahmad and N Munawar. Analysis of SARS-CoV-2 RNA-dependent RNA polymerase as a potential therapeutic drug target using a computational approach. J. Transl. Med. 2020; 18, 275.
AJ Bernal, MMG da Silva, DB Musungaie, E Kovalchuk, A Gonzalez, VD Reyes, A Martín-Quirós, Y Caraco, A Williams-Diaz, ML Brown, J Du, A Pedley, C Assaid, J Strizki, JA Grobler, HH Shamsuddin, R Tipping, H Wan, A Paschke, …, CD Anda. Molnupiravir for oral treatment of covid-19 in nonhospitalized patients. New Engl. J. Med. 2021; 386, 509-20.
JG Julander, JF Demarest, R Taylor, BB Gowen, DM Walling, A Mathis and YS Babu. An update on the progress of galidesivir (BCX4430), a broad-spectrum antiviral. Antivir. Res. 2021; 195, 105180.
JH Beigel, KM Tomashek, LE Dodd, AK Mehta, BS Zingman, AC Kalil, E Hohmann, HY Chu, A Luetkemeyer, S Kline, DL Castilla, RW Finberg, K Dierberg, V Tapson, L Hsieh, TF Patterson, R Paredes, DA Sweeney, WR Short, …, HC Lane. Remdesivir for the treatment of Covid-19 - final report. New Engl. J. Med. 2020; 383, 1813-26.
ZF Udwadia, P Singh, H Barkate, S Patil, S Rangwala, A Pendse, J Kadam, W Wu, CF Caracta and M Tandon. Efficacy and safety of favipiravir, an oral RNA-dependent RNA polymerase inhibitor, in mild-to-moderate COVID-19: A randomized, comparative, open-label, multicenter, phase 3 clinical trial. Int. J. Infect. Dis. 2021; 103, 62-71.
H Li, N Xiong, C Li, Y Gong, L Liu, H Yang, X Tan, N Jiang, Q Zong, J Wang, Z Lu and X Yin. Efficacy of ribavirin and interferon-α therapy for hospitalized patients with COVID-19: A multicenter, retrospective cohort study. Int. J. Infect. Dis. 2021; 104, 641-8.
RA Al-Horani and S Kar. Potential anti-SARS-CoV-2 therapeutics that target the post-entry stages of the viral life cycle: A comprehensive review. Viruses 2020; 12, 1092.
M Bosaeed, A Alharbi, E Mahmoud, S Alrehily, M Bahlaq, Z Gaifer, H Alturkistani, K Alhagan, S Alshahrani, A Tolbah, A Musattat, M Alanazi, R Jaha, K Sultana, H Alqahtani, KA Aamer, S Jaser, A Alsaedy, A Ahmad, …, A Alaskar. Efficacy of favipiravir in adults with mild COVID-19: A randomized, double-blind, multicentre, placebo-controlled clinical trial. Clin. Microbiol. Infect. 2022; 28, 602-8.
L Zhao and W Zhong. Mechanism of action of favipiravir against SARS-CoV-2: Mutagenesis or chain termination. Innovation 2021; 2, 100165.
O Jandourek, M Tauchman, P Paterova, K Konecna, L Navratilova, V Kubicek, O Holas, J Zitko and M Dolezal. Synthesis of novel pyrazinamide derivatives based on 3-chloropyrazine-2-carboxamide and their antimicrobial evaluation. Molecules 2017; 22, 223.
A Sethi, K Joshi, K Sasikala and M Alvala. Molecular docking in modern drug discovery: Principles and recent applications. In: V Gaitonde, P Karmakar and A Trivedi (Ed.). Drug discovery and development - new advances. IntechOpen, London, 2019, p. 1-21.
A Shannon, B Selisko, NT Le, J Huchting, F Touret, G Piorkowski, V Fattorini, F Ferron, E Decroly, C Meier, B Coutard, O Peersen and B Canard. Rapid incorporation of favipiravir by the fast and permissive viral RNA polymerase complex results in SARS-CoV-2 lethal mutagenesis. Nat. Comm. 2020; 11, 4682.
V Zoete, A Daina, C Bovigny and O Michielin. SwissSimilarity: A web tool for low to ultra high throughput ligand-based virtual screening. J. Chem. Inform. Model. 2016; 56, 1399-404.
A Daina, O Michielin and V Zoete. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017; 7, 42717.
ÍF Protti, DR Rodrigues, SK Fonseca, RJ Alves, RB de Oliveira and VG Maltarollo. Do drug-likeness rules apply to oral prodrugs. Chem. Med. Chem. 2021; 16, 1446-56.
W Yin, C Mao, X Luan, DD Shen, Q Shen, H Su, X Wang, F Zhou, W Zhao, M Gao, S Chang, YC Xie, G Tian, HW Jiang, SC Tao, J Shen, Y Jiang, H Jiang, Y Xu, …, HE Xu. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 2020; 368, 1499-504.
WFA Junior, G Bitencourt-Ferreira, JR Godoy, HMA Adriano, WADS Bezerra and AMDS Soares. Protein-ligand docking simulations with AutoDock4 focused on the main protease of SARS-CoV-2. Curr. Med. Chem. 2021; 28, 7614-33.
Q Li and S Shah. Structure-based virtual screening. Meth. Mol. Biol. 2017; 1558, 111-24.
M Merski, J Skrzeczkowski, JK Roth and MW Górna. A Geometric definition of short to medium range hydrogen-mediated interactions in proteins. Molecules 2020; 25, 5326.
RAC Siemieniuk, JJ Bartoszko, L Ge, D Zeraatkar, A Izcovich, E Kum, H Pardo-Hernandez, A Qasim, JPD Martinez, B Rochwerg, F Lamontagne, MA Han, Q Liu, A Agarwal, T Agoritsas, DK Chu, R Couban, E Cusano, A Darzi, …, R Brignardello-Petersen. Drug treatments for covid-19: Living systematic review and network meta-analysis. BMJ 2020; 370, 2980.
I Celik, M Erol and Z Duzgun. In silico evaluation of potential inhibitory activity of remdesivir, favipiravir, ribavirin and galidesivir active forms on SARS-CoV-2 RNA polymerase. Mol. Divers. 2021; 26, 279-92.
MO Rafi, G Bhattacharje, K Al-Khafaji, T Taskin-Tok, MA Alfasane, AK Das, MAK Parvez and MS Rahman. Combination of QSAR, molecular docking, molecular dynamic simulation and MM-PBSA: Analogues of lopinavir and favipiravir as potential drug candidates against COVID-19. J. Biomol. Struct. Dynam. 2020; 40, 3711-30.
P Shende, B Khanolkar and RS Gaud. Drug repurposing: New strategies for addressing COVID-19 outbreak. Expet. Rev. Anti Infect. Ther. 2021; 19, 689-706.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2022 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.







